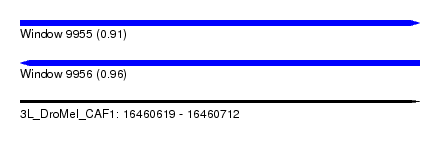
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 16,460,619 – 16,460,712 |
| Length | 93 |
| Max. P | 0.964348 |

| Location | 16,460,619 – 16,460,712 |
|---|---|
| Length | 93 |
| Sequences | 4 |
| Columns | 104 |
| Reading direction | forward |
| Mean pairwise identity | 81.85 |
| Mean single sequence MFE | -28.35 |
| Consensus MFE | -16.40 |
| Energy contribution | -18.15 |
| Covariance contribution | 1.75 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.60 |
| Structure conservation index | 0.58 |
| SVM decision value | 1.05 |
| SVM RNA-class probability | 0.906529 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
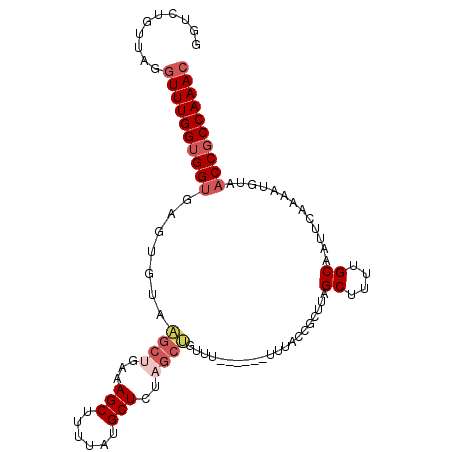
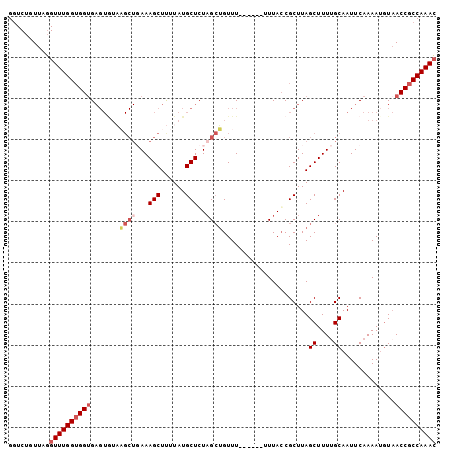
>3L_DroMel_CAF1 16460619 93 + 23771897 GGUCUGUUAGGUUUGGUGGUG-----------AAAGCUUUUAUGCUGUAGAUGUUUUUUUUUUUUACCGCUUAGCUUUUGCAAUUCAAAAUGUCACCGCCAAAC ..........(((((((((((-----------(..(((.....((.(((((...........))))).))..)))(((((.....)))))..)))))))))))) ( -25.90) >DroSec_CAF1 72119 97 + 1 GGUCUGUUAGGUUUGGUGGUGAGUGUAAGCUGAAAGCUUUUAUGCUCUAGCUGUUU-------UUACCGCUUAGCUUUUGCAAUUCAAAAUGUAACCGCCAAAC (((..((((.(((((((((((((....(((((..(((......))).)))))...)-------))))))))..((....))......)))).)))).))).... ( -28.90) >DroSim_CAF1 74334 98 + 1 GUUCUGUUAGGUUUGGUGGUGAGUGUAAGCUGAAAGCUUUUAUGCUCUAGCUGUUU------UUUAGCGCUUAGCUUUUGCAAUUCAAAAAGUAACCGCCAAAC ..........((((((((((((((((((((.....)))..)))))))..((((...------..)))).....((((((........)))))).)))))))))) ( -30.20) >DroYak_CAF1 75918 98 + 1 GGUCCGUUAUGUUUGGGGGAGAGUAUCCGCUCAAAGCUCUUAUGCUCUCGCGGUUU------UUUACCGCUUAGCUUUCGCAAUUCAAAAUUUAACCGCCAAAA (((..((((..((((((.((((((((..((.....))....))))))))(((((..------...)))))...((....))..))))))...)))).))).... ( -28.40) >consensus GGUCUGUUAGGUUUGGUGGUGAGUGUAAGCUGAAAGCUUUUAUGCUCUAGCUGUUU______UUUACCGCUUAGCUUUUGCAAUUCAAAAUGUAACCGCCAAAC ..........((((((((((.......((((...(((......)))..)))).....................((....)).............)))))))))) (-16.40 = -18.15 + 1.75)

