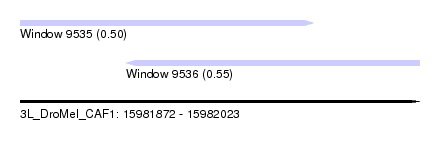
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 15,981,872 – 15,982,023 |
| Length | 151 |
| Max. P | 0.546762 |

| Location | 15,981,872 – 15,981,983 |
|---|---|
| Length | 111 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 89.69 |
| Mean single sequence MFE | -35.20 |
| Consensus MFE | -27.48 |
| Energy contribution | -27.48 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.40 |
| Structure conservation index | 0.78 |
| SVM decision value | -0.07 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
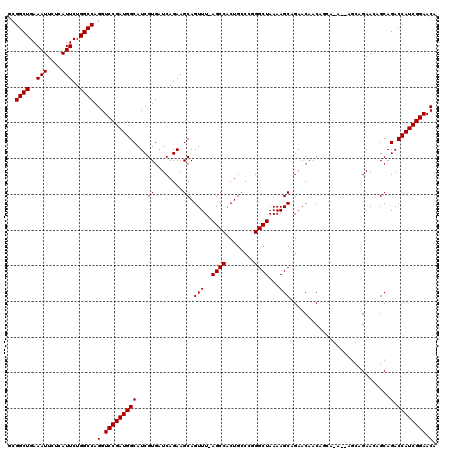
>3L_DroMel_CAF1 15981872 111 + 23771897 GCGGCUGAAAUUCUCAUUCUGGCCAGGUCCGAUGGCAUCGUGAUCAGAAGCAGUUU-AGCCACUGCCCGGGCUAAAAGCAGAACAACAGC--------AGAAGAGCAGACCAUCGGAACA ..((((..(((....)))..)))).(.(((((((((.............((..(((-((((........))))))).)).........((--------......)).).)))))))).). ( -33.20) >DroSec_CAF1 11077 120 + 1 GCGGCUGAAAUUCUCAUUCUGGCCAGGUCCGAUGGCAUCGUGAUCAGAAGCAGUUUAAGCCACUGGCCGGGCUAAAAGCAGAACAACAGCAAAAGAGCAGAAGAGCAGACCAUCGGAACA ..(((((...((((....((((((((.((.(((.((...)).))).)).((.......))..))))))))(((....((.........)).....)))))))...))).))......... ( -32.00) >DroSim_CAF1 11826 111 + 1 GCGGCUGAAAUUCUCAUUCUGGCCAGGUCCGAUGGCAUCGUGAUCAGAAGCAGUUU-AGCCACUGCCCGGGCUAAAAGCAGAACAACAGC--------AGAAAAGCAGACCAUCGGAACA ..((((..(((....)))..)))).(.(((((((((.............((..(((-((((........))))))).)).........((--------......)).).)))))))).). ( -33.20) >DroEre_CAF1 11786 116 + 1 GCGGCUGAAAUUCUGAUUCUGGCCAGGUCCGAUGGCAUCGUGAUCAGAAGCAGUUU-AGCCCCUGCCAUGGCUAAAAGCAGAACAGC---AGAACAGCAGAACUGCAGACCAUCGGAACA ..((((..(((....)))..)))).(.(((((((((.((.......)).(((((((-.((..((((....((.....))......))---))....)).))))))).).)))))))).). ( -38.80) >DroYak_CAF1 11315 116 + 1 GCGGCUGAAAUUCUGAUUCUGGCCAGGUCCGAUGGCAUCGUGAUCAGAAGCAGUUU-AGCCACUGCCAUGGCUAAAAGCAGAACAGC---AGAGCAGCAGAACUGCAGACCAUCGGAACA ..((((..(((....)))..)))).(.(((((((((.((.......)).(((((((-.((..((((....((.....))......))---))....)).))))))).).)))))))).). ( -38.80) >consensus GCGGCUGAAAUUCUCAUUCUGGCCAGGUCCGAUGGCAUCGUGAUCAGAAGCAGUUU_AGCCACUGCCCGGGCUAAAAGCAGAACAACAGCA_A__AGCAGAACAGCAGACCAUCGGAACA ..((((..(((....)))..)))).(.(((((((((...((........)).(((..((((........))))...)))............................).)))))))).). (-27.48 = -27.48 + 0.00)


