
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 19,064,989 – 19,065,093 |
| Length | 104 |
| Max. P | 0.980020 |

| Location | 19,064,989 – 19,065,093 |
|---|---|
| Length | 104 |
| Sequences | 6 |
| Columns | 104 |
| Reading direction | forward |
| Mean pairwise identity | 77.50 |
| Mean single sequence MFE | -35.95 |
| Consensus MFE | -17.17 |
| Energy contribution | -18.40 |
| Covariance contribution | 1.23 |
| Combinations/Pair | 1.25 |
| Mean z-score | -2.02 |
| Structure conservation index | 0.48 |
| SVM decision value | 0.24 |
| SVM RNA-class probability | 0.647045 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
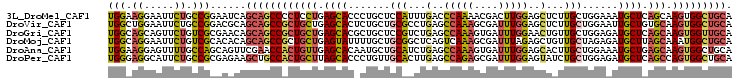
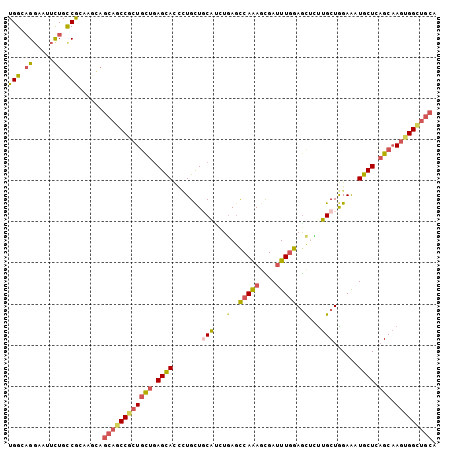
>3L_DroMel_CAF1 19064989 104 + 23771897 UGCAGCCACUUGCUGAGCAUUUCCAGCAAGAGCUCCAAGUCGUUUUGGGUCAAAUGGAGCAGGGUGCUCAGGAGGGGCUGCUGAUUCCGGCAGAAUUCCUUCCA .(((.((.(((((((........))))))).((((((.................)))))).)).)))...(((((((((((((....))))))...))))))). ( -45.33) >DroVir_CAF1 32195 104 + 1 UGCAGCCACUUGCACAGCAAUUCCAGCAAGAGCUCCAAAUCGCUUUGGCUCAGGCGCAGCAGAGUGCUCAGCAGCGGCUGCUGCGUCCGGCAGAAUUCCAGCCA ((.((((((((((............)))))(((........))).)))))))((((((((((..((((....)))).)))))))))).(((.........))). ( -39.90) >DroGri_CAF1 39693 104 + 1 UGCAACCACUUGCUGAGCAUCUCCAGCAACAGUUCCAAAUCACUUUGGCUCAGACGGAGCAGCGUGCUCAGCAGCGGCUGCUGUUCGCGACAGAACUGCUGCCA .........((((((((((((((((((....)))(((((....))))).......))))....))))))))))).(((.((.((((......)))).)).))). ( -37.80) >DroMoj_CAF1 4518 104 + 1 UGCAGCCAUUUGCUAAGCAUCUCUAGCAACAGCUCUAAAUCGCUUUGACUGAGCCGCAGCAAAAUACUCAGCAGCGGCUGCUGUGUGCGACAGAAUUCCUGCCA .((((..((((.....((((...((((..(((.((...........)))))((((((.((..........)).)))))))))).))))....))))..)))).. ( -29.30) >DroAna_CAF1 5393 104 + 1 UGCAGCCACUUGCUCAGCAUUUCCAGCAAGUGCUCCAAAUCACUUUGGCUCAGAUGCAGCAUUGUGCUCAACAGUGGUUCGAACUGCUGGCAAAACUCCUUCCA .........((((.(((((.(((..(((..((..(((((....)))))..))..)))..((((((.....))))))....))).)))))))))........... ( -28.00) >DroPer_CAF1 15354 104 + 1 UGCAGCCACUGGCUGAGCAUCUCCAGCAGAUACUCCAAAUCGCUCUGGCUCAAGUGCAACAGGGUGCUAAGCAGUGGCAGCUUCUCGCGGCAGAAUGCCUCCCA .((.(((((((.((.(((((((.(((..(((.......)))...)))((......))....))))))).))))))))).)).......(((.....)))..... ( -35.40) >consensus UGCAGCCACUUGCUGAGCAUCUCCAGCAAGAGCUCCAAAUCGCUUUGGCUCAGACGCAGCAGAGUGCUCAGCAGCGGCUGCUGCUUCCGGCAGAAUUCCUGCCA .((((((.((.(((.(((((....(((....)))(((((....)))))...............))))).))))).))))))....................... (-17.17 = -18.40 + 1.23)



