
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 19,142,566 – 19,142,766 |
| Length | 200 |
| Max. P | 0.999822 |

| Location | 19,142,566 – 19,142,686 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 98.67 |
| Mean single sequence MFE | -31.66 |
| Consensus MFE | -31.58 |
| Energy contribution | -31.42 |
| Covariance contribution | -0.16 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.24 |
| Structure conservation index | 1.00 |
| SVM decision value | 3.39 |
| SVM RNA-class probability | 0.999132 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 19142566 120 + 23771897 AAAAGGAUUGCGCUCAGGCAGUGAUUUUCAAUUUAAAAAAUUGCACGCAAAAGCGCCACAAAUCAAAGGAAUUGACUUUGCCAGCAGAGGCGGAAAAAAGGGGGAGCCCUCUCCACAAUA .....(((((((((...((.(..(((((........)))))..)..))...)))))....))))...(((......((((((......))))))....((((....)))).)))...... ( -30.90) >DroSec_CAF1 5489 120 + 1 AAAAGGAUUGCGCUCAGGCAGUGAUUUUCAAUUUAAAAAAUUGCACGCAAAAGCGCCACAAAUCAAAGGAAUUGACUUUGCCAGCAGAGGCGGAAAAAAGGGGGACCCCUCUCCACAAUA .....(((((((((...((.(..(((((........)))))..)..))...)))))....))))...(((......((((((......))))))....((((....)))).)))...... ( -30.90) >DroSim_CAF1 5060 120 + 1 AAAAGGAUUGCGCUCAGGCAGUGAUUUUCAAUUUAAAAAAUUGCACGCAAAAGCGCCACAAAUCAAAGGAAUUGACUUUGCCAGCAGAGGCGGAAAAAAGGGGGACCCCUCUCCACAAUA .....(((((((((...((.(..(((((........)))))..)..))...)))))....))))...(((......((((((......))))))....((((....)))).)))...... ( -30.90) >DroEre_CAF1 5920 120 + 1 AAAAGGAUUGCGCUCAGGCAGUGAUUUUCAAUUUAAAAAAUUGCACGCAAAAGCGCCACAAAUCAAAGGAAUUGACUUUGCCAGCAGAGGCGGAAAAAAGGGGGACCCCUCUCCACAAUA .....(((((((((...((.(..(((((........)))))..)..))...)))))....))))...(((......((((((......))))))....((((....)))).)))...... ( -30.90) >DroYak_CAF1 5835 120 + 1 AAAAGGAUUGCGCUGAGGCAGUGAUUUUCAAUUUAAAAAAUUGCACGCAAAAGCGCCACAAAUCAAAGGAAUUGACUUUGCCAGCAGAGGCGGAAAAAAGGGGGACCCCCCUCCAUAAUA .....(((((((((...((.(..(((((........)))))..)..))...)))))....))))...(((......((((((......)))))).....((((....)))))))...... ( -34.70) >consensus AAAAGGAUUGCGCUCAGGCAGUGAUUUUCAAUUUAAAAAAUUGCACGCAAAAGCGCCACAAAUCAAAGGAAUUGACUUUGCCAGCAGAGGCGGAAAAAAGGGGGACCCCUCUCCACAAUA .....(((((((((...((.(..(((((........)))))..)..))...)))))....))))...(((......((((((......)))))).....((((....)))))))...... (-31.58 = -31.42 + -0.16)



| Location | 19,142,566 – 19,142,686 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 98.67 |
| Mean single sequence MFE | -35.90 |
| Consensus MFE | -35.36 |
| Energy contribution | -35.40 |
| Covariance contribution | 0.04 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.79 |
| Structure conservation index | 0.98 |
| SVM decision value | 4.17 |
