
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 16,353,519 – 16,353,620 |
| Length | 101 |
| Max. P | 0.999952 |

| Location | 16,353,519 – 16,353,620 |
|---|---|
| Length | 101 |
| Sequences | 4 |
| Columns | 101 |
| Reading direction | forward |
| Mean pairwise identity | 68.15 |
| Mean single sequence MFE | -26.24 |
| Consensus MFE | -14.66 |
| Energy contribution | -17.60 |
| Covariance contribution | 2.94 |
| Combinations/Pair | 1.20 |
| Mean z-score | -3.39 |
| Structure conservation index | 0.56 |
| SVM decision value | 4.81 |
| SVM RNA-class probability | 0.999952 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
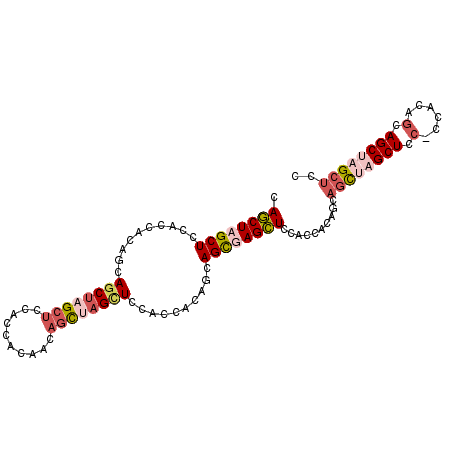
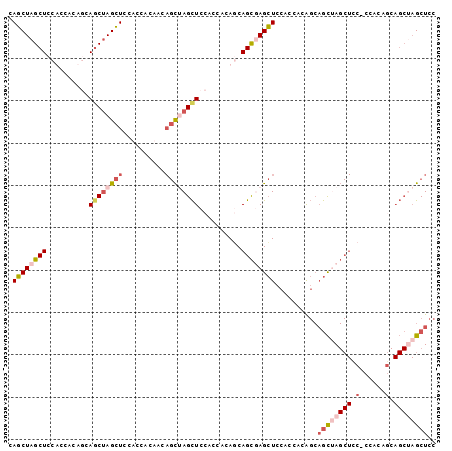
>3L_DroMel_CAF1 16353519 101 + 23771897 CAGCUAGCUCCACCACACCAGCUAGCUCCACCACACCAGCUAGCUCCACCACAGCAGCGAGCUCCGCCACAGCAGCUAGCUCCGCCACAGCAGCUAGCUCC .((((((((..........((((((((..........((((.(((.(......).))).))))..((....))))))))))..((....)))))))))).. ( -31.90) >DroSec_CAF1 36179 101 + 1 CAGCUAGCUCCUCCUCAGCAACGAGCUCCUCCACAACAGCUAGCUCCUCCACAACAGCGAGCUCCACCACAGCAGCUAGCUCCACCACAACAGCGAGCUCC ..(((((......)).)))...((((((((.......((((((((.((.......((....)).......)).))))))))..........)).)))))). ( -23.57) >DroSim_CAF1 39765 101 + 1 CAGCUAGCUCCUCCACAACAGCUAGCUUCUCCACAACAGCGAGCUCCACCACAGCAGCUAGCUCCACCGCAGCAGCGAGCUUCCCCACAGCAGCUAGCUCC .((((((((.((.......((((.(((.((.......(((.((((.(......).)))).))).......)).))).)))).......)).)))))))).. ( -28.88) >DroEre_CAF1 36208 77 + 1 CAUCUAGCUCCACCGCAGCAGCUAGGUACACCUAGA---UCAGUUCCACAACACCAGUUAGGUAUACCGAG---AUCA------------------GCUGC ......((......)).((((((.((((.((((((.---...(((....))).....)))))).))))(..---..))------------------))))) ( -20.60) >consensus CAGCUAGCUCCACCACAGCAGCUAGCUCCACCACAACAGCUAGCUCCACCACAGCAGCGAGCUCCACCACAGCAGCUAGCUCC_CCACAGCAGCUAGCUCC .((((((((..........((((((((..........))))))))..........))))))))..........((((((((.(......).)))))))).. (-14.66 = -17.60 + 2.94)