| Location | 16,460,619 – 16,460,712 |
|---|---|
| Length | 93 |
| Sequences | 4 |
| Columns | 104 |
| Reading direction | reverse |
| Mean pairwise identity | 81.85 |
| Mean single sequence MFE | -23.05 |
| Consensus MFE | -15.15 |
| Energy contribution | -17.15 |
| Covariance contribution | 2.00 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.50 |
| Structure conservation index | 0.66 |
| SVM decision value | 1.57 |
| SVM RNA-class probability | 0.964348 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
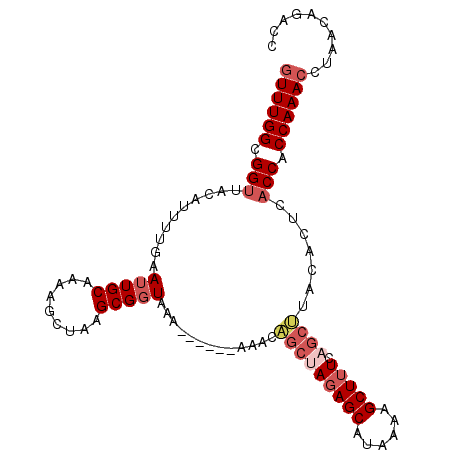
>3L_DroMel_CAF1 16460619 93 - 23771897 GUUUGGCGGUGACAUUUUGAAUUGCAAAAGCUAAGCGGUAAAAAAAAAAAACAUCUACAGCAUAAAAGCUUU-----------CACCACCAAACCUAACAGACC ((((((.(((((..((((..(((((.........)))))..)))).............(((......))).)-----------)))).)))))).......... ( -21.10) >DroSec_CAF1 72119 97 - 1 GUUUGGCGGUUACAUUUUGAAUUGCAAAAGCUAAGCGGUAA-------AAACAGCUAGAGCAUAAAAGCUUUCAGCUUACACUCACCACCAAACCUAACAGACC ((((((.(((..........(((((.........)))))..-------....(((((((((......))))).)))).......))).)))))).......... ( -22.20) >DroSim_CAF1 74334 98 - 1 GUUUGGCGGUUACUUUUUGAAUUGCAAAAGCUAAGCGCUAAA------AAACAGCUAGAGCAUAAAAGCUUUCAGCUUACACUCACCACCAAACCUAACAGAAC ((((((.(((....(((((.....)))))(.((((((((...------....))).(((((......)))))..))))))....))).)))))).......... ( -22.40) >DroYak_CAF1 75918 98 - 1 UUUUGGCGGUUAAAUUUUGAAUUGCGAAAGCUAAGCGGUAAA------AAACCGCGAGAGCAUAAGAGCUUUGAGCGGAUACUCUCCCCCAAACAUAACGGACC .(((((.((..............((....))...(((((...------..)))))(((((.((....((.....))..)).))))))))))))........... ( -26.50) >consensus GUUUGGCGGUUACAUUUUGAAUUGCAAAAGCUAAGCGGUAAA______AAACAGCUAGAGCAUAAAAGCUUUCAGCUUACACUCACCACCAAACCUAACAGACC ((((((.(((..........(((((.........))))).............(((((((((......))))).)))).......))).)))))).......... (-15.15 = -17.15 + 2.00)

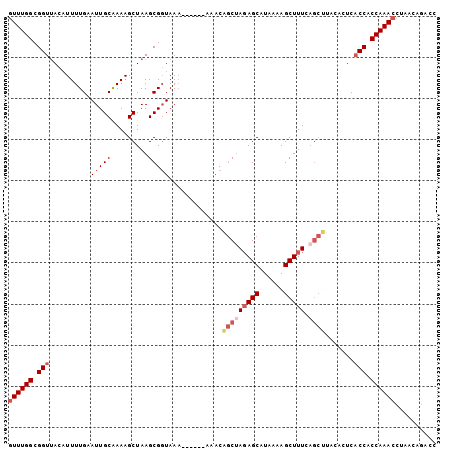
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:12:38 2006