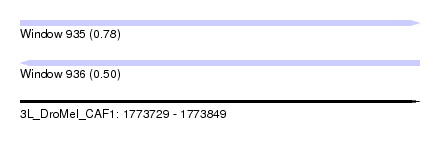

| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 1,773,729 – 1,773,849 |
| Length | 120 |
| Max. P | 0.782815 |

| Location | 1,773,729 – 1,773,849 |
|---|---|
| Length | 120 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 83.56 |
| Mean single sequence MFE | -46.43 |
| Consensus MFE | -34.95 |
| Energy contribution | -34.98 |
| Covariance contribution | 0.03 |
| Combinations/Pair | 1.23 |
| Mean z-score | -2.33 |
| Structure conservation index | 0.75 |
| SVM decision value | 0.56 |
| SVM RNA-class probability | 0.782815 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
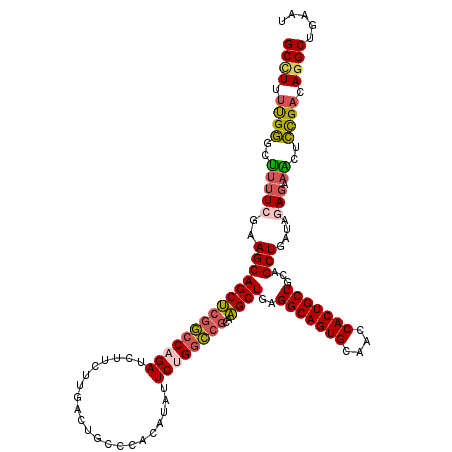
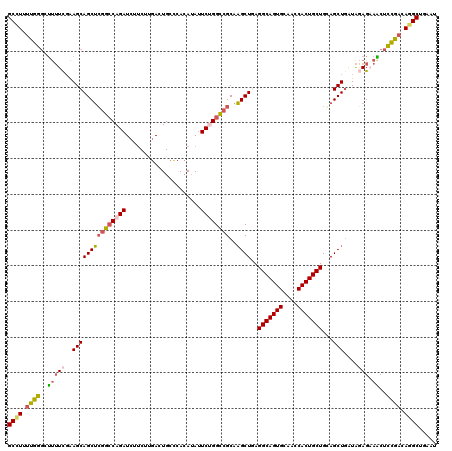
>3L_DroMel_CAF1 1773729 120 + 23771897 GCCUUUUGGGCUUUUCGAAGCAGCUCGGCCAGAUCUUCUUGACUGCCCACAUAUUCUGGCCGCAAGCUGAGGCAGUGCAACCACUGCUGGAGCUGAUAGAGAAACUCCGACAGGCUGAAU ((((.(((((.(((((....(((((((((((((......((......)).....))))))).........(((((((....))))))).))))))...))))).).)))).))))..... ( -49.20) >DroVir_CAF1 11981 120 + 1 GCCUGCCGCGUGCGUUUGAGCAGCGGCAGCAGAUCGCCCAAAUUGGGCAGAAAUUCGGGUCGCAUGCUGAGGCAGUGCACCCACUGCUGCAGCUCGUGUAGAAUCUGCGACAAGCUGAGC ((.((((((.(((......))))))))))).....((((.....)))).....(((((((((((.((((.(((((((....))))))).))))((.....))...))))))...))))). ( -54.70) >DroSec_CAF1 11038 120 + 1 GCUUUUUGGGCUUUUCGAAGCAGCUCGGCCAGAUCUUCUUGACUGCCCACAUAUUCUGGCCGCAAGCUGAGGCAGUGCAACCACUGCUGGAGCUGAUAGAGAAACUCCGACAGGCUGAAU ((((.(((((.(((((....(((((((((((((......((......)).....))))))).........(((((((....))))))).))))))...))))).).)))).))))..... ( -46.50) >DroSim_CAF1 11005 120 + 1 GCUUUUUGGGCUUUUCGAAGCAGCUCGGCCAGAUCUUCUUGACUGCCCACAUAUUCUGGCCGCAAGCUGAGGCAGUGCAACCACUGCUGGAGCUGAUAGAGAAACUCCGACAGGCUGAAU ((((.(((((.(((((....(((((((((((((......((......)).....))))))).........(((((((....))))))).))))))...))))).).)))).))))..... ( -46.50) >DroEre_CAF1 11010 120 + 1 GCCUUUUGGCAUUUUUGAAGCAGCUCGGCCAGAUCUUCACGACUGCCCACAUAUUCUGGCCGCAAGCUUAGGCAGUGCAGCCACUGCUGCAGCUGAUACAGCAGCUCUGACAGGCUGAAU ((((.(..(..........((((((((((((((.....................))))))))..))))..(((((((....)))))))))(((((......))))))..).))))..... ( -42.20) >DroYak_CAF1 10931 120 + 1 GCCUUUUGGGAUUUUCGAAGCAGCUCGGCCAGAUCUUCACGACUGCUCACAUAAUCGGGCCGCAAGCUCAGGCAGUGUAGCCACUGCUGCAGCUGAUAGAGCAGCUCUGACAGGCUGAAU ((((.(..(((((.....(((((.(((............)))))))).....))))(((.(....)))).(((((((....)))))))..(((((......))))))..).))))..... ( -39.50) >consensus GCCUUUUGGGCUUUUCGAAGCAGCUCGGCCAGAUCUUCUUGACUGCCCACAUAUUCUGGCCGCAAGCUGAGGCAGUGCAACCACUGCUGCAGCUGAUAGAGAAACUCCGACAGGCUGAAU ((((.((((..(((((..(((((((((((((((.....................))))))))..))))..(((((((....)))))))...)))....))).))..)))).))))..... (-34.95 = -34.98 + 0.03)

