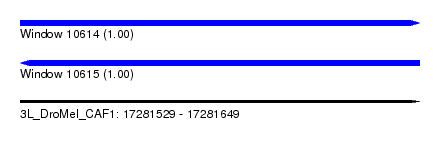
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 17,281,529 – 17,281,649 |
| Length | 120 |
| Max. P | 0.999601 |

| Location | 17,281,529 – 17,281,649 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 94.50 |
| Mean single sequence MFE | -45.70 |
| Consensus MFE | -38.10 |
| Energy contribution | -37.94 |
| Covariance contribution | -0.16 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.83 |
| Structure conservation index | 0.83 |
| SVM decision value | 2.91 |
| SVM RNA-class probability | 0.997713 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 17281529 120 + 23771897 CAAGUGCUCGAAAUAAUAAAGUUAACAGCACUUUAAUAGUUACCGCCCGCCACACCCACGCCGCCCACACGGAGGGCGUAGCGAAUCAGCGGGCGGGGGUGUGGGAAACGGCAACCGAAC .(((((((...(((......)))...))))))).....(((.(((....((((((((.((((((((.......))))...((......)).)))).))))))))....))).)))..... ( -47.10) >DroSec_CAF1 165216 120 + 1 CAAGUGCUCGAAAUAAUAAAGUUAACAGCACUUUAAUAGUUACCGCCCGCCACACCCACGCCGCCCAUACGGUGGGCGCAGCGAAUCAGGGGGCGUGGGAGUGGGAAACGGCAACCGAAC .(((((((...(((......)))...))))))).....(((.(((....((((.((((((((((((((...))))))....(......)..)))))))).))))....))).)))..... ( -45.00) >DroSim_CAF1 161452 120 + 1 CAAGUGCUCGAAAUAAUAAAGUUAACAGCACUUUAAUAGUUACCGCCCGCCACACCCACGCCGCCCACACGGUGGGCGUAGCGAAUCAGGGGGCGUGGGUGUGGGAAACGGCAACCGAAC .(((((((...(((......)))...))))))).....(((.(((....(((((((((((((((((((...))))))....(......)..)))))))))))))....))).)))..... ( -51.00) >DroEre_CAF1 168235 119 + 1 CAAGUGCUCGAAAUAAUAAAGUUAACAGCACUUUAAUAGUUACCGCCUGCCACUCCCACGUCGCCCACACGG-GGGCGUAACGAAUCUGGGGGCGGGGGUGUGGGAAACGGCAACCGAAC .(((((((...(((......)))...)))))))........(((.((((((...((((((((((((......-)))))..)))....)))))))))))))((.(....).))........ ( -41.70) >DroYak_CAF1 173197 119 + 1 CAAGUGCUCGAAAUAAUAAAGUUAACAGCACUUUAAUAGUUACCGCCCGCCACUCCCACGCCGCCCACAAGG-GGGCGUAACGAAUCAGGGGGCGGGGGUGUGGCAAACGGCAACCAAAC .(((((((...(((......)))...))))))).....(((.(((...(((((.(((.((((((((......-))))....(......)..)))).))).)))))...))).)))..... ( -43.70) >consensus CAAGUGCUCGAAAUAAUAAAGUUAACAGCACUUUAAUAGUUACCGCCCGCCACACCCACGCCGCCCACACGG_GGGCGUAGCGAAUCAGGGGGCGGGGGUGUGGGAAACGGCAACCGAAC .(((((((...(((......)))...))))))).....(((.(((....((((.(((.((((((((.......))))....(......)..)))).))).))))....))).)))..... (-38.10 = -37.94 + -0.16)


