| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 1,967,303 – 1,967,401 |
| Length | 98 |
| Max. P | 0.944576 |

| Location | 1,967,303 – 1,967,401 |
|---|---|
| Length | 98 |
| Sequences | 5 |
| Columns | 98 |
| Reading direction | forward |
| Mean pairwise identity | 91.84 |
| Mean single sequence MFE | -30.14 |
| Consensus MFE | -21.76 |
| Energy contribution | -22.16 |
| Covariance contribution | 0.40 |
| Combinations/Pair | 1.12 |
| Mean z-score | -3.45 |
| Structure conservation index | 0.72 |
| SVM decision value | 1.35 |
| SVM RNA-class probability | 0.944576 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 1967303 98 + 23771897 UGCUGACCAUGCUUAAAUGCGGCUCAUUUAUCAAUAGAGCUGCUUAGCAUGGAAAGGACACAAGGAGAAAGCAUCCUGCAUCCCUUUAAGCACUUUAU ((((..(((((((.....(((((((...........)))))))..)))))))(((((.....((((.......)))).....))))).))))...... ( -34.60) >DroSec_CAF1 18412 94 + 1 UGCUGACCAUGCUUAAAUGCGGCUCAUUUAUCAAUA--GCUGCUUAGCAUGGAAAGGACACAAGGAGAAAG--UCCUGCAUCCCUUUAAGCUCUUUUU .(((..(((((((.....((((((.(((....))))--)))))..)))))))(((((.....((((.....--)))).....))))).)))....... ( -29.80) >DroSim_CAF1 32594 94 + 1 UGCUGACCAUGCUUAAAUGCGGCUCAUUUAUCAAUA--GCUGCUUAGCAUGGAAAGGACACAAGGAGAAAG--UCCUGCAUCCCUUUAAGCACUUUUU ((((..(((((((.....((((((.(((....))))--)))))..)))))))(((((.....((((.....--)))).....))))).))))...... ( -30.70) >DroEre_CAF1 18421 94 + 1 UGCUGACCAUGCUUAAAUGCGGCUCAUUUAUCAAUA--GCUGCUUAGCAUGGAAAGGACACGAGAAGAAAG--UCCUGCACCGCCUUAAAUACUUUUA (((...(((((((.....((((((.(((....))))--)))))..)))))))..(((((...........)--))))))).................. ( -26.60) >DroYak_CAF1 18908 94 + 1 UGCUGACCAUGCUUAAAUGCGGCUCAUUUAUCAAUA--GCUGCUUAGCAUGGAAGGCACACGAGGAGAAAG--UCCUGGACCGCUUUAAAUACUUUUU ((((..(((((((.....((((((.(((....))))--)))))..)))))))..))))..(.((((.....--)))).)................... ( -29.00) >consensus UGCUGACCAUGCUUAAAUGCGGCUCAUUUAUCAAUA__GCUGCUUAGCAUGGAAAGGACACAAGGAGAAAG__UCCUGCAUCCCUUUAAGCACUUUUU .(((..(((((((.....(((((...............)))))..)))))))(((((.....((((.......)))).....))))).)))....... (-21.76 = -22.16 + 0.40)

