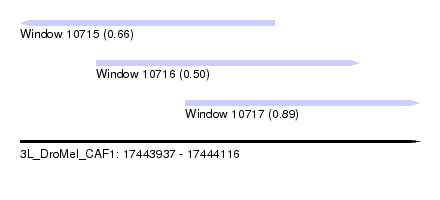

| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 17,443,937 – 17,444,116 |
| Length | 179 |
| Max. P | 0.891084 |

| Location | 17,443,937 – 17,444,051 |
|---|---|
| Length | 114 |
| Sequences | 6 |
| Columns | 114 |
| Reading direction | reverse |
| Mean pairwise identity | 80.70 |
| Mean single sequence MFE | -45.25 |
| Consensus MFE | -28.60 |
| Energy contribution | -29.72 |
| Covariance contribution | 1.12 |
| Combinations/Pair | 1.29 |
| Mean z-score | -1.71 |
| Structure conservation index | 0.63 |
| SVM decision value | 0.26 |
| SVM RNA-class probability | 0.658043 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 17443937 114 - 23771897 UCGAGCCAAAGCCCGAUUGGAGAACGAGGUGGUUCUCCUGCGGAGCGAGGUGUCCUCGCUGCGCUCCAAGCACAACCAACAUCGUAAGUAUCGGCAGGAGCAGGUGGUUCUGCA ..((((((..(((((...(((((((......)))))))..)))(((((((...)))))))))(((((..((...((..((...))..))....)).)))))...)))))).... ( -45.20) >DroSec_CAF1 3082 114 - 1 UCGGGCCAAAGCCCGAUUGGAGAACGAGGUGGUUCUCCUGCGUGGCGAGGUGUCCUCGCUGCGCUCCAAGCACAACCAACAUCGUAAGUAUCGGCAGGAGCAGGCGGUUCUACA ((((((....))))))..(((((((......)))))))((((..((((((...))))))..)(((((..((...((..((...))..))....)).)))))..)))........ ( -46.30) >DroSim_CAF1 3530 114 - 1 UCGGGCCAAAGCCCGAUUGGAGAACGAGGUGGUUCUCCUGCGUGGCGAGGUGUCCUCGCUGCGCUCCAAGCACAAUCAGCAUCGUAAGUAUCGGCAGGAGCAGGCGGUUCUACA ((((((....))))))..(((((((......)))))))((((..((((((...))))))..)(((((..((.......)).(((.......)))..)))))..)))........ ( -48.80) >DroEre_CAF1 3281 114 - 1 CCGGGCCAAAGCCCGUUUGGAAAACGAGGUGGUUCUCCUGCGACGCGAGGUGGCCUCACUGCGCUCCAGGCAUAUCCAGCAUCGUAAGUAUCGACAGGAACAGGCGGUUCUACA ((((((....))))(((((....(((((.(((....((((.((.(((.((((....)))).))))))))).....))).).))))..((....)).....)))))))....... ( -37.10) >DroYak_CAF1 3286 114 - 1 UCGGGCCAAAGCCCGACUGGAAAACGAGGUGGUCCUCCUUCGCCGCGAGGUGGCCUCACUGCGCUCCAAGCAUAUCAAACAUCGUAAGUAUCGCCAGAAUCAGGCGGUGCUACA ((((((....))))))...((....(((((((....))..((((....)))))))))..(((.......)))..)).......(((.((((((((.......))))))))))). ( -43.60) >DroAna_CAF1 3027 114 - 1 UCGCGCCAAGGCGCGACUGGAACACGAGAUAACCAUCUUACGCCGCGAGGUGGCCAGGCUGCGCGCCAAACAGGACCAGCGACGACGUGACAAGCUGGAGAAGGAUGUGGCGCA ..((((((.((((((.((((.....(((((....))))).((((....)))).))))....))))))........(((((.............))))).........)))))). ( -50.52) >consensus UCGGGCCAAAGCCCGAUUGGAAAACGAGGUGGUUCUCCUGCGCCGCGAGGUGGCCUCGCUGCGCUCCAAGCACAACCAACAUCGUAAGUAUCGGCAGGAGCAGGCGGUUCUACA ((((((....))))))(((((....((((....))))..((((.((((((...)))))).)))))))))..............((.((.((((.(.......).)))).)))). (-28.60 = -29.72 + 1.12)



