
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 16,055,212 – 16,055,529 |
| Length | 317 |
| Max. P | 0.999902 |

| Location | 16,055,212 – 16,055,332 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 95.67 |
| Mean single sequence MFE | -36.26 |
| Consensus MFE | -36.16 |
| Energy contribution | -35.28 |
| Covariance contribution | -0.88 |
| Combinations/Pair | 1.17 |
| Mean z-score | -2.28 |
| Structure conservation index | 1.00 |
| SVM decision value | 4.46 |
| SVM RNA-class probability | 0.999902 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 16055212 120 + 23771897 CACAUCAAAACCAUAUCCGCUCUGGCCAGCCGGCCAUCCUUGGAGAAACCCAGGCCGGCAAAGCCUCGUGUGUGAACAGUUACUUCAAAGUAAAUGCUAUUUUCGUUGGCGUUUAUCCAC ...........((((..((....(((..(((((((.....(((......))))))))))...))).))..))))...............((((((((((.......)))))))))).... ( -35.60) >DroSec_CAF1 5237 120 + 1 CACAACAAAACCAUAUCCGCUCUGGCCAGCCGGCCAUCCUUGGAGAAGCCCAGGCCGGCAAAGCCUCGCGUGUGAACAGUUACUUCGAAGUAAAUGCUAUUUUCGUUGGCAUUUAUCCAC ...........((((..(((...(((..(((((((.....(((......))))))))))...)))..)))))))...............((((((((((.......)))))))))).... ( -36.60) >DroSim_CAF1 3752 120 + 1 CACAACAAAACCAUAUCCGCUCUGGCCAGCCGGCCAUCCUUGGAGAAGCCCAGGCCGGCAAAGCCUCGCGUGUGAACAGUUACUUCGAAGUAAAUGCUAUUUUCGUUGGCAUUUAUCCAC ...........((((..(((...(((..(((((((.....(((......))))))))))...)))..)))))))...............((((((((((.......)))))))))).... ( -36.60) >DroEre_CAF1 4340 120 + 1 CACAAUAGAACCAUAUCCGCUCUGGCCAGCCGGCCAUCCUUGGAGAAACCUAGGCCGGCAAAGCCUCGCGUAUGAACAGUAACUUCGAAGUAAAUGCUAUUUUCGUUGGCAUUUAUCCAC ...........((((..(((...(((..(((((((......((.....))..)))))))...)))..)))))))...............((((((((((.......)))))))))).... ( -34.80) >DroYak_CAF1 4260 120 + 1 CACAAUAAAACCACAUCCGCUCUGGCCAGCCGGCCAUCCUUGGAGAAACCCAGGCCGGCAAAGCCUCGCGUAUGAACAGUAACUUCGAAGUAAAUGCUAUUUUCGUUGGCAUUUAUCCAC .............(((.(((...(((..(((((((.....(((......))))))))))...)))..))).)))...............((((((((((.......)))))))))).... ( -37.70) >consensus CACAACAAAACCAUAUCCGCUCUGGCCAGCCGGCCAUCCUUGGAGAAACCCAGGCCGGCAAAGCCUCGCGUGUGAACAGUUACUUCGAAGUAAAUGCUAUUUUCGUUGGCAUUUAUCCAC .............(((.(((...(((..(((((((.....(((......))))))))))...)))..))).)))...............((((((((((.......)))))))))).... (-36.16 = -35.28 + -0.88)



| Location | 16,055,252 – 16,055,369 |
|---|---|
| Length | 117 |
| Sequences | 5 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 95.21 |
| Mean single sequence MFE | -34.74 |
| Consensus MFE | -32.42 |
| Energy contribution | -33.06 |
| Covariance contribution | 0.64 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.31 |
| Structure conservation index | 0.93 |
| SVM decision value | 0.88 |
