
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 16,870,662 – 16,870,762 |
| Length | 100 |
| Max. P | 0.891108 |

| Location | 16,870,662 – 16,870,762 |
|---|---|
| Length | 100 |
| Sequences | 6 |
| Columns | 100 |
| Reading direction | forward |
| Mean pairwise identity | 74.60 |
| Mean single sequence MFE | -34.02 |
| Consensus MFE | -14.69 |
| Energy contribution | -14.42 |
| Covariance contribution | -0.27 |
| Combinations/Pair | 1.52 |
| Mean z-score | -2.10 |
| Structure conservation index | 0.43 |
| SVM decision value | 0.68 |
| SVM RNA-class probability | 0.820867 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
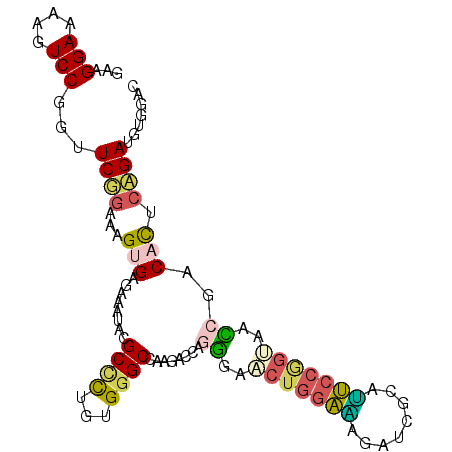
>3L_DroMel_CAF1 16870662 100 + 23771897 GAAGGAAAAGUCCGGUUCUGAUAGUGAGAAAUAUGCCUUGUGGGCCAAGACCAGGGAACUGGAAAGAUCACAUUCCGGUACCCGACACUCAGAUGUUGAU ...(((....)))...(((((..(((........(((.....)))........(((.(((((((........))))))).)))..))))))))....... ( -27.70) >DroPse_CAF1 61639 100 + 1 CAAGGAAAAGUCCGGUUCGGAGAGCGAGCGCUACGCCCUGUGGGCCAAGACACGGGAGCUGGAGAGGUCCAGCUCCCGGCAUCGACACACCGACGUGGAC .........(((((..((((.(.((((((.....((((...)))).......(((((((((((....))))))))))))).))).).).))))..))))) ( -46.20) >DroEre_CAF1 61056 100 + 1 GAAGGAAAAGUCCGGUUCUGAAAGUGAGAAGUACGCCUUGUGGGCCAAGACCAGGCAACUGGAAAGAUCGCAUUCCAGUACCAGGCACUCAGAAGUGGAC .........(((((.((((((..(((.....)))((((.((..(((.......))).(((((((........))))))))).))))..)))))).))))) ( -35.30) >DroYak_CAF1 60411 100 + 1 GAAGGAAAAGUCCGGUUCUGAAAGUGAGAAAUACGCUCUGUGGGCCAAGACCAGGGAACUGGAAAGAUCGCAUUCCAGUACCCGACACUCAGAUGUUGAC ...(((....)))...(((((.(((((....).))))..(((((......)).(((.(((((((........))))))).)))..))))))))....... ( -27.90) >DroMoj_CAF1 66888 100 + 1 CAAGGAAAAAUCCAGUUCGGAAAGCGAUCGCUAUGCGCUGUGGGCCAAGACCAGAGAGCUAACUAAAUCUGAAGAAGGCAAUAGGCAUAGAGACAUAGAC ...(((....)))...(((.....)))((.((((((.((((..(((.....((((.((....))...)))).....))).)))))))))).))....... ( -25.30) >DroAna_CAF1 58087 100 + 1 GAAGGAAAAGUCCGGCUCGGAGAGUGAGCGGUAUGCCCUCUGGGCCAAAACCAGGGAACUGGAGAGAUCUAGUUCCGCGCAUCGUCAUGCCGAUGUGGAU ...(((....)))(((((((((.((.........)).)))))))))........(((((((((....))))))))).(((((((......)))))))... ( -41.70) >consensus GAAGGAAAAGUCCGGUUCGGAAAGUGAGAAAUACGCCCUGUGGGCCAAGACCAGGGAACUGGAAAGAUCGCAUUCCGGUAACCGACACUCAGAUGUGGAC ...(((....)))...((((...(((........((((...))))........((..(((((((........)))))))..))..))).))))....... (-14.69 = -14.42 + -0.27)