| SVM RNA-class probability | 0.999822 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 19142566 120 - 23771897 UAUUGUGGAGAGGGCUCCCCCUUUUUUCCGCCUCUGCUGGCAAAGUCAAUUCCUUUGAUUUGUGGCGCUUUUGCGUGCAAUUUUUUAAAUUGAAAAUCACUGCCUGAGCGCAAUCCUUUU ......((((((((....)))))...)))(((......))).(((((((.....)))))))...((((((..(((((..((((((......))))))))).))..))))))......... ( -34.50) >DroSec_CAF1 5489 120 - 1 UAUUGUGGAGAGGGGUCCCCCUUUUUUCCGCCUCUGCUGGCAAAGUCAAUUCCUUUGAUUUGUGGCGCUUUUGCGUGCAAUUUUUUAAAUUGAAAAUCACUGCCUGAGCGCAAUCCUUUU ......((((((((....)))))...)))(((......))).(((((((.....)))))))...((((((..(((((..((((((......))))))))).))..))))))......... ( -36.70) >DroSim_CAF1 5060 120 - 1 UAUUGUGGAGAGGGGUCCCCCUUUUUUCCGCCUCUGCUGGCAAAGUCAAUUCCUUUGAUUUGUGGCGCUUUUGCGUGCAAUUUUUUAAAUUGAAAAUCACUGCCUGAGCGCAAUCCUUUU ......((((((((....)))))...)))(((......))).(((((((.....)))))))...((((((..(((((..((((((......))))))))).))..))))))......... ( -36.70) >DroEre_CAF1 5920 120 - 1 UAUUGUGGAGAGGGGUCCCCCUUUUUUCCGCCUCUGCUGGCAAAGUCAAUUCCUUUGAUUUGUGGCGCUUUUGCGUGCAAUUUUUUAAAUUGAAAAUCACUGCCUGAGCGCAAUCCUUUU ......((((((((....)))))...)))(((......))).(((((((.....)))))))...((((((..(((((..((((((......))))))))).))..))))))......... ( -36.70) >DroYak_CAF1 5835 120 - 1 UAUUAUGGAGGGGGGUCCCCCUUUUUUCCGCCUCUGCUGGCAAAGUCAAUUCCUUUGAUUUGUGGCGCUUUUGCGUGCAAUUUUUUAAAUUGAAAAUCACUGCCUCAGCGCAAUCCUUUU ......(((((((....))))........((..(((..(((((((((((.....)))))))(((((((....)))).(((((.....))))).....))))))).))).))..))).... ( -34.90) >consensus UAUUGUGGAGAGGGGUCCCCCUUUUUUCCGCCUCUGCUGGCAAAGUCAAUUCCUUUGAUUUGUGGCGCUUUUGCGUGCAAUUUUUUAAAUUGAAAAUCACUGCCUGAGCGCAAUCCUUUU ......((((((((....)))))...)))(((......))).(((((((.....)))))))...((((((..(((((..((((((......))))))))).))..))))))......... (-35.36 = -35.40 + 0.04)

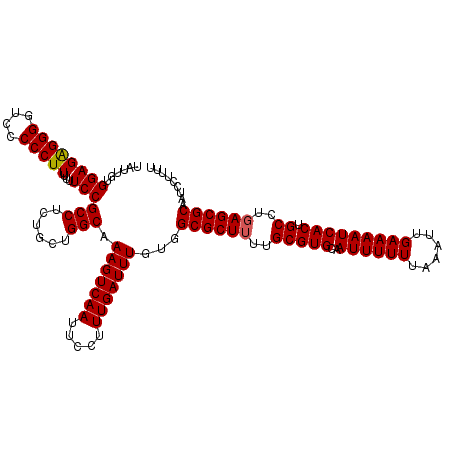

| Location | 19,142,606 – 19,142,726 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 97.75 |
| Mean single sequence MFE | -25.22 |
| Consensus MFE | -24.68 |
| Energy contribution | -24.20 |
| Covariance contribution | -0.48 |
| Combinations/Pair | 1.11 |
| Mean z-score | -1.39 |
| Structure conservation index | 0.98 |
| SVM decision value | 1.12 |
| SVM RNA-class probability | 0.917541 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 19142606 120 + 23771897 UUGCACGCAAAAGCGCCACAAAUCAAAGGAAUUGACUUUGCCAGCAGAGGCGGAAAAAAGGGGGAGCCCUCUCCACAAUAAAUUUCAAUCUAAAAACUUAACUAAAGGCACCAAGAGUAG .(((..((....))(((..........((((((..(((((....)))))(.(((....((((....)))).))).)....))))))....(((....)))......))).......))). ( -23.40) >DroSec_CAF1 5529 120 + 1 UUGCACGCAAAAGCGCCACAAAUCAAAGGAAUUGACUUUGCCAGCAGAGGCGGAAAAAAGGGGGACCCCUCUCCACAAUAAAUUUCAAUCUAAAAACUUAACUAAAGGCACCAAGAGCAC .