| SVM RNA-class probability | 0.873246 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 16055252 117 - 23771897 AAAUGAUGGACAGCUUGGGCUUGUACUGGCUCAAAUGGUGGAUAAACGCCAACGAAAAUAGCAUUUACUUUGAAGUAACUGUUCACACACGAGGCUUUGCCGGCCUGGGUUUCUCCA ......((((.((((..((((.(((..(.(((...(((((......)))))........((((.(((((....))))).)))).......))).)..))).))))..))))..)))) ( -38.60) >DroSec_CAF1 5277 117 - 1 AAAUGAUGGACAGCUUGGGCUUGUACUGGCUCAAAUGGUGGAUAAAUGCCAACGAAAAUAGCAUUUACUUCGAAGUAACUGUUCACACGCGAGGCUUUGCCGGCCUGGGCUUCUCCA ......((((.((((..((((.(((..(.(((.....((((((((((((...........)))))((((....))))..)))))))....))).)..))).))))..))))..)))) ( -38.30) >DroSim_CAF1 3792 117 - 1 AAAUGAUGGACAGCUUGGGCUUGUACUGGCUCAAAUGGUGGAUAAAUGCCAACGAAAAUAGCAUUUACUUCGAAGUAACUGUUCACACGCGAGGCUUUGCCGGCCUGGGCUUCUCCA ......((((.((((..((((.(((..(.(((.....((((((((((((...........)))))((((....))))..)))))))....))).)..))).))))..))))..)))) ( -38.30) >DroEre_CAF1 4380 117 - 1 AAAUGAUGGAUAGCUUGGGAAUGUACUGGCUCAAAUGGUGGAUAAAUGCCAACGAAAAUAGCAUUUACUUCGAAGUUACUGUUCAUACGCGAGGCUUUGCCGGCCUAGGUUUCUCCA ......((((.((((((((...(((..(.(((.(((..(((((((((((...........))))))).))))..)))...((....))..))).)..)))...))))))))..)))) ( -29.30) >DroYak_CAF1 4300 117 - 1 AAAUGAUGGAUAGCUUGGGAAUGUACUGGCUCAAGUGGUGGAUAAAUGCCAACGAAAAUAGCAUUUACUUCGAAGUUACUGUUCAUACGCGAGGCUUUGCCGGCCUGGGUUUCUCCA ......((((.((((..((...(((..((((...((((((((((...((...((((............))))..))...))))))).)))..)))).)))...))..))))..)))) ( -29.20) >consensus AAAUGAUGGACAGCUUGGGCUUGUACUGGCUCAAAUGGUGGAUAAAUGCCAACGAAAAUAGCAUUUACUUCGAAGUAACUGUUCACACGCGAGGCUUUGCCGGCCUGGGUUUCUCCA ......((((.((((((((((.(((..(.(((...(((((......)))))........((((.(((((....))))).)))).......))).)..))).))))))))))..)))) (-32.42 = -33.06 + 0.64)


| Location | 16,055,369 – 16,055,489 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 96.17 |
| Mean single sequence MFE | -28.46 |
| Consensus MFE | -23.42 |
| Energy contribution | -24.06 |
| Covariance contribution | 0.64 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.86 |
| Structure conservation index | 0.82 |
| SVM decision value | 0.86 |
| SVM RNA-class probability | 0.869010 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
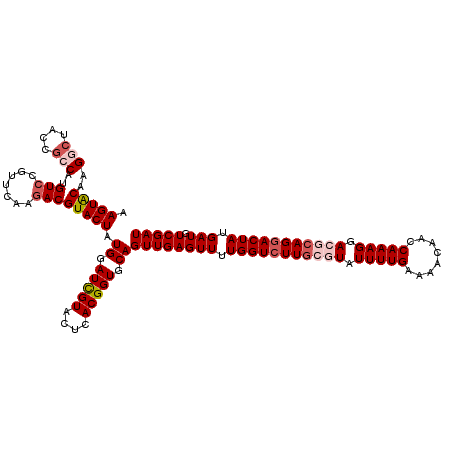
>3L_DroMel_CAF1 16055369 120 + 23771897 CGUUACGCACCCAGUUUUUGUGGUGUACAGAACCAUUUUUAUCGACAUCAUAGUCCUGCGUCCUUUGAUUGUUUUCAAAAUACGCAAGACCAAAACUCAACUGCACCGUGAGUACGAUCC (((.((.(((.(((((..((((((((...((..........)).))))))))(((.(((((..(((((......)))))..))))).)))........)))))....))).))))).... ( -30.60) >DroSec_CAF1 5394 120 + 1 CGUUACGCACCCAGUUUUUGUGGUGUACAGAACCAUUUUUAUCGACAUCAUAGUCCUGCGUCCUUUGGUUGUUUUCAAAAUACGCAAGACCAAAACUCAACUGCACCGUGAGUACGAUCC (((.((.(((.(((((..((((((((...((..........)).))))))))(((.(((((..(((((......)))))..))))).)))........)))))