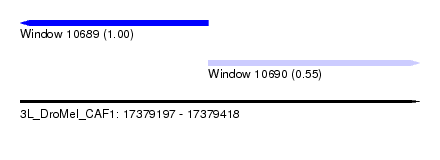
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 17,379,197 – 17,379,418 |
| Length | 221 |
| Max. P | 0.999398 |

| Location | 17,379,197 – 17,379,301 |
|---|---|
| Length | 104 |
| Sequences | 5 |
| Columns | 104 |
| Reading direction | reverse |
| Mean pairwise identity | 96.06 |
| Mean single sequence MFE | -25.28 |
| Consensus MFE | -23.58 |
| Energy contribution | -24.18 |
| Covariance contribution | 0.60 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.24 |
| Structure conservation index | 0.93 |
| SVM decision value | 3.57 |
| SVM RNA-class probability | 0.999398 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
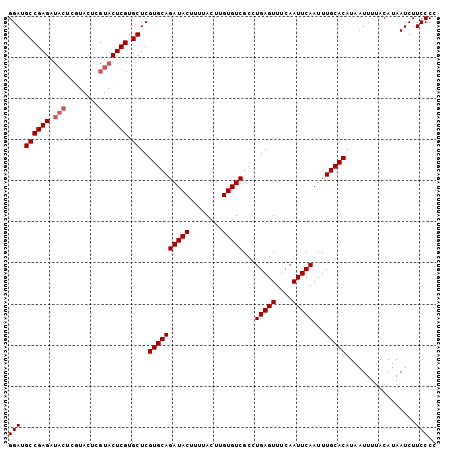
>3L_DroMel_CAF1 17379197 104 - 23771897 GGAUGCCGAGAUACUCGUACUCGUACUCGUGCUCGUGCAGAUACUUUUACUUGUGUCGCCUGAGUUUCAAUUCAAUUUGCACAUAAUUUUACAUAAUCUUCCCC (((.((((((.(((........))))))).))..((((((((((........)))))...(((((....)))))...))))).................))).. ( -25.50) >DroSec_CAF1 6080 104 - 1 GGAUGCCGAGAUACUCGUACUCGUACUCGUGCUCGUGCAGAUACUUUUACUUGUGUCGCCUGAGUUUCAAUUCAAUUUGCACAUAAUUUUACAUAAUCUUCCCC (((.((((((.(((........))))))).))..((((((((((........)))))...(((((....)))))...))))).................))).. ( -25.50) >DroSim_CAF1 5772 104 - 1 GGAUGCCGAGAUACUCGUACUGGUACUCGUGCUCGUGCAGAUACUUUUACUUGUGUCGCCUGAGUUUCAAUUCAAUUUGCACAUAAUUUUACAUAAUCUUCCCC (((.((((((.((((......)))))))).))..((((((((((........)))))...(((((....)))))...))))).................))).. ( -26.40) >DroEre_CAF1 6230 98 - 1 GGAUGCCGAGA------UACUCGUACUCGUGCUCGUGCAGAUACUUUUACUUGUGUCGCCUGAGUUUCAAUUCAAUUUGCACAUAAUUUUACAUAAUCUUCCCG (((.((((((.------((....)))))).))..((((((((((........)))))...(((((....)))))...))))).................))).. ( -23.40) >DroYak_CAF1 7133 104 - 1 GGAUGCCGAGAUACAGAUACUCGUACUCGUGCUCGUGCAGAUACUUUUACUUGUGUCGCCUGAGUUUCAAUUCAAUUUGCACAUAAUUUUACAUAAUCUUCCCC (((.((((((.(((........))))))).))..((((((((((........)))))...(((((....)))))...))))).................))).. ( -25.60) >consensus GGAUGCCGAGAUACUCGUACUCGUACUCGUGCUCGUGCAGAUACUUUUACUUGUGUCGCCUGAGUUUCAAUUCAAUUUGCACAUAAUUUUACAUAAUCUUCCCC (((.((((((.(((........))))))).))..((((((((((........)))))...(((((....)))))...))))).................))).. (-23.58 = -24.18 + 0.60)


| Location | 17,379,301 – 17,379,418 |
|---|---|
| Length | 117 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 79.86 |
