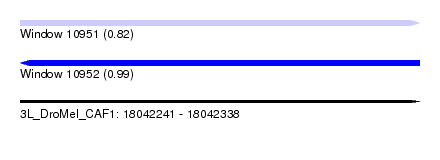

| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 18,042,241 – 18,042,338 |
| Length | 97 |
| Max. P | 0.991109 |

| Location | 18,042,241 – 18,042,338 |
|---|---|
| Length | 97 |
| Sequences | 5 |
| Columns | 97 |
| Reading direction | forward |
| Mean pairwise identity | 78.83 |
| Mean single sequence MFE | -22.18 |
| Consensus MFE | -13.84 |
| Energy contribution | -13.64 |
| Covariance contribution | -0.20 |
| Combinations/Pair | 1.21 |
| Mean z-score | -1.91 |
| Structure conservation index | 0.62 |
| SVM decision value | 0.68 |
| SVM RNA-class probability | 0.820765 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 18042241 97 + 23771897 GCGGCAAGUGGAAAACCAGUUAAUGCCAUAGGAAGACUAACUGGAAGUUUUCCACCAACUAAAGCUAAAGGAAUGCUGAAAAUAUAUUGUGCAAUAC (((.((((((((((((((((((...((...)).....)))))))...)))))))).......(((.........))).........))))))..... ( -22.40) >DroSec_CAF1 157035 97 + 1 GCGGCAAGUGGAAAACCAGUUAAUGGCAUAGGAAGACUAACUAGAAGUUUUCCACCAACUAAAGCUAAAAGAAUGCUGCAAAUAUUUUGUGCAAUGC (((((((((((....(((.....)))....((((((((.......)))))))).)).)))....(.....)..)))))).................. ( -22.00) >DroSim_CAF1 168820 97 + 1 GCGGCAAGUGGAAAACCAGUUAAUGGCAUAGGAAGACUAACUAGAAGUUUUCCACCAACUAAACCUAAAUGAAUGCUGAAAAUUUAUUGUGCAAUUC ((((..(((((....(((.....)))....((((((((.......)))))))).)).)))...))..(((((((.......)))))))..))..... ( -20.30) >DroEre_CAF1 157149 79 + 1 GCGGCAAGUGGAAAACCAGUUAAUGGCUUAGGAGGACCAACUGCAAGUUUUCCACCAGCUAAA-------AUAUA-----------UUUCGCAAUGC ..(((..(((((((((((((...(((((.....)).)))))))...)))))))))..)))...-------.....-----------........... ( -20.80) >DroYak_CAF1 164261 86 + 1 GCGGCAAGUGGAAAACCAGUUAAUGGCAUACGAAGAUCAACUGAAGGUUUUCCACCAGCCAAAGGUAGAAGAAUG-----------CUCCGCAAUGC (((((..((((((((((.((.....)).........((....)).))))))))))..)))...((.((.......-----------))))))..... ( -25.40) >consensus GCGGCAAGUGGAAAACCAGUUAAUGGCAUAGGAAGACUAACUGGAAGUUUUCCACCAACUAAAGCUAAAAGAAUGCUG_AAAU_U_UUGUGCAAUGC (((..((.(((....))).))..(((....((((((((.......)))))))).)))................................)))..... (-13.84 = -13.64 + -0.20)

