
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 15,853,444 – 15,853,641 |
| Length | 197 |
| Max. P | 0.969896 |

| Location | 15,853,444 – 15,853,542 |
|---|---|
| Length | 98 |
| Sequences | 5 |
| Columns | 106 |
| Reading direction | forward |
| Mean pairwise identity | 86.97 |
| Mean single sequence MFE | -38.16 |
| Consensus MFE | -27.28 |
| Energy contribution | -29.48 |
| Covariance contribution | 2.20 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.85 |
| Structure conservation index | 0.71 |
| SVM decision value | 0.27 |
| SVM RNA-class probability | 0.661331 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 15853444 98 + 23771897 --------CAAGUUUUCUUUUCUCUGUACCCAAAUCCUUUGGGUAGUGGGUGGUGGUGGUGCGUUCGUAGGCUUGGCCACCACGCCGCACCGGAUGUCCUGGCAUU --------..................(((((((.....)))))))((((((((((((.(.((.(....).)).).)))))))).)))).((((.....)))).... ( -37.10) >DroSec_CAF1 60658 106 + 1 CAAUUUUAAAAGAUUUCUUUUCUCUGUACCCAAAUCCUUUGGGUAGUGGGUGGUGGUGGUGCGUUCGUAGGCUUGGCCACCGCGCCGCACCGGAUGUCCUGGCAUU .......(((((....))))).....(((((((.....)))))))((((((((((((.(.((.(....).)).).)))))))).)))).((((.....)))).... ( -37.80) >DroSim_CAF1 62226 106 + 1 CAAUUUUAAAAGAUUUCUUUUCUCUGUACCCAAAUCCUUUGGGUAGUGGGUGGUGGUGGUGCGUUCGUAGGCUUGGCCACCGCGCCGCACCGGAUGUCCUGGCAUU .......(((((....))))).....(((((((.....)))))))((((((((((((.(.((.(....).)).).)))))))).)))).((((.....)))).... ( -37.80) >DroEre_CAF1 60476 100 + 1 CAACGUUAGAGGAUUUGUUUUCUGUGCACCCAAAUCCUUGGGGGAGU------UGGUGGUGCGGUCGUAGGCUUGGCCACCGCACCGCACCGGAUGUCCUGGCAUU (((((((((((((....))))))).)).(((((....)))))...))------)).(((((((((((..(((...)))..)).)))))))))(((((....))))) ( -39.30) >DroYak_CAF1 102054 104 + 1 CAAUAUUAGAAGAUUUUUUUUCUCUGUACCCAAAUCCUUGGGGGGGUGGG-GGUGGUGGUGCG-UUGAAGGCUUGGCCACCGCACCGCACCGGAUGUCCUGGCAUU ...(((.((((((....))))))..)))(((((....)))))..((((.(-(((((((((((.-......))...)))))))).)).)))).(((((....))))) ( -38.80) >consensus CAAUUUUAAAAGAUUUCUUUUCUCUGUACCCAAAUCCUUUGGGUAGUGGGUGGUGGUGGUGCGUUCGUAGGCUUGGCCACCGCGCCGCACCGGAUGUCCUGGCAUU ..........................(((((((.....)))))))((((((((((((.(.((........)).).)))))))).)))).((((.....)))).... (-27.28 = -29.48 + 2.20)

