
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 16,155,448 – 16,155,608 |
| Length | 160 |
| Max. P | 0.973478 |

| Location | 16,155,448 – 16,155,568 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 89.58 |
| Mean single sequence MFE | -38.04 |
| Consensus MFE | -32.19 |
| Energy contribution | -32.44 |
| Covariance contribution | 0.25 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.60 |
| Structure conservation index | 0.85 |
| SVM decision value | 0.41 |
| SVM RNA-class probability | 0.726713 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 16155448 120 + 23771897 GCGAGUGGCAAGCGGAUCGGCCGGACUCUCUGACCCGUCUGCGAUUGCGAAUCCAGCUGUGGGACUGUACUUGGAUAAAUAAGAACUAAAGUACAUACUUUAGCAGCAAGAAAGAUAAUU ((((.(.(((.((((((.((.(((.....))).)).))))))...))).).))..((((.((...(((((((................)))))))..)).)))).))............. ( -32.99) >DroSec_CAF1 12793 120 + 1 GCGAGUGGCAAGCGGAUCGGCCGGACUCGCUGACCCGUCUGCGAUUGCAGAUCCAGCUGUGGGACUGUACUUGGAUAAAUAAGAACUAAAGUACAUACUAAAGCAGCAAGAAAGAUAAUU ((.(((.(((..(((.(((((.......))))).)))..))).))))).......(((((.((..(((((((................)))))))..))...)))))............. ( -36.99) >DroSim_CAF1 14162 120 + 1 GCGAGUGGCAAGCGGAUCGGCCGGACUCGCUGACCCGUCGGCGAUUGCGGAUCCAGCUGUGGGACUGUACUUGGAUAAAUAAGAACUAAAGUACAUACUUAAGCAGCAAGAAAGAUAAUU ((.....))....(((((.((.....(((((((....)))))))..)).))))).(((((.((..(((((((................)))))))..))...)))))............. ( -36.59) >DroEre_CAF1 12471 116 + 1 GCGAGUGGCAAGCGGAUCGGCCGGACUCGCUGACCCGUCUGCGAUUGCGAAUCCAGCUCCGUGACUGUACUUGGGUAAAUAAAAACUAAAGUG----CUUCAGGAGCAGGAAAUGCGUUC ...(((.(((..(((.(((((.......))))).)))..))).)))(((..(((.(((((.(((..((((((.(((........))).)))))----).)))))))).)))..))).... ( -45.60) >consensus GCGAGUGGCAAGCGGAUCGGCCGGACUCGCUGACCCGUCUGCGAUUGCGAAUCCAGCUGUGGGACUGUACUUGGAUAAAUAAGAACUAAAGUACAUACUUAAGCAGCAAGAAAGAUAAUU ((.(((.(((.((((.(((((.......))))).)))).))).))))).......(((((.....(((((((................))))))).......)))))............. (-32.19 = -32.44 + 0.25)

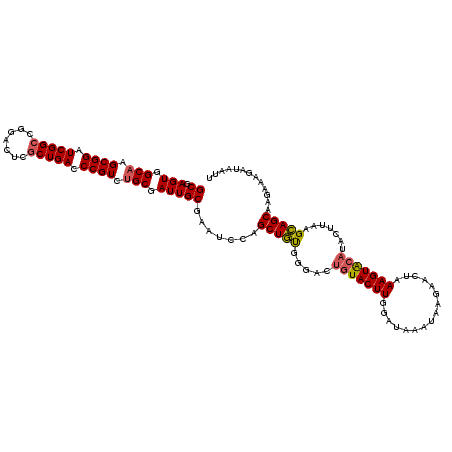

| Location | 16,155,488 – 16,155,608 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 71.67 |
| Mean single sequence MFE | -24.19 |
| Consensus MFE | -13.43 |
| Energy contribution | -12.89 |
| Covariance contribution | -0.54 |
| Combinations/Pair | 1.32 |
| Mean z-score | -2.11 |
| Structure conservation index | 0.56 |
| SVM decision value | 1.71 |
| SVM RNA-class probability | 0.973478 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 16155488 120 + 23771897 GCGAUUGCGAAUCCAGCUGUGGGACUGUACUUGGAUAAAUAAGAACUAAAGUACAUACUUUAGCAGCAAGAAAGAUAAUUAUGUGAAAUCAAGUUAAAUAUCAUCAAUCAUAAAUAUAUA ..(((((........(((((.((..(((((((................)))))))..))...)))))......((((.(((..((....))...))).))))..)))))........... ( -22.19) >DroSec_CAF1 12833 120 + 1 GCGAUUGCAGAUCCAGCUGUGGGACUGUACUUGGAUAAAUAAGAACUAAAGUACAUACUAAAGCAGCAAGAAAGAUAAUUAUGUGAAAUCAAGUUUAAUAUCAUCAAUCAUAAAUAUAUA ..(((((........(((((.((..(((((((................)))))))..))...)))))......((((.(((..((....))....)))))))..)))))........... ( -21.59) >DroEre_CAF1 12511 96 + 1 GCGAUUGCGAAUCCAGCUCCGUGACUGUACUUGGGUAAAUAAAAACUAAAGUG----CUUCAGGAGCAGGAAAUGCGUUCAUUUA-CAUC-------------------UAAAAUAUUAU ..((.((((..(((.(((((.(((..((((((.(((........))).)))))----).)))))))).)))..)))).)).....-....-------------------........... ( -28.80) >consensus GCGAUUGCGAAUCCAGCUGUGGGACUGUACUUGGAUAAAUAAGAACUAAAGUACAUACUUAAGCAGCAAGAAAGAUAAUUAUGUGAAAUCAAGUU_AAUAUCAUCAAUCAUAAAUAUAUA ..(((((....((..(((((.....(((((((................))))))).......)))))..))....)))))........................................ (-13.43 = -12.89 + -0.54)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:09:16 2006