
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 19,167,468 – 19,167,628 |
| Length | 160 |
| Max. P | 0.993279 |

| Location | 19,167,468 – 19,167,588 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 98.75 |
| Mean single sequence MFE | -35.91 |
| Consensus MFE | -34.37 |
| Energy contribution | -34.43 |
| Covariance contribution | 0.06 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.65 |
| Structure conservation index | 0.96 |
| SVM decision value | -0.07 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 19167468 120 + 23771897 AAUCGAGGGGAAAAUGCAUUUUCCCGCACACCCACAAUUGCCAACACGUGUGCGGCACGAAAACUAAACUGGCGUCAUUUAAUUAACCCGCCAGUGCAGCAGAGGGGUCCGCUUAACCCC ..(((..(((((((....)))))))((((((................))))))....)))...((..(((((((..............)))))))..))....(((((.......))))) ( -35.93) >DroSim_CAF1 263 120 + 1 AAUCGAGGGGAAAAUGCAUUUUCCCGCACACCCACAAUUGCCAACACGUGUGCGGCACGGAAACCAAACUGGCGUCAUUUAAUUAACCCGCCAGUGCAGCAGAGGGGUCCGCUUAACCCC ......((((.....((......((((((((................))))))))...(((..((..(((((((..............))))))).(....).))..)))))....)))) ( -37.73) >DroEre_CAF1 11 120 + 1 AAUCGAGGGGAAAAUGCAUUUUCCCGCACACCCACAAUUGCCAACACGUGUGCGGCACGAAAACUAAACUGGCGUCAUUUAAUUAACCCGCCAGUGCAGCAGAGGGGUCCGCUUAACCCC ..(((..(((((((....)))))))((((((................))))))....)))...((..(((((((..............)))))))..))....(((((.......))))) ( -35.93) >DroYak_CAF1 1752 120 + 1 AAUCGAGGGGAAAAUGCAUUUUCCCGCACUCCCACAAUUGCCAACACGUGUGCGGCACGAAAACUAAACUGGCGUCAUUUAAUUAACCCGCCAGUGCAGCAGAGGGGUCCGCUUAACCCC ..(((..(((((((....))))))).............((((.((....))..)))))))...((..(((((((..............)))))))..))....(((((.......))))) ( -34.04) >consensus AAUCGAGGGGAAAAUGCAUUUUCCCGCACACCCACAAUUGCCAACACGUGUGCGGCACGAAAACUAAACUGGCGUCAUUUAAUUAACCCGCCAGUGCAGCAGAGGGGUCCGCUUAACCCC ......((((....(((.(((((((((((((................))))))))...)))))....(((((((..............))))))))))((.((....)).))....)))) (-34.37 = -34.43 + 0.06)