| Location | 17,443,971 – 17,444,089 |
|---|---|
| Length | 118 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 81.01 |
| Mean single sequence MFE | -46.32 |
| Consensus MFE | -34.04 |
| Energy contribution | -33.72 |
| Covariance contribution | -0.33 |
| Combinations/Pair | 1.45 |
| Mean z-score | -1.11 |
| Structure conservation index | 0.73 |
| SVM decision value | -0.07 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 17443971 118 + 23771897 GUUGGUUGUGCUUGGAGCGCAGCGAGGACACCUCGCUCCGCAGGAGAACCACCUCGUUCUCCAAUCGGGCUUUGGCUCGAUCGUGGUAUGGCAUCGCGCUGGACGAA--UGAAAUUUUGG ..(((((.(.((....(((.(((((((...))))))).))))).).)))))..(((((((((((((((((....)))))))((((((.....)))))).)))).)))--)))........ ( -47.00) >DroSec_CAF1 3116 118 + 1 GUUGGUUGUGCUUGGAGCGCAGCGAGGACACCUCGCCACGCAGGAGAACCACCUCGUUCUCCAAUCGGGCUUUGGCCCGAUCGUGGUAUGGCAUCGCGCUGGACAAA--UCAAAUUUUGG .((((((((((.....))))((((..((..((..((((((..(((((((......))))))).(((((((....)))))))))))))..))..)).)))).....))--))))....... ( -49.90) >DroSim_CAF1 3564 118 + 1 GCUGAUUGUGCUUGGAGCGCAGCGAGGACACCUCGCCACGCAGGAGAACCACCUCGUUCUCCAAUCGGGCUUUGGCCCGAUCGUGGUAUGGCAUCGCGCUGGACAAA--UCAAAUUUUGG ..(((((((((.....))))((((..((..((..((((((..(((((((......))))))).(((((((....)))))))))))))..))..)).)))).....))--)))........ ( -49.50) >DroEre_CAF1 3315 118 + 1 GCUGGAUAUGCCUGGAGCGCAGUGAGGCCACCUCGCGUCGCAGGAGAACCACCUCGUUUUCCAAACGGGCUUUGGCCCGGUCGUGGUACGGCAUCGGGCUGGACGAA--UUAAAUUUUGG .........(((((..(((..((((((...))))))..))).(((((((......)))))))...)))))((..(((((((((.....))))..)))))..))((((--......)))). ( -47.50) >DroYak_CAF1 3320 118 + 1 GUUUGAUAUGCUUGGAGCGCAGUGAGGCCACCUCGCGGCGAAGGAGGACCACCUCGUUUUCCAGUCGGGCUUUGGCCCGAUCGUGGUACGGCAUUGGGCUGGAUGAG--UCAAAUUUUGG (((((((...((...(((.(((((..(((((.....((.(((.((((....)))).))).)).(((((((....))))))).)))))....))))).)))))....)--))))))..... ( -45.80) >DroAna_CAF1 3061 118 + 1 GCUGGUCCUGUUUGGCGCGCAGCCUGGCCACCUCGCGGCGUAAGAUGGUUAUCUCGUGUUCCAGUCGCGCCUUGGCGCGAUCAUGAAGGGGCAUCC--UUGGGCAAUGUUUAGAUUCGCG ((..(......)..))((.(((....(((.(((((..(((..((((....))))..)))..).(((((((....)))))))...).))))))....--))).))................ ( -38.20) >consensus GCUGGUUGUGCUUGGAGCGCAGCGAGGACACCUCGCGACGCAGGAGAACCACCUCGUUCUCCAAUCGGGCUUUGGCCCGAUCGUGGUAUGGCAUCGCGCUGGACAAA__UCAAAUUUUGG ....((((((((((..(((..((((((...))))))..))).(((((((......))))))).(((((((....))))))))).))))))))............................ (-34.04 = -33.72 + -0.33)