| Location | 16,353,519 – 16,353,620 |
|---|---|
| Length | 101 |
| Sequences | 4 |
| Columns | 101 |
| Reading direction | reverse |
| Mean pairwise identity | 68.15 |
| Mean single sequence MFE | -38.45 |
| Consensus MFE | -20.81 |
| Energy contribution | -22.88 |
| Covariance contribution | 2.06 |
| Combinations/Pair | 1.32 |
| Mean z-score | -2.64 |
| Structure conservation index | 0.54 |
| SVM decision value | 3.45 |
| SVM RNA-class probability | 0.999238 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
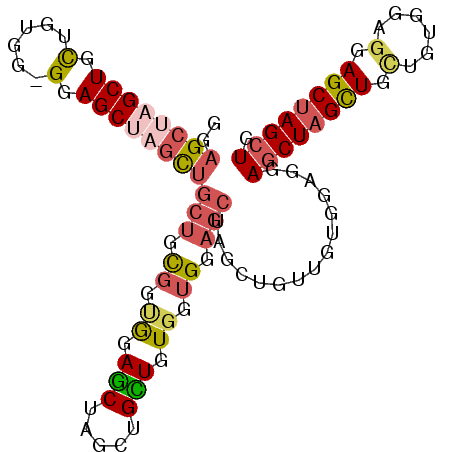
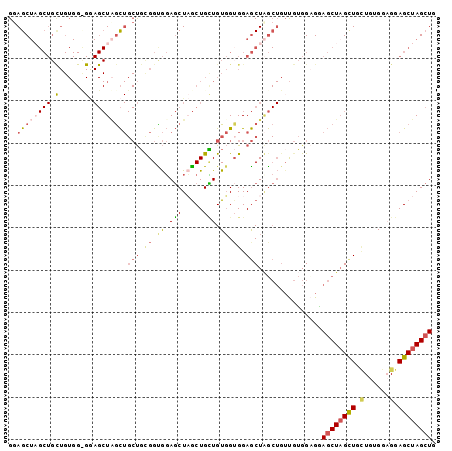
>3L_DroMel_CAF1 16353519 101 - 23771897 GGAGCUAGCUGCUGUGGCGGAGCUAGCUGCUGUGGCGGAGCUCGCUGCUGUGGUGGAGCUAGCUGGUGUGGUGGAGCUAGCUGGUGUGGUGGAGCUAGCUG ..((((((((.(..(.(((.((((((((.(..(.(((.((((.(((.(......).))).))))..))).)..)))))))))..))).)..))))))))). ( -47.80) >DroSec_CAF1 36179 101 - 1 GGAGCUCGCUGUUGUGGUGGAGCUAGCUGCUGUGGUGGAGCUCGCUGUUGUGGAGGAGCUAGCUGUUGUGGAGGAGCUCGUUGCUGAGGAGGAGCUAGCUG .((((((((..(.(..((.......))..).)..)).)))))).............((((((((.((.(..((.((....)).))..).)).)))))))). ( -35.00) >DroSim_CAF1 39765 101 - 1 GGAGCUAGCUGCUGUGGGGAAGCUCGCUGCUGCGGUGGAGCUAGCUGCUGUGGUGGAGCUCGCUGUUGUGGAGAAGCUAGCUGUUGUGGAGGAGCUAGCUG ..((((((((.(((((((....)))))..(..((..(.((((((((.(((..(..(......)..)..)))...)))))))).)))..))).)))))))). ( -44.50) >DroEre_CAF1 36208 77 - 1 GCAGC------------------UGAU---CUCGGUAUACCUAACUGGUGUUGUGGAACUGA---UCUAGGUGUACCUAGCUGCUGCGGUGGAGCUAGAUG (((((------------------((..---...(((((((((((..(((........)))..---).))))))))))))))))).((......))...... ( -26.50) >consensus GGAGCUAGCUGCUGUGG_GGAGCUAGCUGCUGCGGUGGAGCUAGCUGCUGUGGUGGAGCUAGCUGUUGUGGAGGAGCUAGCUGCUGUGGAGGAGCUAGCUG ..((((((((.(......).))))))))(((.((.((.(((.....))).)).)).)))...............((((((((.(......).)))))))). (-20.81 = -22.88 + 2.06)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:11:48 2006