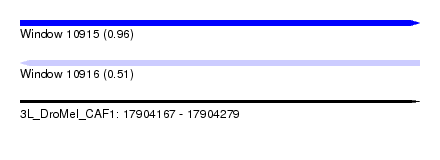

| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 17,904,167 – 17,904,279 |
| Length | 112 |
| Max. P | 0.955935 |

| Location | 17,904,167 – 17,904,279 |
|---|---|
| Length | 112 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 85.64 |
| Mean single sequence MFE | -34.40 |
| Consensus MFE | -28.00 |
| Energy contribution | -28.67 |
| Covariance contribution | 0.67 |
| Combinations/Pair | 1.16 |
| Mean z-score | -3.37 |
| Structure conservation index | 0.81 |
| SVM decision value | 1.46 |
| SVM RNA-class probability | 0.955935 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 17904167 112 + 23771897 CAGCGCAGGUUAUUGACUUCCCCA-AGUAGCCUCACGUGAGGCUGCGAAUUAAAAAUCCAAACAUUUAUAAUUGCUG----UUCAUUAACCGAACGAA-AAACGGCCUAAAGCACU--GA (((.((.(((((.(((........-.((((((((....)))))))).(((((.((((......)))).)))))....----.))).)))))(..((..-...))..)....)).))--). ( -30.70) >DroSec_CAF1 27971 112 + 1 CAGCGCAGGUUAUUGACUUCCCCA-AGUAGCCUCACGUGAGGCUGCGAAUUAAAAAUCCAAACAUUUAUAAUUGCUG----UUCAUUAACCGGCCGAA-AAACGGCCUAAAGCACC--GA ....((.(((((.(((........-.((((((((....)))))))).(((((.((((......)))).)))))....----.))).)))))(((((..-...)))))....))...--.. ( -36.20) >DroSim_CAF1 28228 113 + 1 CAGCGCAGGUUAUUGACUUCCCCAAAGUAGCCUCACGUGAGGCUGCGAAUUAAAAAUCCAAACAUUUAUAAUUGCUG----UUCAUUAAACGGCCGAA-AAACGGCCUAAAGCACC--GA ..((((((..................((((((((....)))))))).(((((.((((......)))).))))).)))----).........(((((..-...)))))....))...--.. ( -31.10) >DroEre_CAF1 29192 114 + 1 CGGCGCAGGUUAUUGACUCCCCGA-AAUAGCCUCACGUGAGGCUGUGAAUUACCAGUCCAAACAUUUGUAAUUGCUG----UUCAUUAACCGGCCGAA-AAGCGGCCUAAAGCGCCUCCA .(((((.(((((.(((........-.((((((((....)))))))).((((((.(((......))).))))))....----.))).)))))(((((..-...)))))....))))).... ( -44.40) >DroYak_CAF1 29796 112 + 1 AAGCGCAGGUUAUUGACUCCCCGA-AAUAGCCUCACGUGAGGCUGUGAAUUAAAAAUACAAACAUUUAUAAUUGCUG----UUCACUAACCGGCCGAA-AAGCGGCCUAAAGCACC--AA ....((.(((((.(((........-.((((((((....)))))))).(((((.((((......)))).)))))....----.))).)))))(((((..-...)))))....))...--.. ( -33.60) >DroAna_CAF1 26877 114 + 1 ---UUCAGGUUAUUGACUUCCCGA-AAUAGCCUCACGUGAGGCUGUGAAUUAAAAAGCCAAAUAAUUAUGAUGGCUAUUUGCUCAUUAAACGGUUGAAAAAACGGCCUAAGCCUCG--AA ---.((((((((((..........-)))))))).....((((((((((..((((.(((((...........))))).)))).)))).....(((((......)))))..)))))))--). ( -30.40) >consensus CAGCGCAGGUUAUUGACUUCCCCA_AAUAGCCUCACGUGAGGCUGCGAAUUAAAAAUCCAAACAUUUAUAAUUGCUG____UUCAUUAACCGGCCGAA_AAACGGCCUAAAGCACC__GA ....((.(((((.(((..........((((((((....)))))))).(((((.((((......)))).))))).........))).)))))(((((......)))))....))....... (-28.00 = -28.67 + 0.67)


