
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 16,120,286 – 16,120,442 |
| Length | 156 |
| Max. P | 0.843246 |

| Location | 16,120,286 – 16,120,402 |
|---|---|
| Length | 116 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 93.33 |
| Mean single sequence MFE | -19.74 |
| Consensus MFE | -15.90 |
| Energy contribution | -15.90 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.26 |
| Structure conservation index | 0.81 |
| SVM decision value | 0.12 |
| SVM RNA-class probability | 0.594265 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
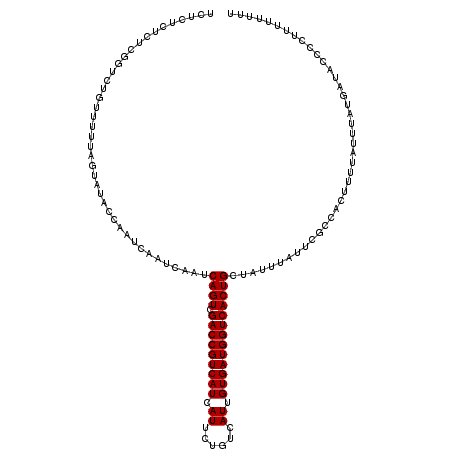
>3L_DroMel_CAF1 16120286 116 + 23771897 UCUCUCUCUCUUGCUGUUUUUAGUAUACCAAUCAAUCAAUCAGUCGACCGUCAUCAUUCUGUCAUCGUGAUGGUCACUGCUAUUUAUUCGCCACUUUUAUUUAUGAUACCCUUU----UU ...........(((((....)))))...............((((.(((((((((.((......)).)))))))))))))...................................----.. ( -18.30) >DroSec_CAF1 6009 120 + 1 UCUCUCUCUCGGUCUGUUUUUAGUAUAGCAAUCAAUCAAUCAGUCGACCGUCAUCAUUCUGUCAUUGUGAUGGUCACUGCUAUUUAUUAGCCACUUUUAUUUAUGAUACCCCCUUUUUUU ..........(((.((((........))))....((((...(((.(((((((((.((......)).))))))))))))((((.....))))............))))))).......... ( -21.60) >DroSim_CAF1 5365 120 + 1 UCUCUCUCUCGGUCUGUUUUUAGUAUACCAAUCAAUCAAUCAGUCGACCGUCAUCAUUCUGUCAUUGUGAUGGUCACUGCUAUUUAUUCGCCACUUUUAUUUAUGAUACCCCUUUUUUUU ..........((..((((...((((........(((.(((((((.(((((((((.((......)).)))))))))))))..))).))).........))))...))))..))........ ( -19.33) >consensus UCUCUCUCUCGGUCUGUUUUUAGUAUACCAAUCAAUCAAUCAGUCGACCGUCAUCAUUCUGUCAUUGUGAUGGUCACUGCUAUUUAUUCGCCACUUUUAUUUAUGAUACCCCUUUUUUUU ........................................((((.(((((((((.((......)).)))))))))))))......................................... (-15.90 = -15.90 + 0.00)



| Location | 16,120,326 – 16,120,442 |
|---|---|
| Length | 116 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 93.33 |
| Mean single sequence MFE | -19.25 |
| Consensus MFE | -16.83 |
| Energy contribution | -17.16 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.36 |
| Structure conservation index | 0.87 |
| SVM decision value | 0.76 |
| SVM RNA-class probability | 0.843246 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 16120326 116 + 23771897 CAGUCGACCGUCAUCAUUCUGUCAUCGUGAUGGUCACUGCUAUUUAUUCGCCACUUUUAUUUAUGAUACCCUUU----UUUUUUCUUCGAUUUUUGCCAUAUUUAGCUUUUAUCUAUCGC ((((.(((((((((.((......)).)))))))))))))...................................----..........(((....((........))....)))...... ( -18.00) >DroSec_CAF1 6049 120 + 1 CAGUCGACCGUCAUCAUUCUGUCAUUGUGAUGGUCACUGCUAUUUAUUAGCCACUUUUAUUUAUGAUACCCCCUUUUUUUUUUUUUGCGAUUUUAACCAUAUUUAGCUUUUAUCUAUCGC ..((((((((((((.((......)).)))))))))...((((.....)))).............)))...................(((((........................))))) ( -19.86) >DroSim_CAF1 5405 120 + 1 CAGUCGACCGUCAUCAUUCUGUCAUUGUGAUGGUCACUGCUAUUUAUUCGCCACUUUUAUUUAUGAUACCCCUUUUUUUUUUUUUUGCGAUUUUUGCCAUAUUUAGCUUUUAUCUAUCGC ((((.(((((((((.((......)).)))))))))))))...............................................(((((....((........))........))))) ( -19.90) >consensus CAGUCGACCGUCAUCAUUCUGUCAUUGUGAUGGUCACUGCUAUUUAUUCGCCACUUUUAUUUAUGAUACCCCUUUUUUUUUUUUUUGCGAUUUUUGCCAUAUUUAGCUUUUAUCUAUCGC ((((.(((((((((.((......)).)))))))))))))...............................................(((((........................))))) (-16.83 = -17.16 + 0.33)

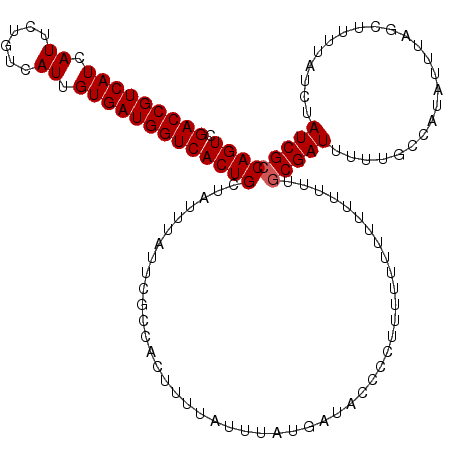

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:08:20 2006