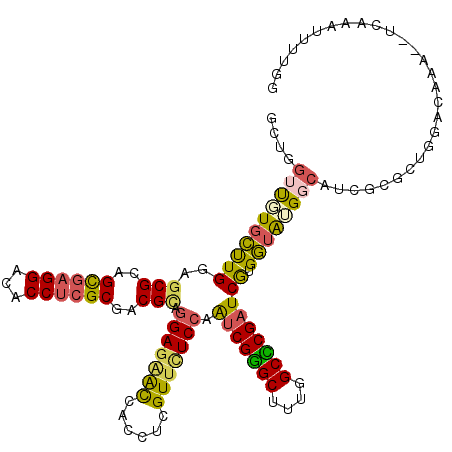

| Location | 17,444,011 – 17,444,116 |
|---|---|
| Length | 105 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 78.42 |
| Mean single sequence MFE | -35.13 |
| Consensus MFE | -25.23 |
| Energy contribution | -24.71 |
| Covariance contribution | -0.52 |
| Combinations/Pair | 1.44 |
| Mean z-score | -1.66 |
| Structure conservation index | 0.72 |
| SVM decision value | 0.96 |
| SVM RNA-class probability | 0.891084 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 17444011 105 + 23771897 CAGGAGAACCACCUCGUUCUCCAAUCGGGCUUUGGCUCGAUCGUGGUAUGGCAUCGCGCUGGACGAA--UGAAAUUUUGGUA-GUCCAAAA---------CACACGGCUUG---UCUAUU ..(((((((......))))))).(((((((....)))))))((((((.....)))))).(((((((.--((...((((((..-..))))))---------....))..)))---)))).. ( -34.60) >DroSec_CAF1 3156 105 + 1 CAGGAGAACCACCUCGUUCUCCAAUCGGGCUUUGGCCCGAUCGUGGUAUGGCAUCGCGCUGGACAAA--UCAAAUUUUGGUA-GUCCAAAA---------CACACGGCUUG---UCUAUU ..(((((((......))))))).(((((((....)))))))((((((.....)))))).(((((((.--.(...((((((..-..))))))---------.....)..)))---)))).. ( -38.80) >DroSim_CAF1 3604 105 + 1 CAGGAGAACCACCUCGUUCUCCAAUCGGGCUUUGGCCCGAUCGUGGUAUGGCAUCGCGCUGGACAAA--UCAAAUUUUGGUA-GUCCUAAA---------CACACGGCUUG---UCUAUU ..(((((((......))))))).(((((((....)))))))((((((..(((.....)))((((..(--(((.....)))).-))))...)---------).)))).....---...... ( -35.80) >DroEre_CAF1 3355 108 + 1 CAGGAGAACCACCUCGUUUUCCAAACGGGCUUUGGCCCGGUCGUGGUACGGCAUCGGGCUGGACGAA--UUAAAUUUUGGUA-GUCCAAAA---------CACACUGCUUGUCGUCUAUU ..(((((((......)))))))..((((((..(((((((((((.....))))..)))))(((((..(--(((.....)))).-)))))...---------..))..))))))........ ( -35.70) >DroYak_CAF1 3360 105 + 1 AAGGAGGACCACCUCGUUUUCCAGUCGGGCUUUGGCCCGAUCGUGGUACGGCAUUGGGCUGGAUGAG--UCAAAUUUUGGUA-GUCCAAAA---------CACACUGCUUG---UCUAGU ...((((....))))(((..((((((((((....)))))))..)))...))).....((((((..((--(....((((((..-..))))))---------......)))..---)))))) ( -37.90) >DroAna_CAF1 3101 115 + 1 UAAGAUGGUUAUCUCGUGUUCCAGUCGCGCCUUGGCGCGAUCAUGAAGGGGCAUCC--UUGGGCAAUGUUUAGAUUCGCGUACGUACAAAGACAAAAUUUCGCAUGGGUUC---UUUACC ......(((...((((((((((.(((((((....)))))))....((((.....))--)))))...(((((.....((....)).....))))).......)))))))...---...))) ( -28.00) >consensus CAGGAGAACCACCUCGUUCUCCAAUCGGGCUUUGGCCCGAUCGUGGUAUGGCAUCGCGCUGGACAAA__UCAAAUUUUGGUA_GUCCAAAA_________CACACGGCUUG___UCUAUU ..(((((((......))))))).(((((((....)))))))((((....(((.....)))((((...................))))...............)))).............. (-25.23 = -24.71 + -0.52)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:24:56 2006