(((..((....))(((..........((((((..(((((....)))))(.(((....((((....)))).))).)....))))))....(((....)))......))).......))). ( -25.10) >DroSim_CAF1 5100 120 + 1 UUGCACGCAAAAGCGCCACAAAUCAAAGGAAUUGACUUUGCCAGCAGAGGCGGAAAAAAGGGGGACCCCUCUCCACAAUAAAUUUCAAUCUAAAAACUUAACUAAAGGCACCAAGAGCAG .(((..((....))(((..........((((((..(((((....)))))(.(((....((((....)))).))).)....))))))....(((....)))......))).......))). ( -25.40) >DroEre_CAF1 5960 120 + 1 UUGCACGCAAAAGCGCCACAAAUCAAAGGAAUUGACUUUGCCAGCAGAGGCGGAAAAAAGGGGGACCCCUCUCCACAAUAAAUUUCAAUCUAAAAACUUAACUAAAGGCACCAAGAGCAG .(((..((....))(((..........((((((..(((((....)))))(.(((....((((....)))).))).)....))))))....(((....)))......))).......))). ( -25.40) >DroYak_CAF1 5875 120 + 1 UUGCACGCAAAAGCGCCACAAAUCAAAGGAAUUGACUUUGCCAGCAGAGGCGGAAAAAAGGGGGACCCCCCUCCAUAAUAAAUUUCAAUCUAAAAACUUAACUAAAGGCACCAAGAGCGA ((((..((....))(((..........((((((...((((((......))))))....(((((....)))))........))))))....(((....)))......))).......)))) ( -26.80) >consensus UUGCACGCAAAAGCGCCACAAAUCAAAGGAAUUGACUUUGCCAGCAGAGGCGGAAAAAAGGGGGACCCCUCUCCACAAUAAAUUUCAAUCUAAAAACUUAACUAAAGGCACCAAGAGCAG .(((..((....))(((..........((((((...((((((......))))))....(((((....)))))........))))))....(((....)))......))).......))). (-24.68 = -24.20 + -0.48)



| Location | 19,142,606 – 19,142,726 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 97.75 |
| Mean single sequence MFE | -35.44 |
| Consensus MFE | -33.48 |
| Energy contribution | -33.36 |
| Covariance contribution | -0.12 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.26 |
| Structure conservation index | 0.94 |
| SVM decision value | 3.28 |
| SVM RNA-class probability | 0.998924 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 19142606 120 - 23771897 CUACUCUUGGUGCCUUUAGUUAAGUUUUUAGAUUGAAAUUUAUUGUGGAGAGGGCUCCCCCUUUUUUCCGCCUCUGCUGGCAAAGUCAAUUCCUUUGAUUUGUGGCGCUUUUGCGUGCAA .(((.(..((((((..((((((((((((......))))))))..((((((((((....)))))...)))))....))))(((((.((((.....)))))))))))))))...).)))... ( -31.80) >DroSec_CAF1 5529 120 - 1 GUGCUCUUGGUGCCUUUAGUUAAGUUUUUAGAUUGAAAUUUAUUGUGGAGAGGGGUCCCCCUUUUUUCCGCCUCUGCUGGCAAAGUCAAUUCCUUUGAUUUGUGGCGCUUUUGCGUGCAA (..(.(..((((((..((((((((((((......))))))))..((((((((((....)))))...)))))....))))(((((.((((.....)))))))))))))))...).)..).. ( -37.30) >DroSim_CAF1 5100 120 - 1 CUGCUCUUGGUGCCUUUAGUUAAGUUUUUAGAUUGAAAUUUAUUGUGGAGAGGGGUCCCCCUUUUUUCCGCCUCUGCUGGCAAAGUCAAUUCCUUUGAUUUGUGGCGCUUUUGCGUGCAA .(((......((((..(((.((((((((......))))))))..((((((((((....)))))...)))))..)))..))))(((((((.....)))))))...