| Location | 1,773,729 – 1,773,849 |
|---|---|
| Length | 120 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 83.56 |
| Mean single sequence MFE | -44.77 |
| Consensus MFE | -29.50 |
| Energy contribution | -30.95 |
| Covariance contribution | 1.45 |
| Combinations/Pair | 1.16 |
| Mean z-score | -1.91 |
| Structure conservation index | 0.66 |
| SVM decision value | -0.07 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
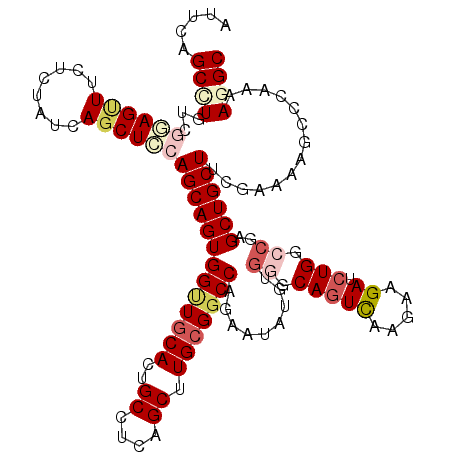
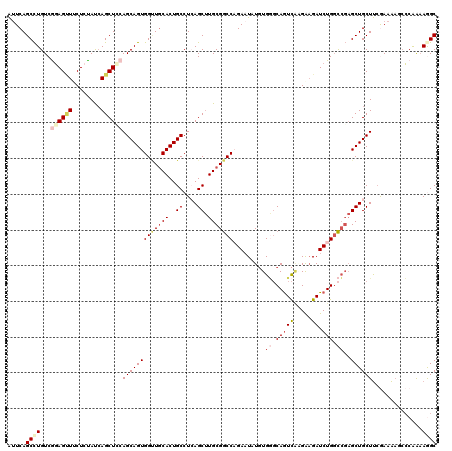
>3L_DroMel_CAF1 1773729 120 - 23771897 AUUCAGCCUGUCGGAGUUUCUCUAUCAGCUCCAGCAGUGGUUGCACUGCCUCAGCUUGCGGCCAGAAUAUGUGGGCAGUCAAGAAGAUCUGGCCGAGCUGCUUCGAAAAGCCCAAAAGGC .....((((...((.((((.((...((((((..((((((....))))))..........(((((((.....(((....)))......)))))))))))))....)).))))))...)))) ( -43.00) >DroVir_CAF1 11981 120 - 1 GCUCAGCUUGUCGCAGAUUCUACACGAGCUGCAGCAGUGGGUGCACUGCCUCAGCAUGCGACCCGAAUUUCUGCCCAAUUUGGGCGAUCUGCUGCCGCUGCUCAAACGCACGCGGCAGGC .....(((((((((............(((.(((((((((((((((.(((....)))))).)))))......(((((.....)))))..))))))).)))((......))..))))))))) ( -54.40) >DroSec_CAF1 11038 120 - 1 AUUCAGCCUGUCGGAGUUUCUCUAUCAGCUCCAGCAGUGGUUGCACUGCCUCAGCUUGCGGCCAGAAUAUGUGGGCAGUCAAGAAGAUCUGGCCGAGCUGCUUCGAAAAGCCCAAAAAGC .....((...(((((((..(((....((((...((((((....))))))...))))...(((((((.....(((....)))......))))))))))..)))))))...))......... ( -41.30) >DroSim_CAF1 11005 120 - 1 AUUCAGCCUGUCGGAGUUUCUCUAUCAGCUCCAGCAGUGGUUGCACUGCCUCAGCUUGCGGCCAGAAUAUGUGGGCAGUCAAGAAGAUCUGGCCGAGCUGCUUCGAAAAGCCCAAAAAGC .....((...(((((((..(((....((((...((((((....))))))...))))...(((((((.....(((....)))......))))))))))..)))))))...))......... ( -41.30) >DroEre_CAF1 11010 120 - 1 AUUCAGCCUGUCAGAGCUGCUGUAUCAGCUGCAGCAGUGGCUGCACUGCCUAAGCUUGCGGCCAGAAUAUGUGGGCAGUCGUGAAGAUCUGGCCGAGCUGCUUCAAAAAUGCCAAAAGGC .....((((.((...(((((((((.....)))))))))(((((((..((....)).))))))).)).......((((....(((((....(((...))).)))))....))))...)))) ( -45.60) >DroYak_CAF1 10931 120 - 1 AUUCAGCCUGUCAGAGCUGCUCUAUCAGCUGCAGCAGUGGCUACACUGCCUGAGCUUGCGGCCCGAUUAUGUGAGCAGUCGUGAAGAUCUGGCCGAGCUGCUUCGAAAAUCCCAAAAGGC .....((((.((..(((.((((....((((.((((((((....))))).)))))))...((((.(((((((........)))...)))).)))))))).)))..))..........)))) ( -43.00) >consensus AUUCAGCCUGUCGGAGUUUCUCUAUCAGCUCCAGCAGUGGUUGCACUGCCUCAGCUUGCGGCCAGAAUAUGUGGGCAGUCAAGAAGAUCUGGCCGAGCUGCUUCGAAAAGCCCAAAAGGC .....((((...((((((........))))))(((((((((((((..((....)).))))))).........((.(((((.....)).))).))..))))))..............)))) (-29.50 = -30.95 + 1.45)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:46:39 2006