| Location | 15,981,912 – 15,982,023 |
|---|---|
| Length | 111 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 89.86 |
| Mean single sequence MFE | -36.32 |
| Consensus MFE | -27.88 |
| Energy contribution | -28.04 |
| Covariance contribution | 0.16 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.50 |
| Structure conservation index | 0.77 |
| SVM decision value | 0.03 |
| SVM RNA-class probability | 0.546762 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
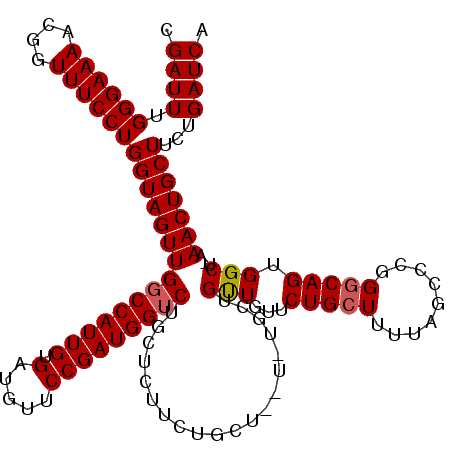
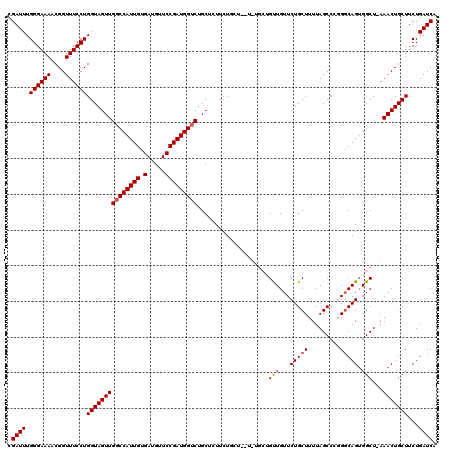
>3L_DroMel_CAF1 15981912 111 - 23771897 CGAUUUGGGAAAACGGUUUCCUGGUAGUUGGCCAUUGUGAUGUUCCGAUGGUCUGCUCUUCU--------GCUGUUGUUCUGCUUUUAGCCCGGGCAGUGGCU-AAACUGCUUCUGAUCA .((((.((((((....))))))(((((((((((((((.(.....))))))))).........--------((..((((((.((.....))..))))))..)).-.)))))))...)))). ( -35.80) >DroSec_CAF1 11117 120 - 1 CGAUUUGGGAAAACGGUUUCCUGGUAGUUGGCCAUUGUGAUGUUCCGAUGGUCUGCUCUUCUGCUCUUUUGCUGUUGUUCUGCUUUUAGCCCGGCCAGUGGCUUAAACUGCUUCUGAUCA .((((.((((((....))))))(((((((((((((((.(.....))))))))).((......))......((..(((..(.((.....))..)..)))..))...)))))))...)))). ( -33.60) >DroSim_CAF1 11866 111 - 1 CGAUUUGGGAAAACGGUUUCCUGGUAGUUGGCCAUUGUGAUGUUCCGAUGGUCUGCUUUUCU--------GCUGUUGUUCUGCUUUUAGCCCGGGCAGUGGCU-AAACUGCUUCUGAUCA .((((.((((((....))))))(((((((((((((((.(.....))))))))).........--------((..((((((.((.....))..))))))..)).-.)))))))...)))). ( -35.80) >DroEre_CAF1 11826 116 - 1 CGAUUUGGGAAAACGGUUUCCUGGUAGUUGCCCAUUGUGAUGUUCCGAUGGUCUGCAGUUCUGCUGUUCU---GCUGUUCUGCUUUUAGCCAUGGCAGGGGCU-AAACUGCUUCUGAUCA .((((.((((((....))))))(((((((((((.(((.......((.(((((..((((..(.((......---)).)..)))).....)))))))))))))).-.)))))))...)))). ( -35.01) >DroYak_CAF1 11355 116 - 1 CGAUUUGGGAAAACGGUUUCCUGGUAGUUGGCCAUUGUGAUGUUCCGAUGGUCUGCAGUUCUGCUGCUCU---GCUGUUCUGCUUUUAGCCAUGGCAGUGGCU-AAACUGCUUCUGAUCA .((((.((((((....))))))(((((((((((((((.(.....))))))))).((((..(.((......---)).)..))))...(((((((....))))))-))))))))...)))). ( -41.40) >consensus CGAUUUGGGAAAACGGUUUCCUGGUAGUUGGCCAUUGUGAUGUUCCGAUGGUCUGCUCUUCUGCU__U_UGCUGUUGUUCUGCUUUUAGCCCGGGCAGUGGCU_AAACUGCUUCUGAUCA .((((.((((((....))))))(((((((((((((((.(.....)))))))))....................(((...(((((.........))))).)))...)))))))...)))). (-27.88 = -28.04 + 0.16)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:05:28 2006