| Mean single sequence MFE | -31.08 |
| Consensus MFE | -21.88 |
| Energy contribution | -22.18 |
| Covariance contribution | 0.31 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.15 |
| Structure conservation index | 0.70 |
| SVM decision value | 0.03 |
| SVM RNA-class probability | 0.548455 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 17379301 117 + 23771897 G--CGCGUGUCUGUAUCGCAUUGAUUGCCCCCUAUUGAUUGUCCAAUUCCAAUCGAGUCUAAUUAAUGCUGUCCGCUCUGGGGACACGGACCGCGGACCACUGCAAA-AUCUCGCCUAGC (--((.((((((((...((((((((((..(....(((((((........))))))))..)))))))))).(((((...((....))))))).))))).))))))...-............ ( -35.30) >DroPse_CAF1 5055 101 + 1 AUAUGUGUAUCUGUAUCGCUUUGAUUGCUCUUUAUUGAUUGUU------CAAUCGAAUGUAAUUAAUGCUGUCCGCUGUGGGGACACGGACUA-------CUGCAAAUAUCUCU------ ...(((((((((((...((.((((((((..((.(((((....)------)))).))..)))))))).)).((((.(...).))))))))).))-------).))).........------ ( -26.30) >DroSec_CAF1 6184 117 + 1 G--CGCGUGUCUGUAUCGCAUUGAUUGCCCCCUAUUGAUUGUCCAAUUCCAAUCGAGUCUAAUUAAUGCUGUCCGCUCUGGGGACACGGACCACGGACCAGUGCAAA-AUCUUGCCGGGC (--(((..((((((...((((((((((..(....(((((((........))))))))..)))))))))).(((((...((....))))))).))))))..))))...-............ ( -37.40) >DroEre_CAF1 6328 110 + 1 G--CGCGUGUCGGUAUCGCAUUGAUUGCCUCUUAUUGAAUGUCCAAUUUCAAUCGAGUCUAAUUAAUGCUGUCCGCUCUGGGGAUACGGACCA-------CAGCAAA-AUCUCGCCGAGC (--(..(((((.(((((((((((((((.(((..((((((........)))))).)))..))))))))))...((......))))))).)).))-------).))...-............ ( -32.00) >DroYak_CAF1 7237 110 + 1 G--CGCGUGUCGGUAUCGCAUUGAUUGCCUCUUAUUGAAUGUCCAAUUCCAAUCGAGUCUAAUUAAUGCUGUCCGCUCUGGGGACACGGACCA-------CUGCAAA-AUCUCGCCGGGC .--......(((((...((((((((((.(((.....((((.....)))).....)))..)))))))))).(((((...((....)))))))..-------.......-.....))))).. ( -29.20) >DroPer_CAF1 5070 100 + 1 AUAUGUGUAUCUGUAUCGCUUUGAUUGCUCUUUAUUGAUUGUU------CAAUCGAAUGUAAUUAAUGCUGUCCGCUGUGGGGACACGGACUA-------CUGCAAAUAUCUC------- ...(((((((((((...((.((((((((..((.(((((....)------)))).))..)))))))).)).((((.(...).))))))))).))-------).)))........------- ( -26.30) >consensus G__CGCGUGUCUGUAUCGCAUUGAUUGCCCCUUAUUGAUUGUCCAAUUCCAAUCGAGUCUAAUUAAUGCUGUCCGCUCUGGGGACACGGACCA_______CUGCAAA_AUCUCGCCG_GC ....((...........((((((((((.......(((((((........)))))))...)))))))))).(((((...((....)))))))...........))................ (-21.88 = -22.18 + 0.31)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:24:31 2006