((((....))))))). ( -35.80) >DroEre_CAF1 5960 120 - 1 CUGCUCUUGGUGCCUUUAGUUAAGUUUUUAGAUUGAAAUUUAUUGUGGAGAGGGGUCCCCCUUUUUUCCGCCUCUGCUGGCAAAGUCAAUUCCUUUGAUUUGUGGCGCUUUUGCGUGCAA .(((......((((..(((.((((((((......))))))))..((((((((((....)))))...)))))..)))..))))(((((((.....)))))))...((((....))))))). ( -35.80) >DroYak_CAF1 5875 120 - 1 UCGCUCUUGGUGCCUUUAGUUAAGUUUUUAGAUUGAAAUUUAUUAUGGAGGGGGGUCCCCCUUUUUUCCGCCUCUGCUGGCAAAGUCAAUUCCUUUGAUUUGUGGCGCUUUUGCGUGCAA .(((....((((((......((((((((......))))))))....(((((((....))))))).....(((......))).(((((((.....)))))))..))))))...)))..... ( -36.50) >consensus CUGCUCUUGGUGCCUUUAGUUAAGUUUUUAGAUUGAAAUUUAUUGUGGAGAGGGGUCCCCCUUUUUUCCGCCUCUGCUGGCAAAGUCAAUUCCUUUGAUUUGUGGCGCUUUUGCGUGCAA .(((....((((((......((((((((......))))))))....((((((((....)))))...)))(((......))).(((((((.....)))))))..))))))...)))..... (-33.48 = -33.36 + -0.12)


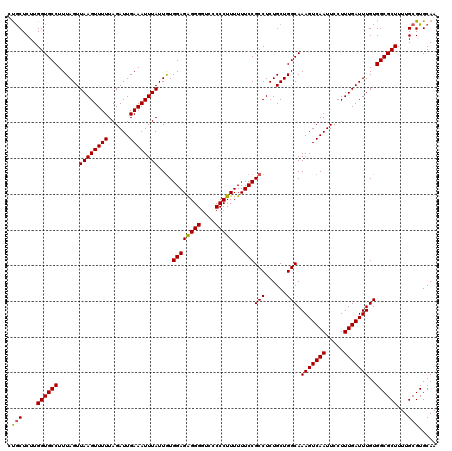
| Location | 19,142,646 – 19,142,766 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 95.07 |
| Mean single sequence MFE | -25.74 |
| Consensus MFE | -22.36 |
| Energy contribution | -22.60 |
| Covariance contribution | 0.24 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.13 |
| Structure conservation index | 0.87 |
| SVM decision value | 2.43 |
| SVM RNA-class probability | 0.993896 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 19142646 120 + 23771897 CCAGCAGAGGCGGAAAAAAGGGGGAGCCCUCUCCACAAUAAAUUUCAAUCUAAAAACUUAACUAAAGGCACCAAGAGUAGAGCUUCUGUUCUGAACUGAGUAAAAAAAAAAUGGAAAAUU ((((((((((((((....((((....)))).)))..........((..(((.....(((.....)))......)))...))))))))))((......))............)))...... ( -24.10) >DroSec_CAF1 5569 120 + 1 CCAGCAGAGGCGGAAAAAAGGGGGACCCCUCUCCACAAUAAAUUUCAAUCUAAAAACUUAACUAAAGGCACCAAGAGCACAGCUUCUGUUCUCAACUGAGUGAAAAAAAAAUGGAAAAUA ((((((((((((((....((((....)))).))).................................((.......))...)))))))).(((....)))...........)))...... ( -26.00) >DroSim_CAF1 5140 119 + 1 CCAGCAGAGGCGGAAAAAAGGGGGACCCCUCUCCACAAUAAAUUUCAAUCUAAAAACUUAACUAAAGGCACCAAGAGCAGAGCUUCUGUUCUGAACUGAGUGAAAA-AAAAUGGAAAAUA (((((....))(((....((((....)))).)))..................................(((..((..((((((....))))))..))..)))....-....)))...... ( -25.00) >DroEre_CAF1 6000 118 + 1 CCAGCAGAGGCGGAAAAAAGGGGGACCCCUCUCCACAAUAAAUUUCAAUCUAAAAACUUAACUAAAGGCACCAAGAGCAGAGCUUCUGUUCUGAGCUGAGUGAAAA--AAAUGGAAAAUA .((((.(..(((((....((((....)))).)))......................(((.....)))))..).(((((((.....)))))))..))))........--............ ( -27.20) >DroYak_CAF1 5915 118 + 1 CCAGCAGAGGCGGAAAAAAGGGGGACCCCCCUCCAUAAUAAAUUUCAAUCUAAAAACUUAACUAAAGGCACCAAGAGCGAAGCUUCUGUUCUGAACUGAGUGAAAA--AAAAGGAAAAUA ((.(((((((((((.....((((....))))))).........(((..(((.....(((.....)))......)))..)))))))))))((......)).......--....))...... ( -26.40) >consensus CCAGCAGAGGCGGAAAAAAGGGGGACCCCUCUCCACAAUAAAUUUCAAUCUAAAAACUUAACUAAAGGCACCAAGAGCAGAGCUUCUGUUCUGAACUGAGUGAAAA_AAAAUGGAAAAUA ((((((((((((((....((((....)))).))).................................((.......))...))))))))((......))............)))...... (-22.36 = -22.60 + 0.24)