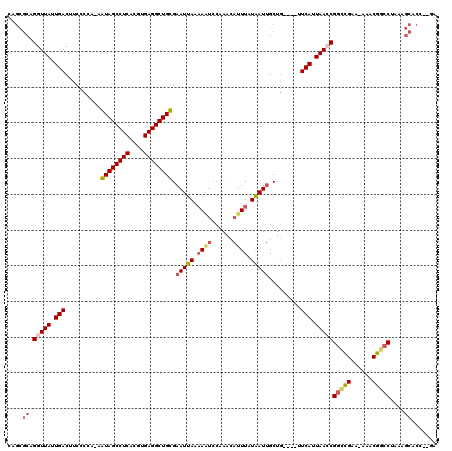
| Location | 17,904,167 – 17,904,279 |
|---|---|
| Length | 112 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 85.64 |
| Mean single sequence MFE | -35.38 |
| Consensus MFE | -26.14 |
| Energy contribution | -27.58 |
| Covariance contribution | 1.45 |
| Combinations/Pair | 1.17 |
| Mean z-score | -2.16 |
| Structure conservation index | 0.74 |
| SVM decision value | -0.05 |
| SVM RNA-class probability | 0.508371 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 17904167 112 - 23771897 UC--AGUGCUUUAGGCCGUUU-UUCGUUCGGUUAAUGAA----CAGCAAUUAUAAAUGUUUGGAUUUUUAAUUCGCAGCCUCACGUGAGGCUACU-UGGGGAAGUCAAUAACCUGCGCUG .(--(((((....(..((...-..))..)(((((.(((.----..(((((((.((((......)))).))))).))((((((....))))))...-........))).))))).)))))) ( -32.30) >DroSec_CAF1 27971 112 - 1 UC--GGUGCUUUAGGCCGUUU-UUCGGCCGGUUAAUGAA----CAGCAAUUAUAAAUGUUUGGAUUUUUAAUUCGCAGCCUCACGUGAGGCUACU-UGGGGAAGUCAAUAACCUGCGCUG .(--(((((....(((((...-..)))))(((((.(((.----..(((((((.((((......)))).))))).))((((((....))))))...-........))).))))).)))))) ( -40.10) >DroSim_CAF1 28228 113 - 1 UC--GGUGCUUUAGGCCGUUU-UUCGGCCGUUUAAUGAA----CAGCAAUUAUAAAUGUUUGGAUUUUUAAUUCGCAGCCUCACGUGAGGCUACUUUGGGGAAGUCAAUAACCUGCGCUG .(--(((((....(((((...-..))))).......(((----((...........))))).(((((((.......((((((....)))))).......)))))))........)))))) ( -33.84) >DroEre_CAF1 29192 114 - 1 UGGAGGCGCUUUAGGCCGCUU-UUCGGCCGGUUAAUGAA----CAGCAAUUACAAAUGUUUGGACUGGUAAUUCACAGCCUCACGUGAGGCUAUU-UCGGGGAGUCAAUAACCUGCGCCG ....(((((...(((..((((-..(((((((((...(((----((...........))))).))))))).......((((((....))))))...-.))..))))......)))))))). ( -41.10) >DroYak_CAF1 29796 112 - 1 UU--GGUGCUUUAGGCCGCUU-UUCGGCCGGUUAGUGAA----CAGCAAUUAUAAAUGUUUGUAUUUUUAAUUCACAGCCUCACGUGAGGCUAUU-UCGGGGAGUCAAUAACCUGCGCUU ..--(((((....(((((...-..)))))((((((((((----.......(((((....))))).......)))))((((((....))))))...-............))))).))))). ( -35.44) >DroAna_CAF1 26877 114 - 1 UU--CGAGGCUUAGGCCGUUUUUUCAACCGUUUAAUGAGCAAAUAGCCAUCAUAAUUAUUUGGCUUUUUAAUUCACAGCCUCACGUGAGGCUAUU-UCGGGAAGUCAAUAACCUGAA--- .(--(.(((.((((((..(((.((((.........)))).)))..)))...........(((((((((..(.....((((((....))))))...-)..))))))))))))))))).--- ( -29.50) >consensus UC__GGUGCUUUAGGCCGUUU_UUCGGCCGGUUAAUGAA____CAGCAAUUAUAAAUGUUUGGAUUUUUAAUUCACAGCCUCACGUGAGGCUACU_UCGGGAAGUCAAUAACCUGCGCUG ....(((((....(((((......)))))(((((.(((......................(((((.....))))).((((((....))))))............))).))))).))))). (-26.14 = -27.58 + 1.45)


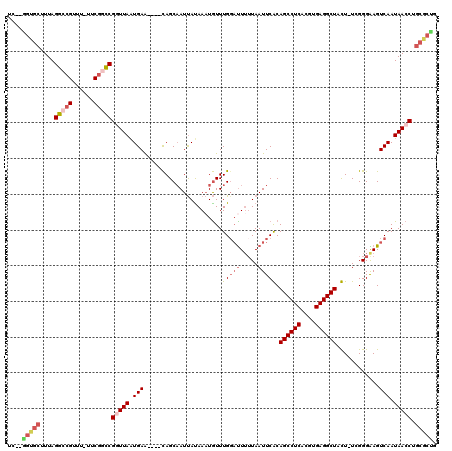
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:28:09 2006