
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 18,221,947 – 18,222,058 |
| Length | 111 |
| Max. P | 0.876646 |

| Location | 18,221,947 – 18,222,058 |
|---|---|
| Length | 111 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 78.18 |
| Mean single sequence MFE | -30.10 |
| Consensus MFE | -18.97 |
| Energy contribution | -18.83 |
| Covariance contribution | -0.14 |
| Combinations/Pair | 1.19 |
| Mean z-score | -1.41 |
| Structure conservation index | 0.63 |
| SVM decision value | 0.18 |
| SVM RNA-class probability | 0.622508 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 18221947 111 + 23771897 GA----GCAA---AGUGAGCAUUUUGCGCGAAAUUGCAUUGCAGAGUUGAACAAUGAUUUAUGUGGCAGCCGGCCGCACCUACAUUUAUGCCAC-CGAGUGGCCAUAAAUCACACCAAA- ..----((((---(((....))))))).(((..((((...))))..))).....(((((((((((((.....)))))............(((((-...))))).)))))))).......- ( -29.90) >DroVir_CAF1 26149 109 + 1 GGUAUCGAGUUACAGCAACGCUUUUCUGCA---ACGCGUUGCA-ACUUGGACGAUGAUUUAUGUG-----CGGCCGCACCUACAUUUAUGCCAU-AAAGUGGCCAUAAAUCACACCAGG- (((.(((((((...(((((((.........---..))))))))-))))))..(.(((((((((((-----(....))))..........(((((-...))))).))))))))))))...- ( -34.40) >DroPse_CAF1 6247 112 + 1 AGCCUCGAAU---GGCGAAAAUUUUGUGCA--AUUGCAA---AGCGUUGAACAAUGAUUUAUGUGGCAGCAGGCCGCACCUACAUUUAUGCCAUGCAAGUGGCCAUAAAUCACACCAAAA .(((......---))).......((((.((--((.((..---.)))))).))))(((((((((((((.....)))))............(((((....))))).))))))))........ ( -29.60) >DroEre_CAF1 4499 111 + 1 GA----GCAA---AGUGAGCAUUUUGCGCGAAUGUGCACUGCAGAGUUGAACAAUGAUUUAUGUGGCAGCUGGCCGCACCUACAUUUAUGCCAC-CGACCGGCCAUAAAUCACACCAAA- ..----....---.(((.(((...(((((....))))).)))............(((((((((((((.....)))))............(((..-.....))).)))))))))))....- ( -27.70) >DroAna_CAF1 10087 111 + 1 GG----GAAA---AGUGAACAUUUUGCGCAAAAUUGCACUGCUGAGCUGAACAAUGAUUUAUGUGGCAGCAGGCCGCACCUACAUUUAUGCCAC-CGAGUGGCCAUAAAUCACACCAAA- ((----....---.((((......((((((....))).((((((.((((((......)))).))..))))))...)))......((((((((((-...)))).)))))))))).))...- ( -29.40) >DroPer_CAF1 6105 112 + 1 AGCCUCGAAU---GGCGAAAAUUUUGUGCA--AUUGCAA---AGCGUUGAACAAUGAUUUAUGUGGCAGCAGGCCGCACCUACAUUUAUGCCAUGCAAGUGGCCAUAAAUCACACCAAAA .(((......---))).......((((.((--((.((..---.)))))).))))(((((((((((((.....)))))............(((((....))))).))))))))........ ( -29.60) >consensus GG____GAAA___AGCGAACAUUUUGCGCA__AUUGCAAUGCAGAGUUGAACAAUGAUUUAUGUGGCAGCAGGCCGCACCUACAUUUAUGCCAC_CAAGUGGCCAUAAAUCACACCAAA_ ..............(((.........))).........................(((((((((((((.....)))))............(((((....))))).))))))))........ (-18.97 = -18.83 + -0.14)