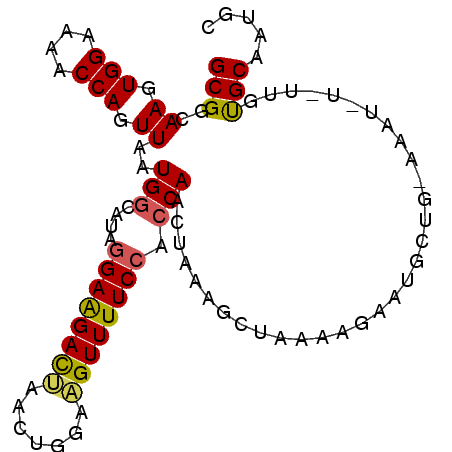
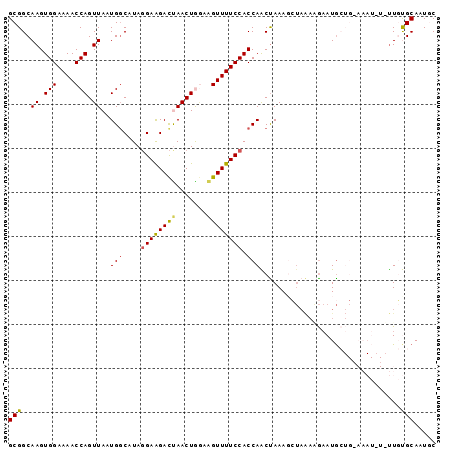
| Location | 18,042,241 – 18,042,338 |
|---|---|
| Length | 97 |
| Sequences | 5 |
| Columns | 97 |
| Reading direction | reverse |
| Mean pairwise identity | 78.83 |
| Mean single sequence MFE | -23.85 |
| Consensus MFE | -20.44 |
| Energy contribution | -19.56 |
| Covariance contribution | -0.88 |
| Combinations/Pair | 1.20 |
| Mean z-score | -2.39 |
| Structure conservation index | 0.86 |
| SVM decision value | 2.25 |
| SVM RNA-class probability | 0.991109 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 18042241 97 - 23771897 GUAUUGCACAAUAUAUUUUCAGCAUUCCUUUAGCUUUAGUUGGUGGAAAACUUCCAGUUAGUCUUCCUAUGGCAUUAACUGGUUUUCCACUUGCCGC .....((.............(((.........)))...((.(((((((((...(((((((((((......)).)))))))))))))))))).)).)) ( -24.20) >DroSec_CAF1 157035 97 - 1 GCAUUGCACAAAAUAUUUGCAGCAUUCUUUUAGCUUUAGUUGGUGGAAAACUUCUAGUUAGUCUUCCUAUGCCAUUAACUGGUUUUCCACUUGCCGC ((..((((.........))))))...............((.(((((((((((...(((((((...........)))))))))))))))))).))... ( -21.90) >DroSim_CAF1 168820 97 - 1 GAAUUGCACAAUAAAUUUUCAGCAUUCAUUUAGGUUUAGUUGGUGGAAAACUUCUAGUUAGUCUUCCUAUGCCAUUAACUGGUUUUCCACUUGCCGC ((((.((..............)))))).....(((......(((((((((((...(((((((...........)))))))))))))))))).))).. ( -22.34) >DroEre_CAF1 157149 79 - 1 GCAUUGCGAAA-----------UAUAU-------UUUAGCUGGUGGAAAACUUGCAGUUGGUCCUCCUAAGCCAUUAACUGGUUUUCCACUUGCCGC .....((((((-----------.....-------))).((.(((((((((((......((((........))))......))))))))))).))))) ( -22.70) >DroYak_CAF1 164261 86 - 1 GCAUUGCGGAG-----------CAUUCUUCUACCUUUGGCUGGUGGAAAACCUUCAGUUGAUCUUCGUAUGCCAUUAACUGGUUUUCCACUUGCCGC .....((((((-----------....)))).......(((.(((((((((((...(((((((...........)))))))))))))))))).))))) ( -28.10) >consensus GCAUUGCACAA_A_AUUU_CAGCAUUCUUUUAGCUUUAGUUGGUGGAAAACUUCCAGUUAGUCUUCCUAUGCCAUUAACUGGUUUUCCACUUGCCGC ......................................((.((((((((((...((((((((...........)))))))))))))))))).))... (-20.44 = -19.56 + -0.88)

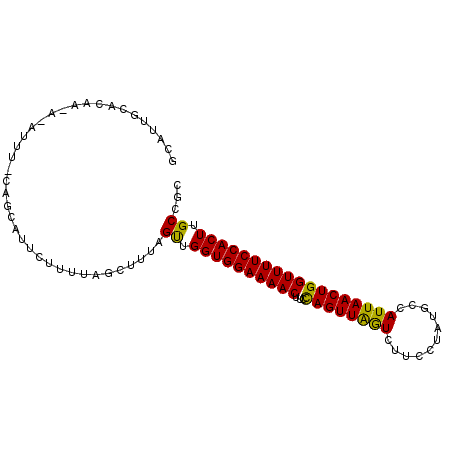
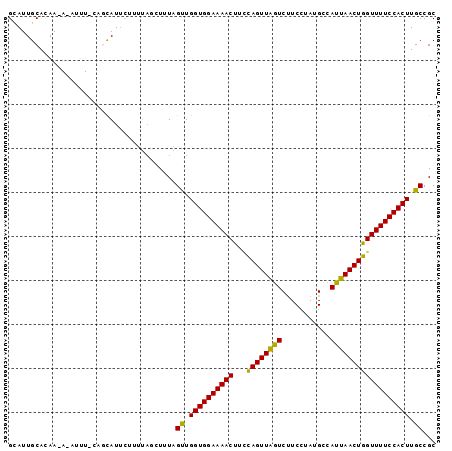
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:28:44 2006