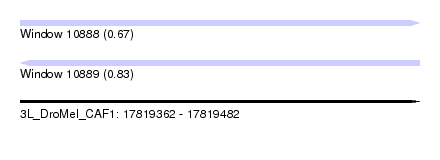

| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 17,819,362 – 17,819,482 |
| Length | 120 |
| Max. P | 0.834982 |

| Location | 17,819,362 – 17,819,482 |
|---|---|
| Length | 120 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 80.39 |
| Mean single sequence MFE | -39.67 |
| Consensus MFE | -22.42 |
| Energy contribution | -22.95 |
| Covariance contribution | 0.53 |
| Combinations/Pair | 1.33 |
| Mean z-score | -2.02 |
| Structure conservation index | 0.57 |
| SVM decision value | 0.28 |
| SVM RNA-class probability | 0.665263 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
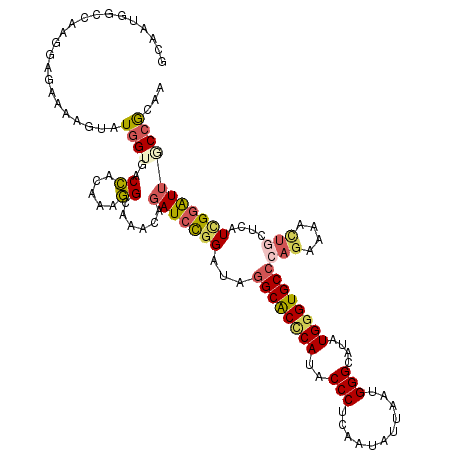
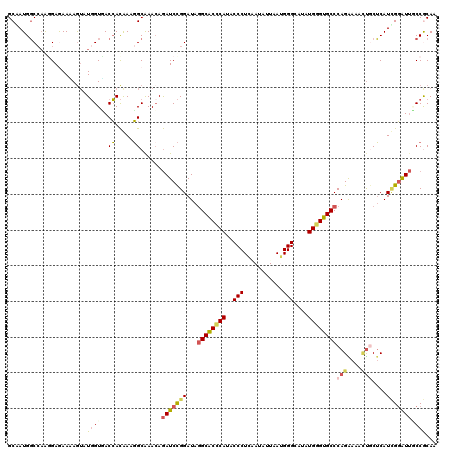
>3L_DroMel_CAF1 17819362 120 + 23771897 GCAAUGGCCAGGGAAAAAAGUAUGGUCACCACGAAGGCAAACAGAUCCGGAUAGGCACCCAUUCCCUCGAUGUUGAGGGGCAUGUGGGUGCCCAGGAAACUGCUCAUCGGAUUGCCGCAA ((..((((((............)))))).......((((.....((((((...((((((((((((((((....)))))))...)))))))))(((....)))....)))))))))))).. ( -52.40) >DroPse_CAF1 1449 120 + 1 GCAAUGGCCAACGAGAACAGUAUGGUGACCACAAAGGCGAACAGAUCCGGAUAGGCACCCAGACCCUGUAUAUUAAUGGGCAUAUGGGUGCCCAGAAAGCUGCUCAUUGGGUUCCCACAA ....(((((((.(((..((((...((..((.....))...))...........((((((((..(((...........)))....))))))))......))))))).))))....)))... ( -32.90) >DroEre_CAF1 1582 120 + 1 GCAAUGGCCAGGGAAAAAAGUAUGGUUACCACGAAGGCAAACAGAUCCGGAUAGGCACUCAUUCCCUCAAUAUUGAGGGGCAUGUGGGUGCCCAGGAAACUACUCAUCGGAUUGCCGCAA ((..((((((............)))))).......((((.....((((((...((((((((((((((((....)))))))...))))))))).((....)).....)))))))))))).. ( -45.70) >DroWil_CAF1 3963 120 + 1 GCUAUGCCCCAGGAGAAUAAUAUCGUGACGAUAAAUGCAAACAAAUCUGGAUACGCACUCAAGCCCUGGACAUUAAUGGGCAAAUGUGUGCCCAGGAAAUCACGCAUGGCAUAACCGCAA ((((((((.((.((........)).)).......((((.......(((((.((((((.....((((...........))))...)))))).))))).......))))))))))...)).. ( -33.24) >DroAna_CAF1 1449 120 + 1 GCGAUGGCAAGGGAGAAGAGAAUGGUCACCACGAAGGCAAAGAGAUCCGGAUAGGCGCCCAUGCCCUCAAUGUUGAUGGGUAGGUGGGUGCCCAGAAAGGUGCUCAUCGGAUUGCCGCAU ..(.((((((....((........(((........)))...(((..((.....((((((((((((((((....))).))))..)))))))))......))..))).))...))))))).. ( -40.90) >DroPer_CAF1 1464 120 + 1 GCAAUGGCCAACGAGAACAGUAUGGUGACCACAAAGGCGAACAGAUCCGGAUAGGCACCCAGACCCUGUAUAUUAAUGGGCAUAUGGGUGCCCAGAAAGCUGCUCAUUGGGUUCCCACAA ....(((((((.(((..((((...((..((.....))...))...........((((((((..(((...........)))....))))))))......))))))).))))....)))... ( -32.90) >consensus GCAAUGGCCAAGGAGAAAAGUAUGGUGACCACAAAGGCAAACAGAUCCGGAUAGGCACCCAUACCCUCAAUAUUAAUGGGCAUAUGGGUGCCCAGAAAACUGCUCAUCGGAUUGCCGCAA ......................((((..((.....))......(((((((...((((((((..(((...........)))....))))))))(((....)))....)))))))))))... (-22.42 = -22.95 + 0.53)