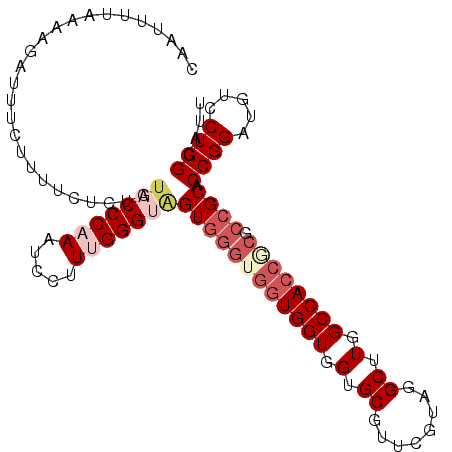

| Location | 15,853,542 – 15,853,641 |
|---|---|
| Length | 99 |
| Sequences | 5 |
| Columns | 99 |
| Reading direction | forward |
| Mean pairwise identity | 98.18 |
| Mean single sequence MFE | -27.40 |
| Consensus MFE | -23.20 |
| Energy contribution | -23.80 |
| Covariance contribution | 0.60 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.33 |
| Structure conservation index | 0.85 |
| SVM decision value | 1.61 |
| SVM RNA-class probability | 0.967103 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
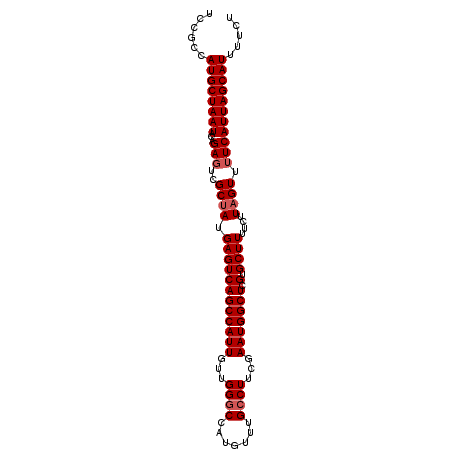
>3L_DroMel_CAF1 15853542 99 + 23771897 UCCGCCAUGCUAAUACAGAGUCGCUAUGAGUCAGCCAUUGUUGGGCCAUGUUUGCCUUCGAAUGGCUCGUGCUUUUCUUAGUUUUCAUUAGCAUUUUCU ......((((((((...(((..((((.((((((((((((...((((.......))))...))))))).).))))....)))).)))))))))))..... ( -26.40) >DroSec_CAF1 60764 99 + 1 UCCGCCAUGCUAAUACAGACUCGCUAUGAGUCAGCCAUUGUUGGGCCAUGUUUGCCUUCGAAUGGCUCGUGCUUUUCUUAGUUUUCAUUAGCAUUUUCU ......((((((((...(((((.....)))))(((((((...((((.......))))...)))))))...................))))))))..... ( -28.10) >DroSim_CAF1 62332 99 + 1 UCCGCCAUGCUAAUACAGAGUCGCUAUGAGUCAGCCAUUGUUGGGCCAUGUUUGCCUUCGAAUGGCUCGUGCUUUUCUUAGUUUUCAUUAGCAUUUUCU ......((((((((...(((..((((.((((((((((((...((((.......))))...))))))).).))))....)))).)))))))))))..... ( -26.40) >DroEre_CAF1 60576 99 + 1 UCCGCCAUGCUAAUACAGAGUCGCAAUGAGUCAGCCAUUGUUGGGCCAUGUUUGGCUUUGAAUGGCUCGUGCUUUUCUUAGUUUUCAUUAGCAUUUUCU ......((((((((((((((..(((.(((((((..((..((..(((...)))..))..))..))))))))))..))))..))....))))))))..... ( -29.50) >DroYak_CAF1 102158 99 + 1 UCCGCCAUGCUAAUACAGAGUCGCUAUGAGUCAGCCAUUGUUGGGCCAUGUUUGCCUUUGAAUGGCUCGUGCUUUUCUUAGUUUUCAUUAGCAUUUUCU ......((((((((((((((..((.((((((((..((.....((((.......)))).))..))))))))))..))))..))....))))))))..... ( -26.60) >consensus UCCGCCAUGCUAAUACAGAGUCGCUAUGAGUCAGCCAUUGUUGGGCCAUGUUUGCCUUCGAAUGGCUCGUGCUUUUCUUAGUUUUCAUUAGCAUUUUCU ......((((((((...(((..((((.((((((((((((...((((.......))))...))))))).).))))....)))).)))))))))))..... (-23.20 = -23.80 + 0.60)



| Location | 15,853,542 – 15,853,641 |
|---|---|
| Length | 99 |
| Sequences | 5 |
| Columns | 99 |
| Reading direction | reverse |
| Mean pairwise identity | 98.18 |
| Mean single sequence MFE | -24.42 |
| Consensus MFE | -21.36 |
| Energy contribution | -21.56 |
| Covariance contribution | 0.20 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.06 |
| Structure conservation index | 0.87 |
| SVM decision value | 1.65 |
| SVM RNA-class probability | 0.969896 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 15853542 99 - 23771897 AGAAAAUGCUAAUGAAAACUAAGAAAAGCACGAGCCAUUCGAAGGCAAACAUGGCCCAACAAUGGCUGACUCAUAGCGACUCUGUAUUAGCAUGGCGGA .(...(((((((((..................(((((((....(((.......)))....)))))))((.((.....)).))..)))))))))..)... ( -22.90) >DroSec_CAF1 60764 99 - 1 AGAAAAUGCUAAUGAAAACUAAGAAAAGCACGAGCCAUUCGAAGGCAAACAUGGCCCAACAAUGGCUGACUCAUAGCGAGUCUGUAUUAGCAUGGCGGA .(...(((((((((..................(((((((....(((.......)))....)))))))(((((.....)))))..)))))))))..)... ( -28.90) >DroSim_CAF1 62332 99 - 1 AGAAAAUGCUAAUGAAAACUAAGAAAAGCACGAGCCAUUCGAAGGCAAACAUGGCCCAACAAUGGCUGACUCAUAGCGACUCUGUAUUAGCAUGGCGGA .(...(((((((((..................(((((((....(((.......)))....)))))))((.((.....)).))..)))))))))..)... ( -22.90) >DroEre_CAF1 60576 99 - 1 AGAAAAUGCUAAUGAAAACUAAGAAAAGCACGAGCCAUUCAAAGCCAAACAUGGCCCAACAAUGGCUGACUCAUUGCGACUCUGUAUUAGCAUGGCGGA .(...(((((((((.......(((...(((..(((((((....((((....)))).....))))))).......)))...))).)))))))))..)... ( -24.50) >DroYak_CAF1 102158 99 - 1 AGAAAAUGCUAAUGAAAACUAAGAAAAGCACGAGCCAUUCAAAGGCAAACAUGGCCCAACAAUGGCUGACUCAUAGCGACUCUGUAUUAGCAUGGCGGA .(...(((((((((..................(((((((....(((.......)))....)))))))((.((.....)).))..)))))))))..)... ( -22.90) >consensus AGAAAAUGCUAAUGAAAACUAAGAAAAGCACGAGCCAUUCGAAGGCAAACAUGGCCCAACAAUGGCUGACUCAUAGCGACUCUGUAUUAGCAUGGCGGA .(...(((((((((..................(((((((....(((.......)))....)))))))((.((.....)).))..)))))))))..)... (-21.36 = -21.56 + 0.20)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:03:22 2006