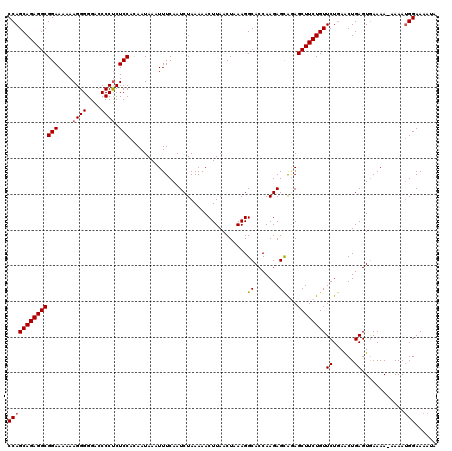
| Location | 19,142,646 – 19,142,766 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 95.07 |
| Mean single sequence MFE | -26.32 |
| Consensus MFE | -24.32 |
| Energy contribution | -24.04 |
| Covariance contribution | -0.28 |
| Combinations/Pair | 1.11 |
| Mean z-score | -1.07 |
| Structure conservation index | 0.92 |
| SVM decision value | -0.05 |
| SVM RNA-class probability | 0.506902 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 19142646 120 - 23771897 AAUUUUCCAUUUUUUUUUUACUCAGUUCAGAACAGAAGCUCUACUCUUGGUGCCUUUAGUUAAGUUUUUAGAUUGAAAUUUAUUGUGGAGAGGGCUCCCCCUUUUUUCCGCCUCUGCUGG ......................(((..((((..(((((((..(((...(....)...)))..)))))))...............((((((((((....)))))...))))).))))))). ( -23.10) >DroSec_CAF1 5569 120 - 1 UAUUUUCCAUUUUUUUUUCACUCAGUUGAGAACAGAAGCUGUGCUCUUGGUGCCUUUAGUUAAGUUUUUAGAUUGAAAUUUAUUGUGGAGAGGGGUCCCCCUUUUUUCCGCCUCUGCUGG ......................((((.((((((.((((..(..(.....)..))))).)))(((((((......)))))))...((((((((((....)))))...)))))))).)))). ( -28.40) >DroSim_CAF1 5140 119 - 1 UAUUUUCCAUUUU-UUUUCACUCAGUUCAGAACAGAAGCUCUGCUCUUGGUGCCUUUAGUUAAGUUUUUAGAUUGAAAUUUAUUGUGGAGAGGGGUCCCCCUUUUUUCCGCCUCUGCUGG .............-....((((.((..((((........))))..)).))))((..(((.((((((((......))))))))..((((((((((....)))))...)))))..)))..)) ( -26.80) >DroEre_CAF1 6000 118 - 1 UAUUUUCCAUUU--UUUUCACUCAGCUCAGAACAGAAGCUCUGCUCUUGGUGCCUUUAGUUAAGUUUUUAGAUUGAAAUUUAUUGUGGAGAGGGGUCCCCCUUUUUUCCGCCUCUGCUGG ............--........((((((((...(((((((..(((...(....)...)))..)))))))...))))........((((((((((....)))))...)))))....)))). ( -27.60) >DroYak_CAF1 5915 118 - 1 UAUUUUCCUUUU--UUUUCACUCAGUUCAGAACAGAAGCUUCGCUCUUGGUGCCUUUAGUUAAGUUUUUAGAUUGAAAUUUAUUAUGGAGGGGGGUCCCCCUUUUUUCCGCCUCUGCUGG ............--........(((..((((..(((.((...))))).((((((((((..((((((((......))))))))...))))))(((....))).......))))))))))). ( -25.70) >consensus UAUUUUCCAUUUU_UUUUCACUCAGUUCAGAACAGAAGCUCUGCUCUUGGUGCCUUUAGUUAAGUUUUUAGAUUGAAAUUUAUUGUGGAGAGGGGUCCCCCUUUUUUCCGCCUCUGCUGG ......................((((..((...(((((((..(((...(....)...)))..)))))))...............((((((((((....)))))...)))))))..)))). (-24.32 = -24.04 + -0.28)


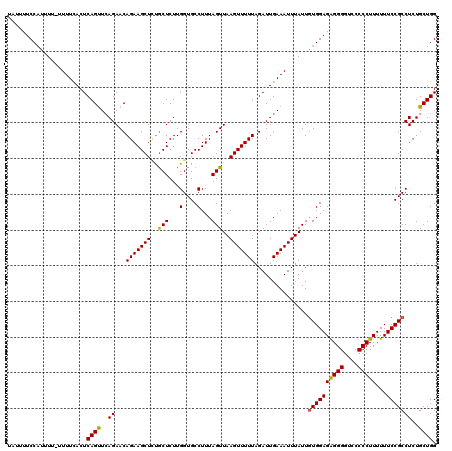
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:37:15 2006