| Location | 17,819,362 – 17,819,482 |
|---|---|
| Length | 120 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 80.39 |
| Mean single sequence MFE | -39.86 |
| Consensus MFE | -25.58 |
| Energy contribution | -25.58 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.27 |
| Mean z-score | -2.05 |
| Structure conservation index | 0.64 |
| SVM decision value | 0.73 |
| SVM RNA-class probability | 0.834982 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
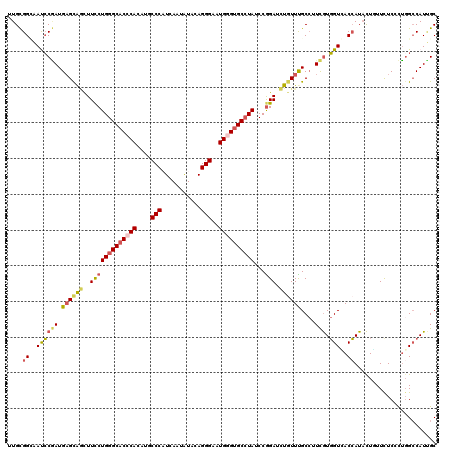
>3L_DroMel_CAF1 17819362 120 - 23771897 UUGCGGCAAUCCGAUGAGCAGUUUCCUGGGCACCCACAUGCCCCUCAACAUCGAGGGAAUGGGUGCCUAUCCGGAUCUGUUUGCCUUCGUGGUGACCAUACUUUUUUCCCUGGCCAUUGC ....(((((........((((..(((((((((((((.....(((((......)))))..))))))))))...))).)))))))))...(..(((.(((............))).)))..) ( -44.10) >DroPse_CAF1 1449 120 - 1 UUGUGGGAACCCAAUGAGCAGCUUUCUGGGCACCCAUAUGCCCAUUAAUAUACAGGGUCUGGGUGCCUAUCCGGAUCUGUUCGCCUUUGUGGUCACCAUACUGUUCUCGUUGGCCAUUGC .(((((..(((((((((((((..(((((((((((((...((((...........)))).))))))))))...))).)))))))...))).)))..))))).((..(.....)..)).... ( -43.10) >DroEre_CAF1 1582 120 - 1 UUGCGGCAAUCCGAUGAGUAGUUUCCUGGGCACCCACAUGCCCCUCAAUAUUGAGGGAAUGAGUGCCUAUCCGGAUCUGUUUGCCUUCGUGGUAACCAUACUUUUUUCCCUGGCCAUUGC ....(((.....((.(((((...((((((((((...(((..((((((....)))))).))).)))))))...)))..((.(((((.....))))).)))))))...))....)))..... ( -38.20) >DroWil_CAF1 3963 120 - 1 UUGCGGUUAUGCCAUGCGUGAUUUCCUGGGCACACAUUUGCCCAUUAAUGUCCAGGGCUUGAGUGCGUAUCCAGAUUUGUUUGCAUUUAUCGUCACGAUAUUAUUCUCCUGGGGCAUAGC .....(((((((((((((.......((((..((.(((((((((...........))))..))))).))..)))).......))))).(((((...)))))............)))))))) ( -35.94) >DroAna_CAF1 1449 120 - 1 AUGCGGCAAUCCGAUGAGCACCUUUCUGGGCACCCACCUACCCAUCAACAUUGAGGGCAUGGGCGCCUAUCCGGAUCUCUUUGCCUUCGUGGUGACCAUUCUCUUCUCCCUUGCCAUCGC ....(((((((((..(((.....)))(((((.((((....(((.(((....))))))..)))).)))))..)))).....(..((.....))..)...............)))))..... ( -34.70) >DroPer_CAF1 1464 120 - 1 UUGUGGGAACCCAAUGAGCAGCUUUCUGGGCACCCAUAUGCCCAUUAAUAUACAGGGUCUGGGUGCCUAUCCGGAUCUGUUCGCCUUUGUGGUCACCAUACUGUUCUCGUUGGCCAUUGC .(((((..(((((((((((((..(((((((((((((...((((...........)))).))))))))))...))).)))))))...))).)))..))))).((..(.....)..)).... ( -43.10) >consensus UUGCGGCAAUCCGAUGAGCAGCUUCCUGGGCACCCACAUGCCCAUCAAUAUACAGGGAAUGGGUGCCUAUCCGGAUCUGUUUGCCUUCGUGGUCACCAUACUGUUCUCCCUGGCCAUUGC ....((..((((((.((((((..(((((((((((((....(((...........)))..))))))))))...))).))))))....))).)))..))....................... (-25.58 = -25.58 + 0.00)
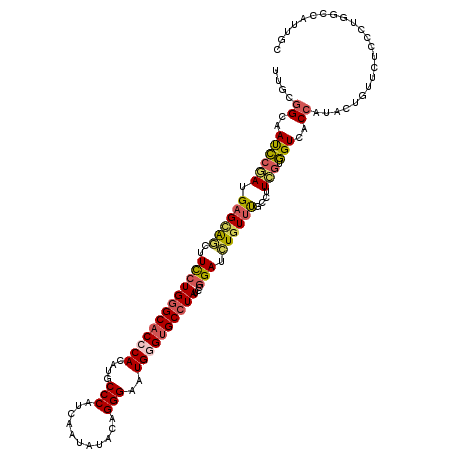


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:27:42 2006