| Location | 16,870,662 – 16,870,762 |
|---|---|
| Length | 100 |
| Sequences | 6 |
| Columns | 100 |
| Reading direction | reverse |
| Mean pairwise identity | 74.60 |
| Mean single sequence MFE | -32.42 |
| Consensus MFE | -13.62 |
| Energy contribution | -14.27 |
| Covariance contribution | 0.64 |
| Combinations/Pair | 1.43 |
| Mean z-score | -2.24 |
| Structure conservation index | 0.42 |
| SVM decision value | 0.96 |
| SVM RNA-class probability | 0.891108 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
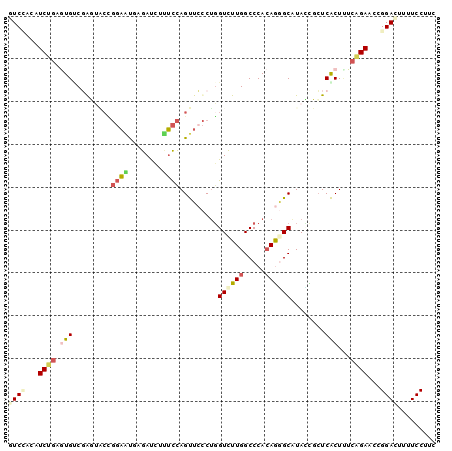
>3L_DroMel_CAF1 16870662 100 - 23771897 AUCAACAUCUGAGUGUCGGGUACCGGAAUGUGAUCUUUCCAGUUCCCUGGUCUUGGCCCACAAGGCAUAUUUCUCACUAUCAGAACCGGACUUUUCCUUC .......((((((((..(((.((.((((........)))).)).)))..((((((.....))))))........)))..)))))...(((....)))... ( -27.00) >DroPse_CAF1 61639 100 - 1 GUCCACGUCGGUGUGUCGAUGCCGGGAGCUGGACCUCUCCAGCUCCCGUGUCUUGGCCCACAGGGCGUAGCGCUCGCUCUCCGAACCGGACUUUUCCUUG ((((...((((((.(((((.(((((((((((((....))))))))))).)).))))).)))((((((.......)))))).)))...))))......... ( -50.20) >DroEre_CAF1 61056 100 - 1 GUCCACUUCUGAGUGCCUGGUACUGGAAUGCGAUCUUUCCAGUUGCCUGGUCUUGGCCCACAAGGCGUACUUCUCACUUUCAGAACCGGACUUUUCCUUC ((((..(((((((((...((((((((((........)))))).))))..((((((.....))))))........)))..))))))..))))......... ( -34.10) >DroYak_CAF1 60411 100 - 1 GUCAACAUCUGAGUGUCGGGUACUGGAAUGCGAUCUUUCCAGUUCCCUGGUCUUGGCCCACAGAGCGUAUUUCUCACUUUCAGAACCGGACUUUUCCUUC (((....((((((((..(((.(((((((........))))))).))).(((....)))))).(((.......)))....)))))....)))......... ( -30.50) >DroMoj_CAF1 66888 100 - 1 GUCUAUGUCUCUAUGCCUAUUGCCUUCUUCAGAUUUAGUUAGCUCUCUGGUCUUGGCCCACAGCGCAUAGCGAUCGCUUUCCGAACUGGAUUUUUCCUUG ......(((.((((((((...(((....(((((...((....)).)))))....)))....)).)))))).))).............(((....)))... ( -22.00) >DroAna_CAF1 58087 100 - 1 AUCCACAUCGGCAUGACGAUGCGCGGAACUAGAUCUCUCCAGUUCCCUGGUUUUGGCCCAGAGGGCAUACCGCUCACUCUCCGAGCCGGACUUUUCCUUC .(((.(((((......)))))...((((((.((....)).))))))..(((....((((...))))..)))((((.......)))).))).......... ( -30.70) >consensus GUCCACAUCUGAGUGUCGAGUACCGGAAUGAGAUCUUUCCAGUUCCCUGGUCUUGGCCCACAGGGCAUACCGCUCACUUUCAGAACCGGACUUUUCCUUC ((((...((((.(((.........((((........)))).........((((((.....))))))........)))...))))...))))......... (-13.62 = -14.27 + 0.64)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:15:27 2006