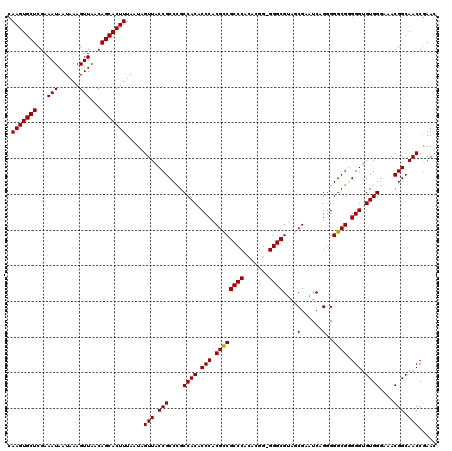
| Location | 17,281,529 – 17,281,649 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 94.50 |
| Mean single sequence MFE | -55.27 |
| Consensus MFE | -47.46 |
| Energy contribution | -47.34 |
| Covariance contribution | -0.12 |
| Combinations/Pair | 1.05 |
| Mean z-score | -4.40 |
| Structure conservation index | 0.86 |
| SVM decision value | 3.77 |
| SVM RNA-class probability | 0.999601 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
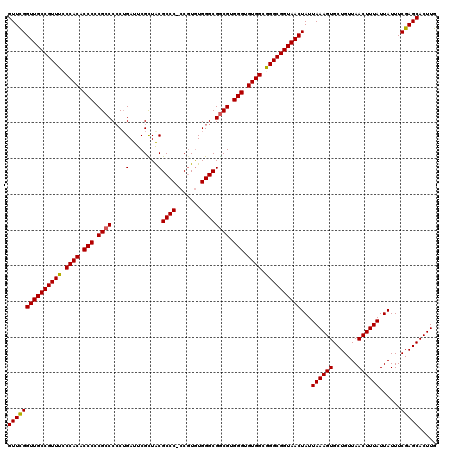
>3L_DroMel_CAF1 17281529 120 - 23771897 GUUCGGUUGCCGUUUCCCACACCCCCGCCCGCUGAUUCGCUACGCCCUCCGUGUGGGCGGCGUGGGUGUGGCGGGCGGUAACUAUUAAAGUGCUGUUAACUUUAUUAUUUCGAGCACUUG ((((((((((((((..((((((((.((((.((......))...((((.......)))))))).))))))))..)))))))))...((((((.......))))))......)))))..... ( -58.60) >DroSec_CAF1 165216 120 - 1 GUUCGGUUGCCGUUUCCCACUCCCACGCCCCCUGAUUCGCUGCGCCCACCGUAUGGGCGGCGUGGGUGUGGCGGGCGGUAACUAUUAAAGUGCUGUUAACUUUAUUAUUUCGAGCACUUG ((((((((((((((..((((.(((((((((((.....((.((....)).))...))).)))))))).))))..)))))))))...((((((.......))))))......)))))..... ( -56.60) >DroSim_CAF1 161452 120 - 1 GUUCGGUUGCCGUUUCCCACACCCACGCCCCCUGAUUCGCUACGCCCACCGUGUGGGCGGCGUGGGUGUGGCGGGCGGUAACUAUUAAAGUGCUGUUAACUUUAUUAUUUCGAGCACUUG ((((((((((((((..(((((((((((((....(........)((((((...)))))))))))))))))))..)))))))))...((((((.......))))))......)))))..... ( -62.20) >DroEre_CAF1 168235 119 - 1 GUUCGGUUGCCGUUUCCCACACCCCCGCCCCCAGAUUCGUUACGCCC-CCGUGUGGGCGACGUGGGAGUGGCAGGCGGUAACUAUUAAAGUGCUGUUAACUUUAUUAUUUCGAGCACUUG (((((((((((((((.((((.(((((((((...........((((..-..)))))))))..).))).)))).))))))))))...((((((.......))))))......)))))..... ( -44.40) >DroYak_CAF1 173197 119 - 1 GUUUGGUUGCCGUUUGCCACACCCCCGCCCCCUGAUUCGUUACGCCC-CCUUGUGGGCGGCGUGGGAGUGGCGGGCGGUAACUAUUAAAGUGCUGUUAACUUUAUUAUUUCGAGCACUUG ...(((((((((((((((((.(((.((((...(((....))).((((-......)))))))).))).)))))))))))))))))...(((((((..................))))))). ( -54.57) >consensus GUUCGGUUGCCGUUUCCCACACCCCCGCCCCCUGAUUCGCUACGCCC_CCGUGUGGGCGGCGUGGGUGUGGCGGGCGGUAACUAUUAAAGUGCUGUUAACUUUAUUAUUUCGAGCACUUG (((((((((((((((.((((.(((.((((....(........)((((.......)))))))).))).)))).))))))))))...((((((.......))))))......)))))..... (-47.46 = -47.34 + -0.12)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:23:20 2006