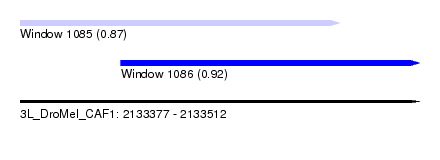
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 2,133,377 – 2,133,512 |
| Length | 135 |
| Max. P | 0.919020 |

| Location | 2,133,377 – 2,133,485 |
|---|---|
| Length | 108 |
| Sequences | 6 |
| Columns | 114 |
| Reading direction | forward |
| Mean pairwise identity | 80.59 |
| Mean single sequence MFE | -39.49 |
| Consensus MFE | -29.82 |
| Energy contribution | -30.83 |
| Covariance contribution | 1.01 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.90 |
| Structure conservation index | 0.76 |
| SVM decision value | 0.88 |
| SVM RNA-class probability | 0.872435 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 2133377 108 + 23771897 UGGUGCCGGUG------CCGGUGCCCGUGCCCGUGGACAUGGCUGUGGACAUGGCCUGACUUGGAGUCGGCGACGAUGAUGACAUUGUUGCUGAAUAAUUGAAGGCGUCUAGCU .((..(((...------.)))..))((((.(((..(......)..))).))))((((.........((((((((((((....))))))))))))........))))........ ( -45.13) >DroGri_CAF1 180921 87 + 1 UGUUGCU------------GGUGCUGGUG---------------AUGUUCGUUGCCUGGCUCGGCGUUGGCGACGAUGAUGACAUUGUUGCUGAAUAAUUGAAGGCGUCUAGCU ....(((------------((((((..((---------------((((((...(((......)))...((((((((((....)))))))))))))).))))..)))).))))). ( -32.60) >DroSec_CAF1 185434 102 + 1 UGGUGCC------------GGUGCCCGUUCCCGUGGACAUGGCUGUGGACAUGGCCUGACUUGGAGUCGGCGACGAUGAUGACAUUGUUGCUGAAUAAUUGAAGGCGUCUAGCU ....((.------------.(((((..((((...((.((.((((((....)))))))).)).))))((((((((((((....)))))))))))).........)))).)..)). ( -36.50) >DroEre_CAF1 190308 102 + 1 UGGUGCCG------------GUGCCCGUGCCCGUGGACAUGGCCGUGGACAUGGCCUGGCUUGGAGUCGGCGACGAUGAUGACAUUGUUGCUGAAUAAUUGAAGGCGUCUAGCU .((.((((------------(...))).)))).(((((..((((((....)))))).(.(((.((.((((((((((((....))))))))))))....)).))).))))))... ( -44.20) >DroYak_CAF1 183203 114 + 1 UGGUGCCGGUGCCGUUGCCGGUGCCCGUGCCCGUGGACAUGGCUGUGGACAUGGCCUGACUUGGAGUCGGCGACGAUGAUGACAUUGUUGCUGAAUAAUUGAAGGCGUCUAGCU .((..(.((..(((....)))..)).)..))..(((((..((((((....))))))...(((.((.((((((((((((....))))))))))))....)).)))..)))))... ( -49.80) >DroAna_CAF1 184621 84 + 1 UGGAGGU------------GGUGCCC------------------GUGGACAUGGCCUGGCUCGGAGUCGGCGACGAUGAUGACAUUGUUGCUGAAUAAUUGAAGGCGUCUAGCU (((((((------------.(((((.------------------..)).))).))).(.(((((..((((((((((((....))))))))))))....))).)).).))))... ( -28.70) >consensus UGGUGCC____________GGUGCCCGUGCCCGUGGACAUGGCUGUGGACAUGGCCUGACUUGGAGUCGGCGACGAUGAUGACAUUGUUGCUGAAUAAUUGAAGGCGUCUAGCU .((((((((........))))))))....................(((((...((((.........((((((((((((....))))))))))))........)))))))))... (-29.82 = -30.83 + 1.01)