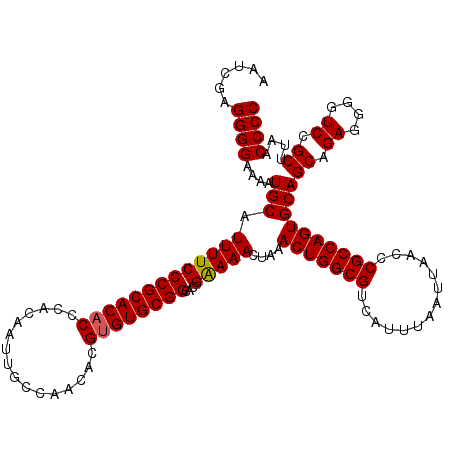
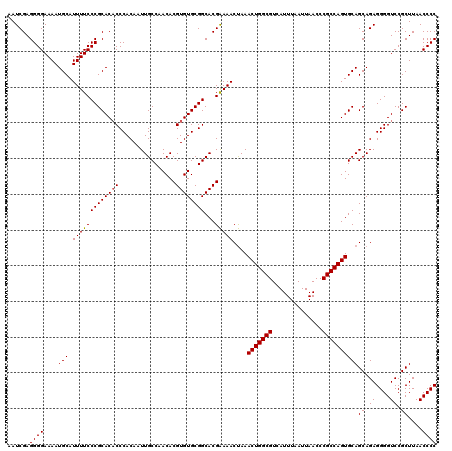
| Location | 19,167,468 – 19,167,588 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 98.75 |
| Mean single sequence MFE | -44.33 |
| Consensus MFE | -43.89 |
| Energy contribution | -43.70 |
| Covariance contribution | -0.19 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.65 |
| Structure conservation index | 0.99 |
| SVM decision value | 0.03 |
| SVM RNA-class probability | 0.548122 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 19167468 120 - 23771897 GGGGUUAAGCGGACCCCUCUGCUGCACUGGCGGGUUAAUUAAAUGACGCCAGUUUAGUUUUCGUGCCGCACACGUGUUGGCAAUUGUGGGUGUGCGGGAAAAUGCAUUUUCCCCUCGAUU (((.....(((.((((....((((.(((((((.(((.....)))..))))))).)))).....(((((((....)).))))).....)))).)))(((((((....)))))))))).... ( -44.10) >DroSim_CAF1 263 120 - 1 GGGGUUAAGCGGACCCCUCUGCUGCACUGGCGGGUUAAUUAAAUGACGCCAGUUUGGUUUCCGUGCCGCACACGUGUUGGCAAUUGUGGGUGUGCGGGAAAAUGCAUUUUCCCCUCGAUU ((((...((((((....))))))(((((((((.(((.....)))..)))))))....((((((..((.((((..((....))..)))).).)..))))))...)).....))))...... ( -45.80) >DroEre_CAF1 11 120 - 1 GGGGUUAAGCGGACCCCUCUGCUGCACUGGCGGGUUAAUUAAAUGACGCCAGUUUAGUUUUCGUGCCGCACACGUGUUGGCAAUUGUGGGUGUGCGGGAAAAUGCAUUUUCCCCUCGAUU (((.....(((.((((....((((.(((((((.(((.....)))..))))))).)))).....(((((((....)).))))).....)))).)))(((((((....)))))))))).... ( -44.10) >DroYak_CAF1 1752 120 - 1 GGGGUUAAGCGGACCCCUCUGCUGCACUGGCGGGUUAAUUAAAUGACGCCAGUUUAGUUUUCGUGCCGCACACGUGUUGGCAAUUGUGGGAGUGCGGGAAAAUGCAUUUUCCCCUCGAUU ((((((.....))))))...((((.(((((((.(((.....)))..))))))).)))).....(((((((....)).)))))((((.(((((((((......)))))..))))..)))). ( -43.30) >consensus GGGGUUAAGCGGACCCCUCUGCUGCACUGGCGGGUUAAUUAAAUGACGCCAGUUUAGUUUUCGUGCCGCACACGUGUUGGCAAUUGUGGGUGUGCGGGAAAAUGCAUUUUCCCCUCGAUU ((((...((((((....))))))(((((((((.(((.....)))..)))))))....((((((..((.((((..((....))..)))).).)..))))))...)).....))))...... (-43.89 = -43.70 + -0.19)



| Location | 19,167,508 – 19,167,628 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 95.42 |
| Mean single sequence MFE | -44.24 |
| Consensus MFE | -41.14 |
| Energy contribution | -42.14 |
| Covariance contribution | 1.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.54 |
| Structure conservation index | 0.93 |
| SVM decision value | 1.84 |
| SVM RNA-class probability | 0.979403 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 19167508 120 + 23771897 CCAACACGUGUGCGGCACGAAAACUAAACUGGCGUCAUUUAAUUAACCCGCCAGUGCAGCAGAGGGGUCCGCUUAACCCCUGCCGCACACGGCGACCACAUGAUGAUUACGAUGGCGACG ......((((((((((.......((..(((((((..............)))))))..))...((((((.......)))))))))))))))).((.((((.(((...))).).)))))... ( -44.54) >DroSim_CAF1 303 120 + 1 CCAACACGUGUGCGGCACGGAAACCAAACUGGCGUCAUUUAAUUAACCCGCCAGUGCAGCAGAGGGGUCCGCUUAACCCCUGCCGCACACGGCGACCACAUGAUGAUUACGAUGGCGACG ......((((((((((..(....)...(((((((..............))))))).......((((((.......)))))))))))))))).((.((((.(((...))).).)))))... ( -44.24) >DroEre_CAF1 51 114 + 1 CCAACACGUGUGCGGCACGAAAACUAAACUGGCGUCAUUUAAUUAACCCGCCAGUGCAGCAGAGGGGUCCGCUUAACCCCUGCCGCACACGGCGACCACAUGAUGAUUGCGAUG------ ......((((((((((.......((..(((((((..............)))))))..))...((((((.......))))))))))))))))((((.((.....)).))))....------ ( -45.74) >DroYak_CAF1 1792 120 + 1 CCAACACGUGUGCGGCACGAAAACUAAACUGGCGUCAUUUAAUUAACCCGCCAGUGCAGCAGAGGGGUCCGCUUAACCCCUGCCACACACGGCGACCACAUGAUGAUUGCGAUGGCGAUG (((.(.((((((.(((.......((..(((((((..............)))))))..))...((((((.......))))))))).))))))((((.((.....)).))))).)))..... ( -42.44) >consensus CCAACACGUGUGCGGCACGAAAACUAAACUGGCGUCAUUUAAUUAACCCGCCAGUGCAGCAGAGGGGUCCGCUUAACCCCUGCCGCACACGGCGACCACAUGAUGAUUACGAUGGCGACG (((.(.((((((((((...........(((((((..............))))))).......((((((.......))))))))))))))))((((.((.....)).))))).)))..... (-41.14 = -42.14 + 1.00)