....))).))))).... ( -29.70) >DroSim_CAF1 3909 120 + 1 CGUUACGCACCCAGUUUUUGUGGUGUACAGAACCAUUUUUAUCGACAUCAUAGUCCUGCGUCCUUUGGUUGUUUUCAAAAUACGCAAGACCAAAACUCAACUGCACCGUGAGUACGAUCC (((.((.(((.(((((..((((((((...((..........)).))))))))(((.(((((..(((((......)))))..))))).)))........)))))....))).))))).... ( -29.70) >DroEre_CAF1 4497 120 + 1 CGUCACGCACCCAGUUUUUGUGGUGUACAGAACCAUUUUUAUCGACAUCAUAGUCCUGCGUCCUUUGGUUAUUUUCAAAAUAACCAAGACCAAAACUCAACUGCACCGUGAGUACAAUCC ..(((((....(((((..((((((((...((..........)).))))))))((..((.(((..(((((((((.....))))))))))))))..))..)))))...)))))......... ( -28.40) >DroYak_CAF1 4417 120 + 1 CGUCACGCACCCAGUUUUUGUGGUGUACAGAACCAUUUUUAUCGACAUCGUAGUCCUGCGUCCUUUGGUUAUUUUCAAAAUAACAAACACCAAAACUCAACUGCACCGUGAGUACAAUCC ..(((((....(((((....((((((................(((...((((....))))....)))((((((.....))))))..))))))......)))))...)))))......... ( -23.90) >consensus CGUUACGCACCCAGUUUUUGUGGUGUACAGAACCAUUUUUAUCGACAUCAUAGUCCUGCGUCCUUUGGUUGUUUUCAAAAUACGCAAGACCAAAACUCAACUGCACCGUGAGUACGAUCC ..(((((....(((((..((((((((...((..........)).))))))))(((.(((((..(((((......)))))..))))).)))........)))))...)))))......... (-23.42 = -24.06 + 0.64)



| Location | 16,055,369 – 16,055,489 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 96.17 |
| Mean single sequence MFE | -36.65 |
| Consensus MFE | -32.54 |
| Energy contribution | -33.26 |
| Covariance contribution | 0.72 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.25 |
| Structure conservation index | 0.89 |
| SVM decision value | 2.84 |
| SVM RNA-class probability | 0.997322 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 16055369 120 - 23771897 GGAUCGUACUCACGGUGCAGUUGAGUUUUGGUCUUGCGUAUUUUGAAAACAAUCAAAGGACGCAGGACUAUGAUGUCGAUAAAAAUGGUUCUGUACACCACAAAAACUGGGUGCGUAACG (((((((((.....)))).((((((((.(((((((((((.((((((......)))))).))))))))))).))).)))))......))))).((((.(((.......)))))))...... ( -39.90) >DroSec_CAF1 5394 120 - 1 GGAUCGUACUCACGGUGCAGUUGAGUUUUGGUCUUGCGUAUUUUGAAAACAACCAAAGGACGCAGGACUAUGAUGUCGAUAAAAAUGGUUCUGUACACCACAAAAACUGGGUGCGUAACG (((((((((.....)))).((((((((.(((((((((((.(((((........))))).))))))))))).))).)))))......))))).((((.(((.......)))))))...... ( -38.50) >DroSim_CAF1 3909 120 - 1 GGAUCGUACUCACGGUGCAGUUGAGUUUUGGUCUUGCGUAUUUUGAAAACAACCAAAGGACGCAGGACUAUGAUGUCGAUAAAAAUGGUUCUGUACACCACAAAAACUGGGUGCGUAACG (((((((((.....)))).((((((((.(((((((((((.(((((........))))).))))))))))).))).)))))......))))).((((.(((.......)))))))...... ( -38.50) >DroEre_CAF1 4497 120 - 1 GGAUUGUACUCACGGUGCAGUUGAGUUUUGGUCUUGGUUAUUUUGAAAAUAACCAAAGGACGCAGGACUAUGAUGUCGAUAAAAAUGGUUCUGUACACCACAAAAACUGGGUGCGUGACG .((((((((.....))))))))..(((.((((((((((((((.....))))))))..))))((((((((((.............)))))))))).((((.