
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 16,017,731 – 16,017,821 |
| Length | 90 |
| Max. P | 0.928173 |

| Location | 16,017,731 – 16,017,821 |
|---|---|
| Length | 90 |
| Sequences | 5 |
| Columns | 95 |
| Reading direction | forward |
| Mean pairwise identity | 83.85 |
| Mean single sequence MFE | -28.08 |
| Consensus MFE | -19.07 |
| Energy contribution | -20.31 |
| Covariance contribution | 1.24 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.08 |
| Structure conservation index | 0.68 |
| SVM decision value | 0.61 |
| SVM RNA-class probability | 0.799875 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
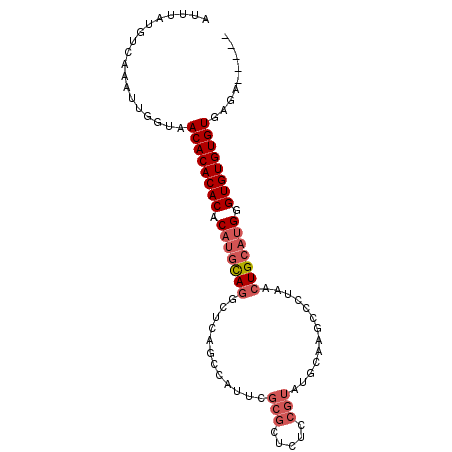
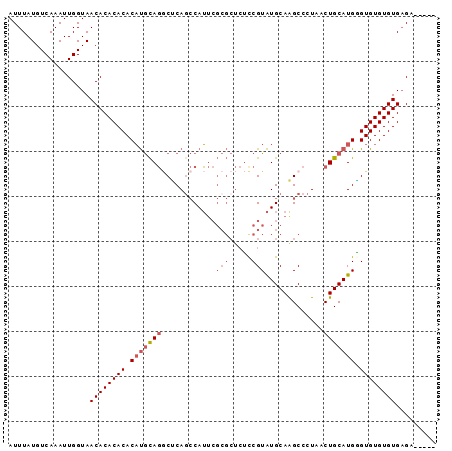
>3L_DroMel_CAF1 16017731 90 + 23771897 AUUUAUGUCAAAUUGGUAACACACACACAUGCAGGCUCAGCCAUUCGCGCUCUCCGUAUGCAAGCCCUAACUGCAUGAGUGUGUGUGAGC----- ..................((((((((.((((((((((..((.....))((.........)).)))).....)))))).))))))))....----- ( -27.20) >DroSec_CAF1 17568 90 + 1 AUUUAUGUCAAAUUGGUAACACACACACAUGCAGGCUCAGCCAUUCGCGCUCUCCGUAUGCAAGCCCUAACUGCAUGGGUGUGUGUGAGA----- ..................((((((((.((((((((((..((.....))((.........)).)))).....)))))).))))))))....----- ( -27.20) >DroSim_CAF1 26209 90 + 1 AUUUAUGUCAAAUUGGUAACACACACACAUGCAGGCUCAACCAUUCGCGCUCUCCGUAUGCAAGCCCUAACUGCAUGGGUGUGUGUGAGA----- ..................((((((((.(((((((((..........))((.........)).........))))))).))))))))....----- ( -26.30) >DroEre_CAF1 20803 95 + 1 AUUUAUGUCAAAUUGGUAACACACACACAUGCAGCCCCAGCCAUUCGCGCUCUUCGUAUGCAGGCCCUAACUGCAUGGGUGUGUGUGAGUGUGCC ...................(((((.((((((((.(((.((((....).)))......((((((.......))))))))))))))))).))))).. ( -30.90) >DroAna_CAF1 18721 85 + 1 AUUUAUGUCAAAUUGGUAACACACACACCCAUACACACAGACAG--------AUCGUGUGCAGGCCCUAGCUGCAUGUGUGUGUGUGUGU--UGG ................((((((((((((.((((.((((.((...--------.))))))((((.......)))).)))).))))))))))--)). ( -28.80) >consensus AUUUAUGUCAAAUUGGUAACACACACACAUGCAGGCUCAGCCAUUCGCGCUCUCCGUAUGCAAGCCCUAACUGCAUGGGUGUGUGUGAGA_____ ..................((((((((.(((((((............(((.....))).............))))))).))))))))......... (-19.07 = -20.31 + 1.24)