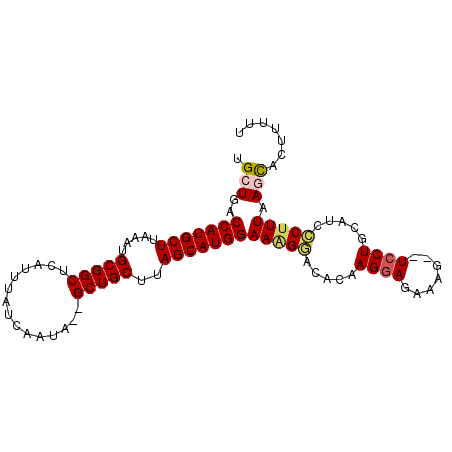
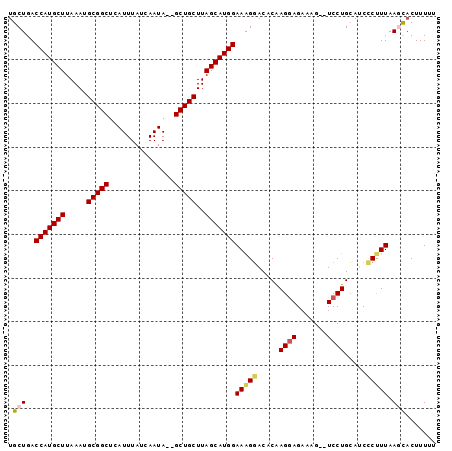
| Location | 1,967,303 – 1,967,401 |
|---|---|
| Length | 98 |
| Sequences | 5 |
| Columns | 98 |
| Reading direction | reverse |
| Mean pairwise identity | 91.84 |
| Mean single sequence MFE | -27.94 |
| Consensus MFE | -21.10 |
| Energy contribution | -20.98 |
| Covariance contribution | -0.12 |
| Combinations/Pair | 1.15 |
| Mean z-score | -2.37 |
| Structure conservation index | 0.76 |
| SVM decision value | 1.05 |
| SVM RNA-class probability | 0.906839 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
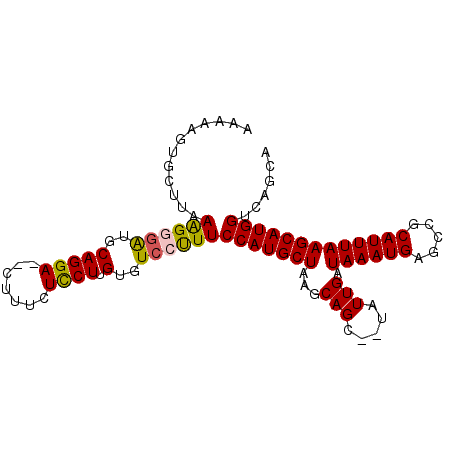
>3L_DroMel_CAF1 1967303 98 - 23771897 AUAAAGUGCUUAAAGGGAUGCAGGAUGCUUUCUCCUUGUGUCCUUUCCAUGCUAAGCAGCUCUAUUGAUAAAUGAGCCGCAUUUAAGCAUGGUCAGCA ......((((..(((((((((((((.......))).))))))))))(((((((..((.((((.(((....))))))).)).....)))))))..)))) ( -33.30) >DroSec_CAF1 18412 94 - 1 AAAAAGAGCUUAAAGGGAUGCAGGA--CUUUCUCCUUGUGUCCUUUCCAUGCUAAGCAGC--UAUUGAUAAAUGAGCCGCAUUUAAGCAUGGUCAGCA .......(((..((((((((((((.--.......))))))))))))(((((((..((.((--((((....))).))).)).....)))))))..))). ( -30.40) >DroSim_CAF1 32594 94 - 1 AAAAAGUGCUUAAAGGGAUGCAGGA--CUUUCUCCUUGUGUCCUUUCCAUGCUAAGCAGC--UAUUGAUAAAUGAGCCGCAUUUAAGCAUGGUCAGCA ......((((..((((((((((((.--.......))))))))))))(((((((..((.((--((((....))).))).)).....)))))))..)))) ( -30.30) >DroEre_CAF1 18421 94 - 1 UAAAAGUAUUUAAGGCGGUGCAGGA--CUUUCUUCUCGUGUCCUUUCCAUGCUAAGCAGC--UAUUGAUAAAUGAGCCGCAUUUAAGCAUGGUCAGCA .............(((.((((((((--(...........)))))...........((.((--((((....))).))).))......)))).))).... ( -22.10) >DroYak_CAF1 18908 94 - 1 AAAAAGUAUUUAAAGCGGUCCAGGA--CUUUCUCCUCGUGUGCCUUCCAUGCUAAGCAGC--UAUUGAUAAAUGAGCCGCAUUUAAGCAUGGUCAGCA ..............((((..((.((--........)).))..))..(((((((..((.((--((((....))).))).)).....)))))))...)). ( -23.60) >consensus AAAAAGUGCUUAAAGGGAUGCAGGA__CUUUCUCCUUGUGUCCUUUCCAUGCUAAGCAGC__UAUUGAUAAAUGAGCCGCAUUUAAGCAUGGUCAGCA ............((((((..(((((.......)))).)..))))))(((((((...(((.....))).((((((.....)))))))))))))...... (-21.10 = -20.98 + -0.12)

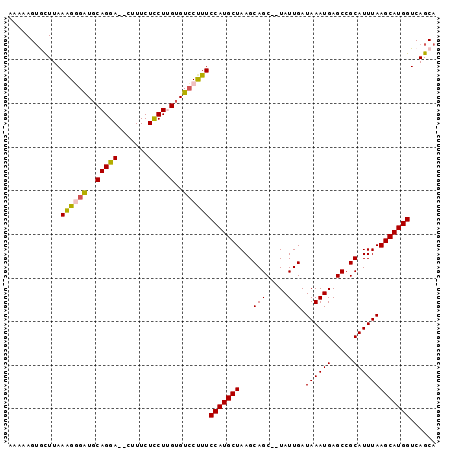
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:47:56 2006