| Location | 19,064,989 – 19,065,093 |
|---|---|
| Length | 104 |
| Sequences | 6 |
| Columns | 104 |
| Reading direction | reverse |
| Mean pairwise identity | 77.50 |
| Mean single sequence MFE | -41.55 |
| Consensus MFE | -25.42 |
| Energy contribution | -26.07 |
| Covariance contribution | 0.65 |
| Combinations/Pair | 1.37 |
| Mean z-score | -2.55 |
| Structure conservation index | 0.61 |
| SVM decision value | 1.85 |
| SVM RNA-class probability | 0.980020 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 19064989 104 - 23771897 UGGAAGGAAUUCUGCCGGAAUCAGCAGCCCCUCCUGAGCACCCUGCUCCAUUUGACCCAAAACGACUUGGAGCUCUUGCUGGAAAUGCUCAGCAAGUGGCUGCA .((.((.....)).)).......((((((.((.(((((((....((((((.(((........)))..))))))(((....)))..)))))))..)).)))))). ( -36.30) >DroVir_CAF1 32195 104 - 1 UGGCUGGAAUUCUGCCGGACGCAGCAGCCGCUGCUGAGCACUCUGCUGCGCCUGAGCCAAAGCGAUUUGGAGCUCUUGCUGGAAUUGCUGUGCAAGUGGCUGCA .(((.((....)))))((.((((((((..(((....)))...))))))))))..(((((.((((((((.(.((....))).)))))))).......)))))... ( -41.90) >DroGri_CAF1 39693 104 - 1 UGGCAGCAGUUCUGUCGCGAACAGCAGCCGCUGCUGAGCACGCUGCUCCGUCUGAGCCAAAGUGAUUUGGAACUGUUGCUGGAGAUGCUCAGCAAGUGGUUGCA .(((((.....))))).......(((((((((((((((((.....(((((.(.(..(((((....)))))..)....).))))).)))))))).))))))))). ( -47.20) >DroMoj_CAF1 4518 104 - 1 UGGCAGGAAUUCUGUCGCACACAGCAGCCGCUGCUGAGUAUUUUGCUGCGGCUCAGUCAAAGCGAUUUAGAGCUGUUGCUAGAGAUGCUUAGCAAAUGGCUGCA .(((((.....))))).......(((((((.(((((((((((((((.(((((((((((.....))))..))))))).))..)))))))))))))..))))))). ( -49.00) >DroAna_CAF1 5393 104 - 1 UGGAAGGAGUUUUGCCAGCAGUUCGAACCACUGUUGAGCACAAUGCUGCAUCUGAGCCAAAGUGAUUUGGAGCACUUGCUGGAAAUGCUGAGCAAGUGGCUGCA .(((.(.(((..((((((((((.......))))))).)))....))).).)))...(((((....)))))((((((((((.(......).))))))).)))... ( -34.70) >DroPer_CAF1 15354 104 - 1 UGGGAGGCAUUCUGCCGCGAGAAGCUGCCACUGCUUAGCACCCUGUUGCACUUGAGCCAGAGCGAUUUGGAGUAUCUGCUGGAGAUGCUCAGCCAGUGGCUGCA .....(((.....))).......((.(((((((((.(((((((....(((..((..(((((....)))))..))..))).)).).)))).)).))))))).)). ( -40.20) >consensus UGGCAGGAAUUCUGCCGCAAGCAGCAGCCGCUGCUGAGCACCCUGCUGCAUCUGAGCCAAAGCGAUUUGGAGCUCUUGCUGGAAAUGCUCAGCAAGUGGCUGCA (((.((.....)).)))......((((((((((((.((((.......(((...(..(((((....)))))..)...)))......)))).))).))))))))). (-25.42 = -26.07 + 0.65)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:36:12 2006