| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 16,401,254 – 16,401,348 |
| Length | 94 |
| Max. P | 0.999283 |

| Location | 16,401,254 – 16,401,348 |
|---|---|
| Length | 94 |
| Sequences | 4 |
| Columns | 102 |
| Reading direction | forward |
| Mean pairwise identity | 82.22 |
| Mean single sequence MFE | -26.31 |
| Consensus MFE | -21.34 |
| Energy contribution | -22.59 |
| Covariance contribution | 1.25 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.67 |
| Structure conservation index | 0.81 |
| SVM decision value | 2.38 |
| SVM RNA-class probability | 0.993145 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
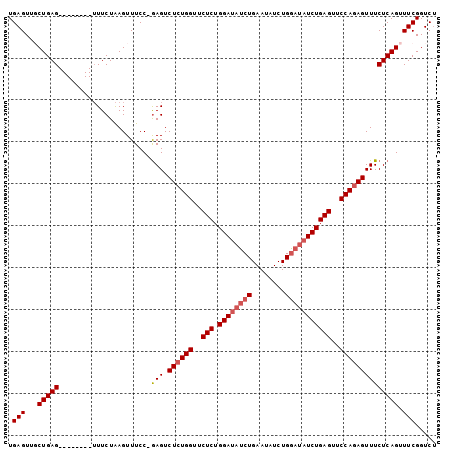
>3L_DroMel_CAF1 16401254 94 + 23771897 UGAGUUUCUGAG--------UUUCUAAGUUUCCAGAGUCUCUGGUUCUCUGGAUAUCUGAAUAUCUGACUAUCUGAGUUCCAGAGUUUCUCAGUUUCGGUCU .(((...(((((--------.((((........)))).((((((..(((.(((((((.(.....).)).))))))))..))))))...))))).)))..... ( -25.60) >DroSec_CAF1 16229 93 + 1 UGAGUUGCUGAG--------UUUCUAAGUUUCC-GAGUCUCUGGUUCUCUGGAUAUCUGAAUAUCUGGAUAUCUGAGUUCCAGAGUUUCUCAGUUUCGGUCU ...(..((((((--------.............-....((((((..(((.((((((((........)))))))))))..))))))...))))))..)..... ( -30.15) >DroSim_CAF1 16256 101 + 1 UGAGUUGCUGAGUUUCUGAGUUUCUAAGUUUCC-AAGUCUUUGGUUCUCUGGAUAUCUGAAUAUCUGGAUAUCUGAGUUCCACAGUUUCUCAGUUUCGGUCG ...(..((((((...(((.......(((.....-....)))(((..(((.((((((((........)))))))))))..))))))...))))))..)..... ( -27.10) >DroEre_CAF1 18017 77 + 1 UGAGUUUCUGAG--------AUUCUGAGUUUCC-GAGUCUCUGGUUCUCUGGAUA----------------UCUGAGCUCCAGAGUUUCUCAGUUUCGGUCU .......(((((--------(..(((((.....-(((.((((((..(((.((...----------------.)))))..))))))))))))))))))))... ( -22.40) >consensus UGAGUUGCUGAG________UUUCUAAGUUUCC_GAGUCUCUGGUUCUCUGGAUAUCUGAAUAUCUGGAUAUCUGAGUUCCAGAGUUUCUCAGUUUCGGUCU .(((...(((((......................(((.((((((..(((.((((((((........)))))))))))..)))))))))))))).)))..... (-21.34 = -22.59 + 1.25)


| Location | 16,401,254 – 16,401,348 |
|---|---|
| Length | 94 |
| Sequences | 4 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 82.22 |
| Mean single sequence MFE | -23.23 |
| Consensus MFE | -20.30 |
| Energy contribution | -21.30 |
| Covariance contribution | 1.00 |
| Combinations/Pair | 1.05 |
| Mean z-score | -3.51 |
| Structure conservation index | 0.87 |
| SVM decision value | 3.48 |
| SVM RNA-class probability | 0.999283 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
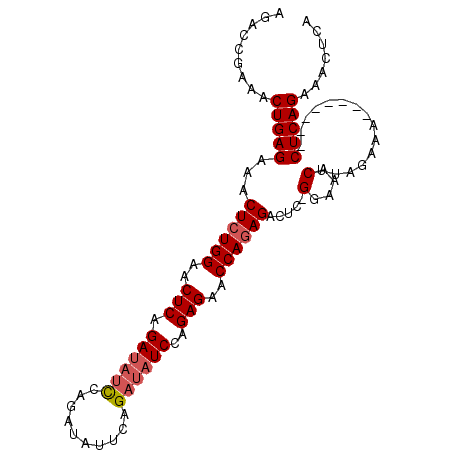
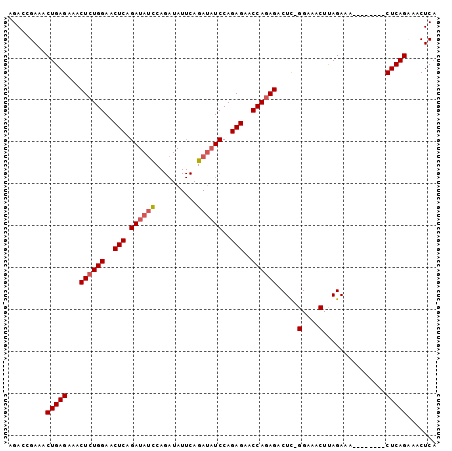
>3L_DroMel_CAF1 16401254 94 - 23771897 AGACCGAAACUGAGAAACUCUGGAACUCAGAUAGUCAGAUAUUCAGAUAUCCAGAGAACCAGAGACUCUGGAAACUUAGAAA--------CUCAGAAACUCA .........(((((...(((((((..((.((((......))))..))..)))))))..((((.....))))...........--------)))))....... ( -24.70) >DroSec_CAF1 16229 93 - 1 AGACCGAAACUGAGAAACUCUGGAACUCAGAUAUCCAGAUAUUCAGAUAUCCAGAGAACCAGAGACUC-GGAAACUUAGAAA--------CUCAGCAACUCA .........(((((...((((((..(((.((((((..........))))))..)))..))))))....-(....).......--------)))))....... ( -25.20) >DroSim_CAF1 16256 101 - 1 CGACCGAAACUGAGAAACUGUGGAACUCAGAUAUCCAGAUAUUCAGAUAUCCAGAGAACCAAAGACUU-GGAAACUUAGAAACUCAGAAACUCAGCAACUCA .........(((((...((.(((..(((.((((((..........))))))..)))..))).)).((.-(....)..))...)))))............... ( -19.90) >DroEre_CAF1 18017 77 - 1 AGACCGAAACUGAGAAACUCUGGAGCUCAGA----------------UAUCCAGAGAACCAGAGACUC-GGAAACUCAGAAU--------CUCAGAAACUCA .........((((((..((((((..(((...----------------......)))..))))))....-(....)......)--------)))))....... ( -23.10) >consensus AGACCGAAACUGAGAAACUCUGGAACUCAGAUAUCCAGAUAUUCAGAUAUCCAGAGAACCAGAGACUC_GGAAACUUAGAAA________CUCAGAAACUCA .........(((((...((((((..(((.((((((..........))))))..)))..)))))).....(....)...............)))))....... (-20.30 = -21.30 + 1.00)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:12:24 2006