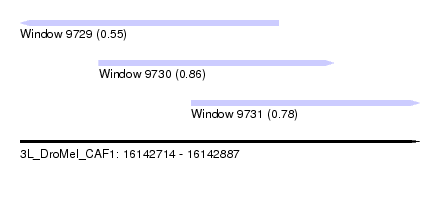
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 16,142,714 – 16,142,887 |
| Length | 173 |
| Max. P | 0.862437 |

| Location | 16,142,714 – 16,142,826 |
|---|---|
| Length | 112 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 80.19 |
| Mean single sequence MFE | -36.76 |
| Consensus MFE | -22.36 |
| Energy contribution | -23.08 |
| Covariance contribution | 0.72 |
| Combinations/Pair | 1.16 |
| Mean z-score | -1.72 |
| Structure conservation index | 0.61 |
| SVM decision value | 0.03 |
| SVM RNA-class probability | 0.547740 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 16142714 112 - 23771897 --CGGCUCUCCCCUCAUCACCACGAAUGCUCUCCACAUCGUCAGUGGCCGCCGUCUGGCGGGGAGAGCUGUAAUCGUGGGCACACCGAAAGAAU------AUAUUCCCCAAUGAUCUGAA --(((((((((((......((((..(((..........)))..))))..(((....))))))))))))))..((((((((.....(....)...------......))).)))))..... ( -38.04) >DroSec_CAF1 4905 104 - 1 --CGGCUCUCCCCUC--------GAAUGCUCUUCACAUCGUCAGUGGCCGCCGUCUGGCGGGGAGAGCUGUAAUCGUGGGCACACCGAAAGAAU------AUAUUCCCCAAUGAUCUGAA --(((((((((((..--------(((.....))).....((((((((...))).))))))))))))))))..((((((((.....(....)...------......))).)))))..... ( -37.04) >DroSim_CAF1 5689 112 - 1 --CGGCUCUCCCCUCGCCACCACGAAUGCUCUCCACAUCGUCAGUGGCCGCCGUCUGGCGGGGAGAGCUGUAAUCGUGGGCACACCGAAAGAAU------AUAUUCCCCAAUGAUCUGAA --(((((((((((..(((((.((((.((.......))))))..))))).(((....))))))))))))))..((((((((.....(....)...------......))).)))))..... ( -44.04) >DroEre_CAF1 4268 111 - 1 CCCGAUUCUCCCCUCUUC-CCACGGAUGCUCCACACAUUGG--AGUGCCGCCGUCUGGCG--GAGAGCUGUAAUCGUGGGCACACCGAAAGAAUACUUGUAUAUUCCCCACUGAUC---- ...((((..(..((((((-((((((.(((((((.....)))--))))...))))..)).)--)))))..).))))(((((.....(....).(((....)))....))))).....---- ( -32.60) >DroYak_CAF1 4957 108 - 1 CCAG-CGCUCCCCUCUUU-CCGUGGAUGCUCCACUCAUUGG--GGAGCCGUCGUCUGGCA--GAGAGCUUUAAUCGUGGGCACACCAAAAGAAU------AUAUUCCCCAUUGAUCUGGU ((((-.............-..((((.....))))(((.(((--(((((((.....)))).--...........((.(((.....)))...))..------....)))))).))).)))). ( -32.10) >consensus __CGGCUCUCCCCUCUUC_CCACGAAUGCUCUACACAUCGUCAGUGGCCGCCGUCUGGCGGGGAGAGCUGUAAUCGUGGGCACACCGAAAGAAU______AUAUUCCCCAAUGAUCUGAA ..(((((((((((.........(((.((.......)))))......((((.....)))))))))))))))..((((((((..........((((........)))))))).))))..... (-22.36 = -23.08 + 0.72)


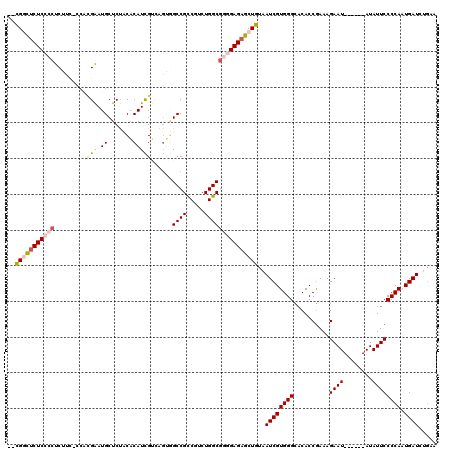
| Location | 16,142,748 – 16,142,850 |
|---|---|
| Length | 102 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 79.85 |
| Mean single sequence MFE | -43.20 |
| Consensus MFE | -27.49 |
| Energy contribution | -28.55 |
| Covariance contribution | 1.06 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.06 |
| Structure conservation index | 0.64 |
| SVM decision value | 0.83 |
| SVM RNA-class probability | 0.862437 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 16142748 102 + 23771897 CCCACGAUUACAGCUCUCCCCGCCAGACGGCGGCCACUGACGAUGUGGAGAGCAUUCGUGGUGAUGAGGGGAGAGCCG--------------GGAGACUGCAGACAGGAGACUGGA---- ............((((((((((((....)))...((((.(((((((.....))).))))))))....)))))))))((--------------(....)))....(((....)))..---- ( -44.10) >DroSec_CAF1 4939 94 + 1 CCCACGAUUACAGCUCUCCCCGCCAGACGGCGGCCACUGACGAUGUGAAGAGCAUUC--------GAGGGGAGAGCCG--------------GGAGACUGCAGACAGGAGACUGGA---- ............((((((((((((....))).........((((((.....))).))--------).)))))))))((--------------(....)))....(((....)))..---- ( -38.50) >DroSim_CAF1 5723 102 + 1 CCCACGAUUACAGCUCUCCCCGCCAGACGGCGGCCACUGACGAUGUGGAGAGCAUUCGUGGUGGCGAGGGGAGAGCCG--------------GGAGACUGCAGACAGGAGACUGGA---- ............((((((((((((....))).((((((.(((((((.....))).))))))))))..)))))))))((--------------(....)))....(((....)))..---- ( -51.10) >DroEre_CAF1 4304 115 + 1 CCCACGAUUACAGCUCUC--CGCCAGACGGCGGCACU--CCAAUGUGUGGAGCAUCCGUGG-GAAGAGGGGAGAAUCGGGUGAUUGGAGACAGGAGACUGGAGACAGGAGACAGGAGACC .((.(((((((..(((((--(......(.((((..((--(((.....)))))...)))).)-.....))))))......)))))))....(((....)))......(....).))..... ( -39.10) >consensus CCCACGAUUACAGCUCUCCCCGCCAGACGGCGGCCACUGACGAUGUGGAGAGCAUUCGUGG_GA_GAGGGGAGAGCCG______________GGAGACUGCAGACAGGAGACUGGA____ ............((((((((((((....)))..........(((((.....)))))...........)))))))))................(....)......(((....)))...... (-27.49 = -28.55 + 1.06)