| Location | 2,133,411 – 2,133,512 |
|---|---|
| Length | 101 |
| Sequences | 6 |
| Columns | 104 |
| Reading direction | forward |
| Mean pairwise identity | 76.89 |
| Mean single sequence MFE | -31.52 |
| Consensus MFE | -22.99 |
| Energy contribution | -23.13 |
| Covariance contribution | 0.14 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.55 |
| Structure conservation index | 0.73 |
| SVM decision value | 1.12 |
| SVM RNA-class probability | 0.919020 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
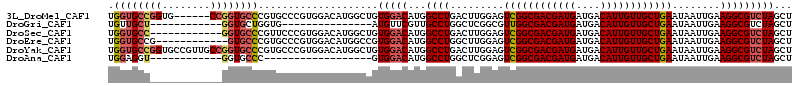
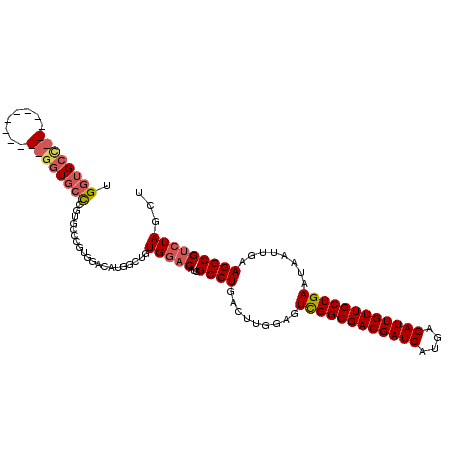
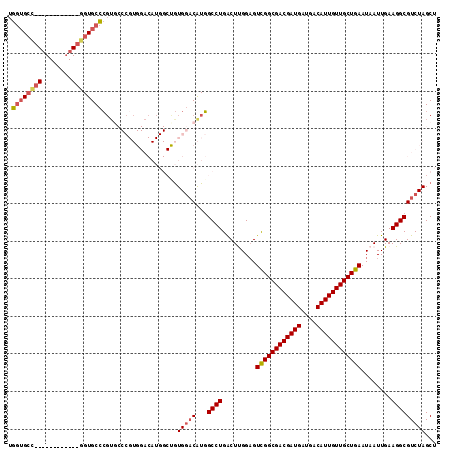
>3L_DroMel_CAF1 2133411 101 + 23771897 GGCUGUGGACAUGGCCUGACUUGGAGUCGGCGACGAUGAUGACAUUGUUGCUGAAUAAUUGAAGGCGUCUAGCUGGGACUGC---UGAGAUUGCUGGGAUUGCU ((((((....))))))...((..(..((((((((((((....)))))))))))).(((((...(((((((.....)))).))---)..))))))..))...... ( -35.60) >DroVir_CAF1 205185 87 + 1 --------UC---GCCUGGCUUGGCGUUGGCGACGAUGAUGACAUUGUUGCUGAAUAAUUGAAGGCGUCUAGCUGGGAUUGUGACUGAGAGUGU---G---ACU --------..---.((..(((.((((((((((((((((....))))))))))...........)))))).)))..))...((.((.......))---.---)). ( -30.50) >DroGri_CAF1 180938 94 + 1 ----AUGUUCGUUGCCUGGCUCGGCGUUGGCGACGAUGAUGACAUUGUUGCUGAAUAAUUGAAGGCGUCUAGCUGCGACUGCGACUGUAACUGC---A---ACU ----......(((((..((((.((((((((((((((((....))))))))))...........)))))).)))))))))((((........)))---)---... ( -30.40) >DroYak_CAF1 183243 83 + 1 GGCUGUGGACAUGGCCUGACUUGGAGUCGGCGACGAUGAUGACAUUGUUGCUGAAUAAUUGAAGGCGUCUAGCUGGGA---------------------CUGCU ((((((....))))))...(((.((.((((((((((((....))))))))))))....)).)))((((((.....)))---------------------).)). ( -31.20) >DroMoj_CAF1 183937 90 + 1 --------UCGUGGCCUGGCUCGGCGUUGGCGACGAUGAUGACAUUGUUGCUGAAUAAUUGAAGGCGUCUAGCUGCGACUGCGAGUGCGAGUGC---G---ACU --------..((.((((.((.(.(((((((((((((((....)))))))))............(((.....))).)))).))..).)).)).))---.---)). ( -30.90) >DroAna_CAF1 184635 97 + 1 ----GUGGACAUGGCCUGGCUCGGAGUCGGCGACGAUGAUGACAUUGUUGCUGAAUAAUUGAAGGCGUCUAGCUGGGACUGU---UGAGAUUGCUGCGACUGCU ----(..(.(..(((....(((((((((((((((((((....))))))))))...........(((.....)))..)))).)---))))...)))..).)..). ( -30.50) >consensus ____GUGGACAUGGCCUGGCUCGGAGUCGGCGACGAUGAUGACAUUGUUGCUGAAUAAUUGAAGGCGUCUAGCUGGGACUGC___UGAGAGUGC___G___ACU .............((((.........((((((((((((....))))))))))))........))))((((.....))))......................... (-22.99 = -23.13 + 0.14)

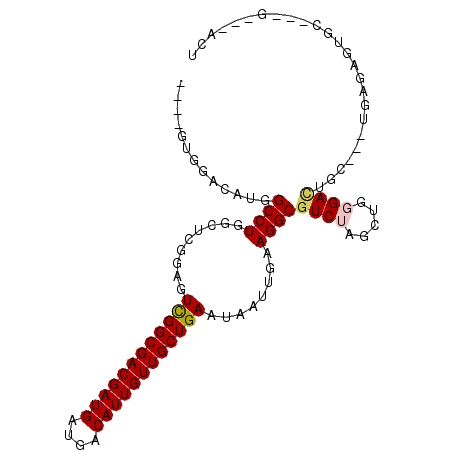
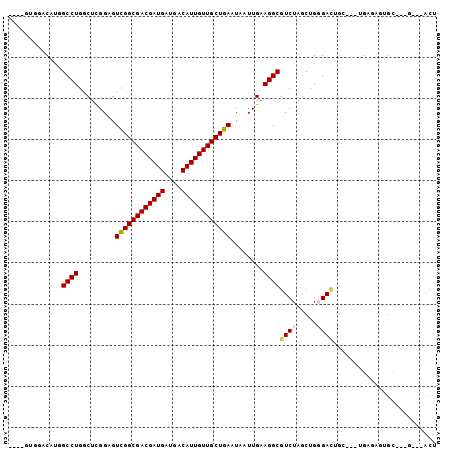
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:49:07 2006