| Location | 16,017,731 – 16,017,821 |
|---|---|
| Length | 90 |
| Sequences | 5 |
| Columns | 95 |
| Reading direction | reverse |
| Mean pairwise identity | 83.85 |
| Mean single sequence MFE | -29.82 |
| Consensus MFE | -22.03 |
| Energy contribution | -22.11 |
| Covariance contribution | 0.08 |
| Combinations/Pair | 1.20 |
| Mean z-score | -2.26 |
| Structure conservation index | 0.74 |
| SVM decision value | 1.19 |
| SVM RNA-class probability | 0.928173 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
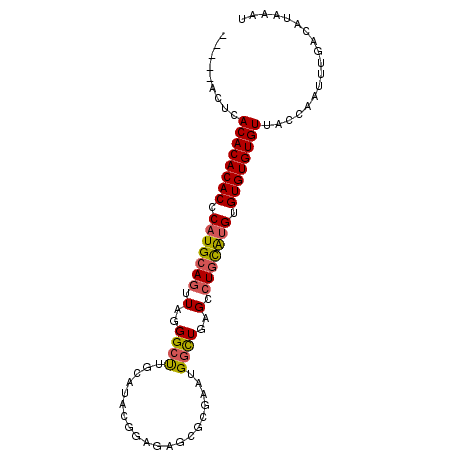
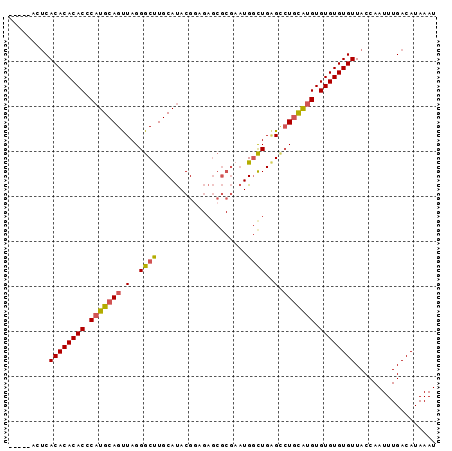
>3L_DroMel_CAF1 16017731 90 - 23771897 -----GCUCACACACACUCAUGCAGUUAGGGCUUGCAUACGGAGAGCGCGAAUGGCUGAGCCUGCAUGUGUGUGUGUUACCAAUUUGACAUAAAU -----....((((((((.((((((....(.(((..(.....)..))).)....(((...))))))))).)))))))).................. ( -29.30) >DroSec_CAF1 17568 90 - 1 -----UCUCACACACACCCAUGCAGUUAGGGCUUGCAUACGGAGAGCGCGAAUGGCUGAGCCUGCAUGUGUGUGUGUUACCAAUUUGACAUAAAU -----....((((((((.((((((....(.(((..(.....)..))).)....(((...))))))))).)))))))).................. ( -28.50) >DroSim_CAF1 26209 90 - 1 -----UCUCACACACACCCAUGCAGUUAGGGCUUGCAUACGGAGAGCGCGAAUGGUUGAGCCUGCAUGUGUGUGUGUUACCAAUUUGACAUAAAU -----....((((((((.((((((.....(((((.(((.((.......)).)))...))))))))))).)))))))).................. ( -27.50) >DroEre_CAF1 20803 95 - 1 GGCACACUCACACACACCCAUGCAGUUAGGGCCUGCAUACGAAGAGCGCGAAUGGCUGGGGCUGCAUGUGUGUGUGUUACCAAUUUGACAUAAAU .........((((((((.(((((((((..(((((((...........)))...))))..))))))))).)))))))).................. ( -34.10) >DroAna_CAF1 18721 85 - 1 CCA--ACACACACACACACAUGCAGCUAGGGCCUGCACACGAU--------CUGUCUGUGUGUAUGGGUGUGUGUGUUACCAAUUUGACAUAAAU ..(--((((((((((...(((((.((....))..(((((.((.--------...)))))))))))).)))))))))))................. ( -29.70) >consensus _____ACUCACACACACCCAUGCAGUUAGGGCUUGCAUACGGAGAGCGCGAAUGGCUGAGCCUGCAUGUGUGUGUGUUACCAAUUUGACAUAAAU .........((((((((.(((((((.(..((((....................))))..).))))))).)))))))).................. (-22.03 = -22.11 + 0.08)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:06:19 2006