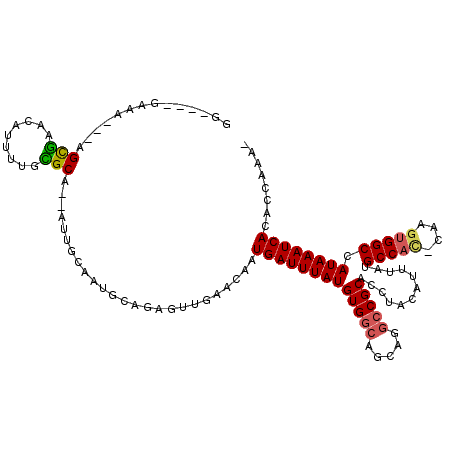

| Location | 18,221,947 – 18,222,058 |
|---|---|
| Length | 111 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 78.18 |
| Mean single sequence MFE | -33.68 |
| Consensus MFE | -21.94 |
| Energy contribution | -22.05 |
| Covariance contribution | 0.11 |
| Combinations/Pair | 1.13 |
| Mean z-score | -1.78 |
| Structure conservation index | 0.65 |
| SVM decision value | 0.90 |
| SVM RNA-class probability | 0.876646 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 18221947 111 - 23771897 -UUUGGUGUGAUUUAUGGCCACUCG-GUGGCAUAAAUGUAGGUGCGGCCGGCUGCCACAUAAAUCAUUGUUCAACUCUGCAAUGCAAUUUCGCGCAAAAUGCUCACU---UUGC----UC -.((((.(((((((((((.((..((-((.((((........)))).))))..)).).))))))))))...))))....((((.((......))((.....)).....---))))----.. ( -33.10) >DroVir_CAF1 26149 109 - 1 -CCUGGUGUGAUUUAUGGCCACUUU-AUGGCAUAAAUGUAGGUGCGGCCG-----CACAUAAAUCAUCGUCCAAGU-UGCAACGCGU---UGCAGAAAAGCGUUGCUGUAACUCGAUACC -..(((.((((((((((((((....-.)))).....(((.((.....)))-----))))))))))))...)))(((-((((..(((.---(((......))).))))))))))....... ( -32.40) >DroPse_CAF1 6247 112 - 1 UUUUGGUGUGAUUUAUGGCCACUUGCAUGGCAUAAAUGUAGGUGCGGCCUGCUGCCACAUAAAUCAUUGUUCAACGCU---UUGCAAU--UGCACAAAAUUUUCGCC---AUUCGAGGCU ..((((.((((((((((((.((((((((.......))))))))))(((.....))).))))))))))...)))).(((---(((.(((--.((...........)).---))))))))). ( -35.00) >DroEre_CAF1 4499 111 - 1 -UUUGGUGUGAUUUAUGGCCGGUCG-GUGGCAUAAAUGUAGGUGCGGCCAGCUGCCACAUAAAUCAUUGUUCAACUCUGCAGUGCACAUUCGCGCAAAAUGCUCACU---UUGC----UC -.((((.(((((((((((.((((.(-((.((((........)))).))).)))).).))))))))))...))))....(((.(((.(....).)))...))).....---....----.. ( -34.60) >DroAna_CAF1 10087 111 - 1 -UUUGGUGUGAUUUAUGGCCACUCG-GUGGCAUAAAUGUAGGUGCGGCCUGCUGCCACAUAAAUCAUUGUUCAGCUCAGCAGUGCAAUUUUGCGCAAAAUGUUCACU---UUUC----CC -.((((.(((((((((((((....)-))((((.....(((((.....))))))))).))))))))))...)))).......(((((....)))))............---....----.. ( -32.00) >DroPer_CAF1 6105 112 - 1 UUUUGGUGUGAUUUAUGGCCACUUGCAUGGCAUAAAUGUAGGUGCGGCCUGCUGCCACAUAAAUCAUUGUUCAACGCU---UUGCAAU--UGCACAAAAUUUUCGCC---AUUCGAGGCU ..((((.((((((((((((.((((((((.......))))))))))(((.....))).))))))))))...)))).(((---(((.(((--.((...........)).---))))))))). ( -35.00) >consensus _UUUGGUGUGAUUUAUGGCCACUCG_AUGGCAUAAAUGUAGGUGCGGCCUGCUGCCACAUAAAUCAUUGUUCAACUCUGCAAUGCAAU__UGCACAAAAUGUUCACU___AUUC____CC ..((((.((((((((((((((......))))..............(((.....))).))))))))))...)))).........((......))........................... (-21.94 = -22.05 + 0.11)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:29:51 2006