((.....)))))))).))). ( -36.62) >DroYak_CAF1 4417 120 - 1 GGAUUGUACUCACGGUGCAGUUGAGUUUUGGUGUUUGUUAUUUUGAAAAUAACCAAAGGACGCAGGACUACGAUGUCGAUAAAAAUGGUUCUGUACACCACAAAAACUGGGUGCGUGACG .........(((((((.(((((...((.((((((..(((.(((((........))))))))(((((((((((....)).......))))))))))))))).)).))))).)).))))).. ( -29.71) >consensus GGAUCGUACUCACGGUGCAGUUGAGUUUUGGUCUUGCGUAUUUUGAAAACAACCAAAGGACGCAGGACUAUGAUGUCGAUAAAAAUGGUUCUGUACACCACAAAAACUGGGUGCGUAACG (.(((((....))))).).((((((((.(((((((((((.(((((........))))).))))))))))).))).))))).......(((.(((((.(((.......)))))))).))). (-32.54 = -33.26 + 0.72)



| Location | 16,055,409 – 16,055,529 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 92.83 |
| Mean single sequence MFE | -37.14 |
| Consensus MFE | -30.00 |
| Energy contribution | -31.60 |
| Covariance contribution | 1.60 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.19 |
| Structure conservation index | 0.81 |
| SVM decision value | 1.65 |
| SVM RNA-class probability | 0.970150 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 16055409 120 - 23771897 AAGUACAAGGCUAACGCCAUGUCCGUUCAAGACGUACUAUGGAUCGUACUCACGGUGCAGUUGAGUUUUGGUCUUGCGUAUUUUGAAAACAAUCAAAGGACGCAGGACUAUGAUGUCGAU .(((((..(((....)))..(((.......)))))))).((.(((((....))))).))((((((((.(((((((((((.((((((......)))))).))))))))))).))).))))) ( -44.10) >DroSec_CAF1 5434 120 - 1 AAGUACAAGGCUACCGACAUGUCCGUUCAAGACGUACUAUGGAUCGUACUCACGGUGCAGUUGAGUUUUGGUCUUGCGUAUUUUGAAAACAACCAAAGGACGCAGGACUAUGAUGUCGAU .(((.....)))..((((((.......((((((......((.(((((....))))).)).....))))))(((((((((.(((((........))))).)))))))))....)))))).. ( -37.20) >DroSim_CAF1 3949 120 - 1 AAGUACAAGGCUACCGACAUGUCCGUUCAAGACGUACUAUGGAUCGUACUCACGGUGCAGUUGAGUUUUGGUCUUGCGUAUUUUGAAAACAACCAAAGGACGCAGGACUAUGAUGUCGAU .(((.....)))..((((((.......((((((......((.(((((....))))).)).....))))))(((((((((.(((((........))))).)))))))))....)))))).. ( -37.20) >DroEre_CAF1 4537 120 - 1 AAGUGCAAGGAUUCCGCCAUGUCCUAUCAAAACGUACUAUGGAUUGUACUCACGGUGCAGUUGAGUUUUGGUCUUGGUUAUUUUGAAAAUAACCAAAGGACGCAGGACUAUGAUGUCGAU ...(((..((......))..(((((((((((((......(.((((((((.....)))))))).))))))))).(((((((((.....))))))))))))))))).(((......)))... ( -35.30) >DroYak_CAF1 4457 120 - 1 AAGUACAAGGCAUCCGCCAUGUCCUUGCAAGACGUACUAUGGAUUGUACUCACGGUGCAGUUGAGUUUUGGUGUUUGUUAUUUUGAAAAUAACCAAAGGACGCAGGACUACGAUGUCGAU (((.(((.(((....))).))).)))....(((((......((((((((.....)))))))).(((((((((.(((((((((.....))))))..))).)).)))))))...)))))... ( -31.90) >consensus AAGUACAAGGCUACCGCCAUGUCCGUUCAAGACGUACUAUGGAUCGUACUCACGGUGCAGUUGAGUUUUGGUCUUGCGUAUUUUGAAAACAACCAAAGGACGCAGGACUAUGAUGUCGAU .(((((..(((....)))..(((.......)))))))).((.(((((....))))).))((((((((.(((((((((((.(((((........))))).))))))))))).))).))))) (-30.00 = -31.60 + 1.60)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:06:59 2006