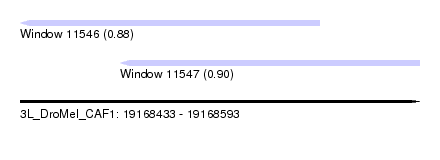
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 19,168,433 – 19,168,593 |
| Length | 160 |
| Max. P | 0.899814 |

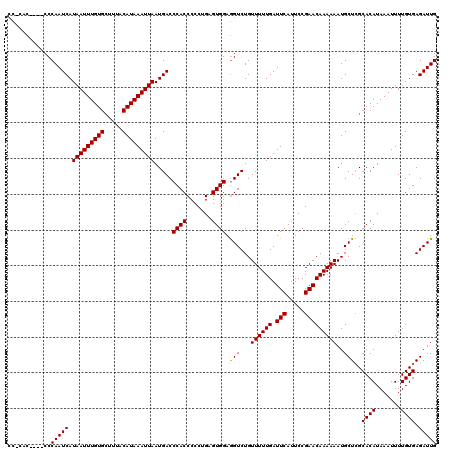
| Location | 19,168,433 – 19,168,553 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 92.60 |
| Mean single sequence MFE | -24.48 |
| Consensus MFE | -21.54 |
| Energy contribution | -21.35 |
| Covariance contribution | -0.19 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.86 |
| Structure conservation index | 0.88 |
| SVM decision value | 0.92 |
| SVM RNA-class probability | 0.882618 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 19168433 120 - 23771897 CCUUACUCCACCCAAUCAUAAUUUGUGAUUUACAUAAAUUAAUGACCCACCCCCUGAGUGGAGGUCUGUUUUUGAUUCAUUCCGAACAAAAAAUGCUCGCACAUAAAUUUUGUGAGAUUG .....((((((.((.((((((((((((.....)))))))).)))).........)).)))))).....((((((.(((.....)))))))))...((((((.........)))))).... ( -25.10) >DroSim_CAF1 1223 120 - 1 CCCCACUCCACCCAAUCAUAAUUUGUGCUUUACAUAAAUUAAUGACCCACCCCCAGAGUGGAGGUCUGUUUUUGAUUCAUUCCGAACAAAAAAUGCUCGCACAUAAAUUUUGUGAGAUUG ((((((((.......((((((((((((.....)))))))).))))..........)))))).))....((((((.(((.....)))))))))...((((((.........)))))).... ( -26.53) >DroEre_CAF1 942 110 - 1 ----------CCCAAUCAUAAUUUGUGCUUUACAUAAAUUAAUGACCCACCCCCUGAGUGGAAGUCUGUUUUUGAUUCAUUCCGAACAAAAAAUGCUAGCACAUAAAUUUUGUGAGAUUG ----------.....((((((..((((((...(((........(((((((.......))))..)))..((((((.(((.....))))))))))))..))))))......))))))..... ( -22.30) >DroYak_CAF1 2674 116 - 1 CCACAC----CCCAAUCAUAAUUUGUGCUUUACAUAAAUUAAUGACCCACCCCCCGAGUGGAGGUCUGUUUUUGAUUCAUUCCGAACAAAAAAUGCUCGCACAUAAAUUUUGUGAGAUUG ..(((.----.((..((((((((((((.....)))))))).)))).((((.......)))).))..)))(((((.(((.....))))))))....((((((.........)))))).... ( -24.00) >consensus CC_CAC____CCCAAUCAUAAUUUGUGCUUUACAUAAAUUAAUGACCCACCCCCUGAGUGGAGGUCUGUUUUUGAUUCAUUCCGAACAAAAAAUGCUCGCACAUAAAUUUUGUGAGAUUG ............(((((.(((((((((.....))))))))).....((((.......)))).(((...((((((.(((.....)))))))))..)))..((((.......)))).))))) (-21.54 = -21.35 + -0.19)


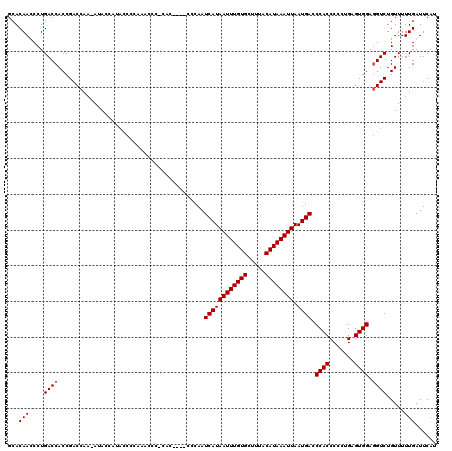
| Location | 19,168,473 – 19,168,593 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 85.87 |
| Mean single sequence MFE | -17.83 |
| Consensus MFE | -16.88 |
| Energy contribution | -17.12 |
| Covariance contribution | 0.25 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.69 |
| Structure conservation index | 0.95 |
| SVM decision value | 1.01 |
| SVM RNA-class probability | 0.899814 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 19168473 120 - 23771897 GCACAAUCCUGACCACCGACCAAAAUACUAUACCCCAAGCCCUUACUCCACCCAAUCAUAAUUUGUGAUUUACAUAAAUUAAUGACCCACCCCCUGAGUGGAGGUCUGUUUUUGAUUCAU ...(((....((((.........................................((((((((((((.....)))))))).)))).((((.......)))).)))).....)))...... ( -18.90) >DroSim_CAF1 1263 120 - 1 GCACAAUCCUGACCACCGACCAAAAUACUACACCCCAAACCCCCACUCCACCCAAUCAUAAUUUGUGCUUUACAUAAAUUAAUGACCCACCCCCAGAGUGGAGGUCUGUUUUUGAUUCAU ...................................((.(((.((((((.......((((((((((((.....)))))))).))))..........)))))).))).))............ ( -20.23) >DroEre_CAF1 982 99 - 1 GCACAACCCUGACCACCGACCAA-AUACCA--------------------CCCAAUCAUAAUUUGUGCUUUACAUAAAUUAAUGACCCACCCCCUGAGUGGAAGUCUGUUUUUGAUUCAU ..............((.(((...-......--------------------.....((((((((((((.....)))))))).)))).((((.......))))..))).))........... ( -14.00) >DroYak_CAF1 2714 115 - 1 GCACAACCCUGACCACCGACCAA-AUACCGUACCCUAAACCCACAC----CCCAAUCAUAAUUUGUGCUUUACAUAAAUUAAUGACCCACCCCCCGAGUGGAGGUCUGUUUUUGAUUCAU ..............((.((((..-......................----.....((((((((((((.....)))))))).)))).((((.......)))).)))).))........... ( -18.20) >consensus GCACAACCCUGACCACCGACCAA_AUACCAUACCCCAAACCC_CAC____CCCAAUCAUAAUUUGUGCUUUACAUAAAUUAAUGACCCACCCCCUGAGUGGAGGUCUGUUUUUGAUUCAU ...(((....((((.........................................((((((((((((.....)))))))).)))).((((.......)))).)))).....)))...... (-16.88 = -17.12 + 0.25)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:38:17 2006