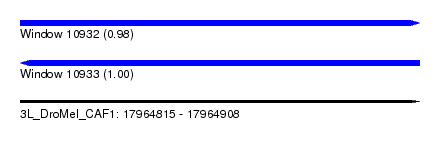

| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 17,964,815 – 17,964,908 |
| Length | 93 |
| Max. P | 0.999437 |

| Location | 17,964,815 – 17,964,908 |
|---|---|
| Length | 93 |
| Sequences | 5 |
| Columns | 95 |
| Reading direction | forward |
| Mean pairwise identity | 93.28 |
| Mean single sequence MFE | -41.46 |
| Consensus MFE | -33.66 |
| Energy contribution | -33.78 |
| Covariance contribution | 0.12 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.33 |
| Structure conservation index | 0.81 |
| SVM decision value | 1.94 |
| SVM RNA-class probability | 0.983300 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 17964815 93 + 23771897 GUCGACGUCGGCGCUCUCCGCUUUGCACUC--UCUCGCACUUCGAACAUGUGCUGCCAUCCACCCAGCGAGUGCGAGCAGGAGCGCGCGAUCCCA ...(..(((((((((((..((((.((((((--....((((.........)))).((..........)))))))))))).))))))).))))..). ( -37.10) >DroSec_CAF1 87552 93 + 1 GUCGACGUCGGCGCUCUCCGCUUCGCACUC--UCUCGGACUUCGAACAUGUGCUGCCAUCCACCCAGCGAGUGCGAGAAGGAGCGCGCGAUCCCA ...(..((((.(((.((((..(((((((((--..(((.....)))......((((.........)))))))))))))..)))).)))))))..). ( -37.60) >DroSim_CAF1 94488 93 + 1 GUCGACGUCGGCGCUCUCCGCUUCGCACUC--UCUCGCACGUCGAACAUGUGCUGCCAUCCACCCAGCGAGUGCGAGCAGGAGCGCGCGAUCCCA ...(..(((((((((((..(((.(((((((--....((((((.....)))))).((..........)))))))))))).))))))).))))..). ( -40.90) >DroEre_CAF1 89873 93 + 1 GUCGACGUCGGCGCUCUCCGGCUCGCACUC--UCUCGCACUUGGAACAUGUGCUGCCGUCCAGCCAGCGAGUGCGAGCAGGAGCGCGCGAUCCGA .(((..(((((((((((...((((((((((--.((.((...((((.((.....))...)))))).)).)))))))))).))))))).)))).))) ( -46.20) >DroYak_CAF1 93920 95 + 1 GUCGACGUCGGCGCUCUCCGCGUCGCACUCUCUCUCGCACUUGAAACAUGUGCUGCCAUCCAGCCAGCGAGUGCGAGCGGGAGCGCGCGAUCCCA ((((.(...(((((.....)))))((.(((((.((((((((((.....((.((((.....)))))).)))))))))).))))).))))))).... ( -45.50) >consensus GUCGACGUCGGCGCUCUCCGCUUCGCACUC__UCUCGCACUUCGAACAUGUGCUGCCAUCCACCCAGCGAGUGCGAGCAGGAGCGCGCGAUCCCA ......((((.(((.((((..(((((((((......((((.........)))).((..........)))))))))))..)))).))))))).... (-33.66 = -33.78 + 0.12)



| Location | 17,964,815 – 17,964,908 |
|---|---|
| Length | 93 |
| Sequences | 5 |
| Columns | 95 |
| Reading direction | reverse |
| Mean pairwise identity | 93.28 |
| Mean single sequence MFE | -45.74 |
| Consensus MFE | -41.34 |
| Energy contribution | -42.02 |
| Covariance contribution | 0.68 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.33 |
| Structure conservation index | 0.90 |
| SVM decision value | 3.60 |
| SVM RNA-class probability | 0.999437 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 17964815 93 - 23771897 UGGGAUCGCGCGCUCCUGCUCGCACUCGCUGGGUGGAUGGCAGCACAUGUUCGAAGUGCGAGA--GAGUGCAAAGCGGAGAGCGCCGACGUCGAC .((..(((.(((((((((((.((((((((((......)))).((((.........))))....--))))))..))))).)))))))))..))... ( -41.70) >DroSec_CAF1 87552 93 - 1 UGGGAUCGCGCGCUCCUUCUCGCACUCGCUGGGUGGAUGGCAGCACAUGUUCGAAGUCCGAGA--GAGUGCGAAGCGGAGAGCGCCGACGUCGAC .((..(((.((((((((.(((((((((.((...(((((..((((....))).)..))))))).--))))))).)).)).)))))))))..))... ( -39.60) >DroSim_CAF1 94488 93 - 1 UGGGAUCGCGCGCUCCUGCUCGCACUCGCUGGGUGGAUGGCAGCACAUGUUCGACGUGCGAGA--GAGUGCGAAGCGGAGAGCGCCGACGUCGAC .((..(((.((((((((((((((((((((((......)))).((((.........))))....--))))))).))))).)))))))))..))... ( -45.80) >DroEre_CAF1 89873 93 - 1 UCGGAUCGCGCGCUCCUGCUCGCACUCGCUGGCUGGACGGCAGCACAUGUUCCAAGUGCGAGA--GAGUGCGAGCCGGAGAGCGCCGACGUCGAC ((((.((((((.((((.((((((((((((((......)))).((((.((...)).))))....--)))))))))).)))).))).)))..)))). ( -51.50) >DroYak_CAF1 93920 95 - 1 UGGGAUCGCGCGCUCCCGCUCGCACUCGCUGGCUGGAUGGCAGCACAUGUUUCAAGUGCGAGAGAGAGUGCGACGCGGAGAGCGCCGACGUCGAC .((..(((.((((((((((((((((((((((.(.....).))))...(.((((......)))).))))))))).)))).)))))))))..))... ( -50.10) >consensus UGGGAUCGCGCGCUCCUGCUCGCACUCGCUGGGUGGAUGGCAGCACAUGUUCGAAGUGCGAGA__GAGUGCGAAGCGGAGAGCGCCGACGUCGAC (((..(((.((((((((((((((((((((((......)))).((((.........))))......)))))))).)))).)))))))))..))).. (-41.34 = -42.02 + 0.68)

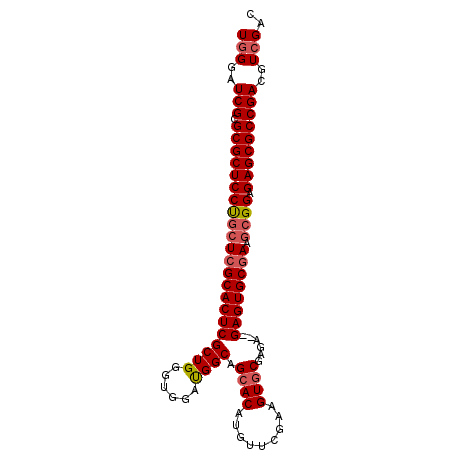

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:28:25 2006