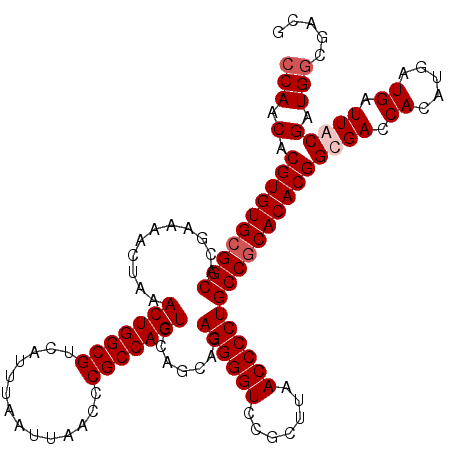

| Location | 19,167,508 – 19,167,628 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 95.42 |
| Mean single sequence MFE | -51.62 |
| Consensus MFE | -50.12 |
| Energy contribution | -49.75 |
| Covariance contribution | -0.38 |
| Combinations/Pair | 1.07 |
| Mean z-score | -2.85 |
| Structure conservation index | 0.97 |
| SVM decision value | 2.39 |
| SVM RNA-class probability | 0.993279 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 19167508 120 - 23771897 CGUCGCCAUCGUAAUCAUCAUGUGGUCGCCGUGUGCGGCAGGGGUUAAGCGGACCCCUCUGCUGCACUGGCGGGUUAAUUAAAUGACGCCAGUUUAGUUUUCGUGCCGCACACGUGUUGG .((.((((.(((.......))))))).))(((((((((((((((((.....))))))...((((.(((((((.(((.....)))..))))))).)))).....)))))))))))...... ( -52.40) >DroSim_CAF1 303 120 - 1 CGUCGCCAUCGUAAUCAUCAUGUGGUCGCCGUGUGCGGCAGGGGUUAAGCGGACCCCUCUGCUGCACUGGCGGGUUAAUUAAAUGACGCCAGUUUGGUUUCCGUGCCGCACACGUGUUGG .((.((((.(((.......))))))).))(((((((((((((((((.....))))))...((((.(((((((.(((.....)))..))))))).)))).....)))))))))))...... ( -51.10) >DroEre_CAF1 51 114 - 1 ------CAUCGCAAUCAUCAUGUGGUCGCCGUGUGCGGCAGGGGUUAAGCGGACCCCUCUGCUGCACUGGCGGGUUAAUUAAAUGACGCCAGUUUAGUUUUCGUGCCGCACACGUGUUGG ------.((((((.......)))))).(((((((((((((((((((.....))))))...((((.(((((((.(((.....)))..))))))).)))).....))))))))))).))... ( -51.20) >DroYak_CAF1 1792 120 - 1 CAUCGCCAUCGCAAUCAUCAUGUGGUCGCCGUGUGUGGCAGGGGUUAAGCGGACCCCUCUGCUGCACUGGCGGGUUAAUUAAAUGACGCCAGUUUAGUUUUCGUGCCGCACACGUGUUGG ....((.((((((.......)))))).))(((((((((((((((((.....))))))...((((.(((((((.(((.....)))..))))))).)))).....)))))))))))...... ( -51.80) >consensus CGUCGCCAUCGCAAUCAUCAUGUGGUCGCCGUGUGCGGCAGGGGUUAAGCGGACCCCUCUGCUGCACUGGCGGGUUAAUUAAAUGACGCCAGUUUAGUUUUCGUGCCGCACACGUGUUGG .....(((.(((.(((((...))))).))(((((((((((((((((.....))))))...((((.(((((((.(((.....)))..))))))).)))).....))))))))))).).))) (-50.12 = -49.75 + -0.38)

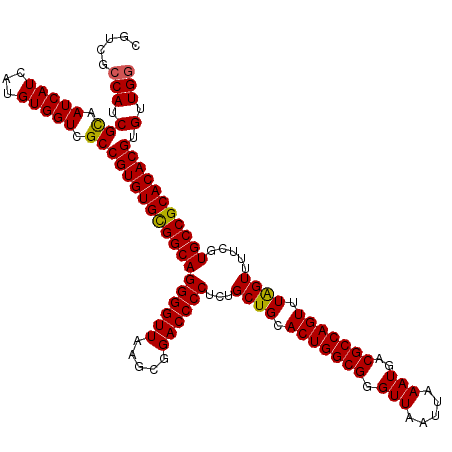
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:38:11 2006