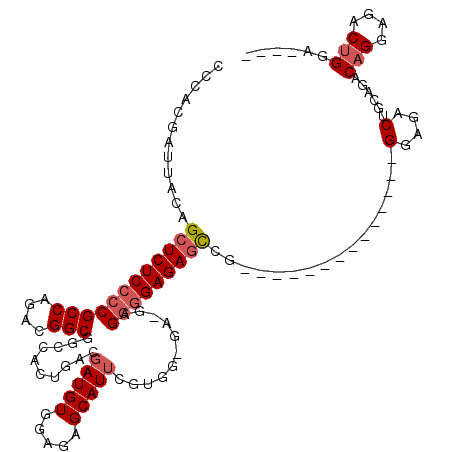
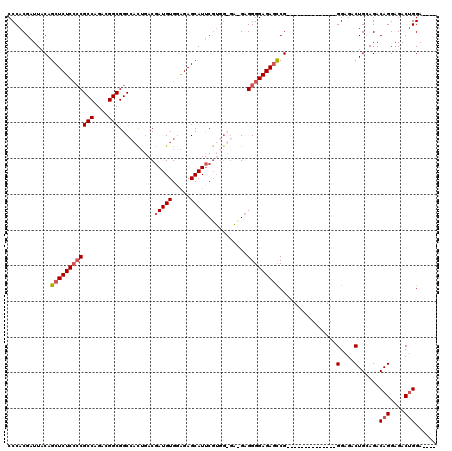
| Location | 16,142,788 – 16,142,887 |
|---|---|
| Length | 99 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 80.03 |
| Mean single sequence MFE | -31.85 |
| Consensus MFE | -20.41 |
| Energy contribution | -21.47 |
| Covariance contribution | 1.06 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.92 |
| Structure conservation index | 0.64 |
| SVM decision value | 0.55 |
| SVM RNA-class probability | 0.779830 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 16142788 99 + 23771897 CGAUGUGGAGAGCAUUCGUGGUGAUGAGGGGAGAGCCG--------------GGAGACUGCAGACAGGAGACUGGA-------GACAGGAGACAGGCGACCGGUCGCAAAGUGGCUCAAU .........((((...(...(((((..((.....((((--------------(....)).....(((....)))..-------....(....).)))..)).)))))...)..))))... ( -30.90) >DroSec_CAF1 4979 87 + 1 CGAUGUGAAGAGCAUUC--------GAGGGGAGAGCCG--------------GGAGACUGCAGACAGGAGACUGGA-------GACAGGAGACAGGC----GGUCGCAAAGUGGCUCAAU ((((((.....))).))--------)......((((((--------------.(.((((((...(((....)))..-------....(....)..))----)))).)....))))))... ( -30.80) >DroSim_CAF1 5763 99 + 1 CGAUGUGGAGAGCAUUCGUGGUGGCGAGGGGAGAGCCG--------------GGAGACUGCAGACAGGAGACUGGA-------GACAGGAGACAGGCGACCGGUCGCAAAGUGGCUCAAU .........((((...(...(((((..((.....((((--------------(....)).....(((....)))..-------....(....).)))..)).)))))...)..))))... ( -31.90) >DroEre_CAF1 4340 119 + 1 CAAUGUGUGGAGCAUCCGUGG-GAAGAGGGGAGAAUCGGGUGAUUGGAGACAGGAGACUGGAGACAGGAGACAGGAGACCGGAGACAGGAGACAGGCGACCGGUCGCAAAGUGGCUCAAU .........((((.(((....-..........(..((.(((..(((....)))...))).))..).(....).)))((((((.(...(....)...)..))))))........))))... ( -33.80) >consensus CGAUGUGGAGAGCAUUCGUGG_GA_GAGGGGAGAGCCG______________GGAGACUGCAGACAGGAGACUGGA_______GACAGGAGACAGGCGACCGGUCGCAAAGUGGCUCAAU .(((((.....)))))................(((((...............(....)......(((....)))..........((.(....)..((((....))))...)))))))... (-20.41 = -21.47 + 1.06)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:08:55 2006