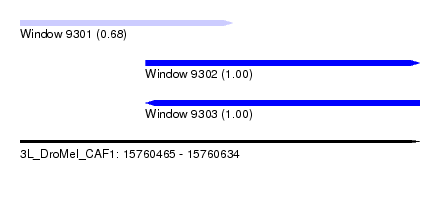

| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 15,760,465 – 15,760,634 |
| Length | 169 |
| Max. P | 0.999407 |

| Location | 15,760,465 – 15,760,555 |
|---|---|
| Length | 90 |
| Sequences | 4 |
| Columns | 90 |
| Reading direction | forward |
| Mean pairwise identity | 95.37 |
| Mean single sequence MFE | -24.31 |
| Consensus MFE | -21.25 |
| Energy contribution | -21.50 |
| Covariance contribution | 0.25 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.73 |
| Structure conservation index | 0.87 |
| SVM decision value | 0.31 |
| SVM RNA-class probability | 0.682337 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
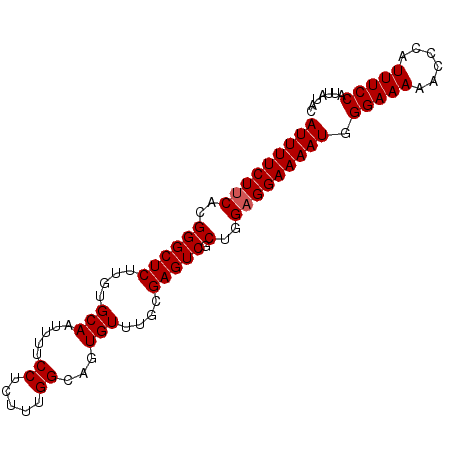
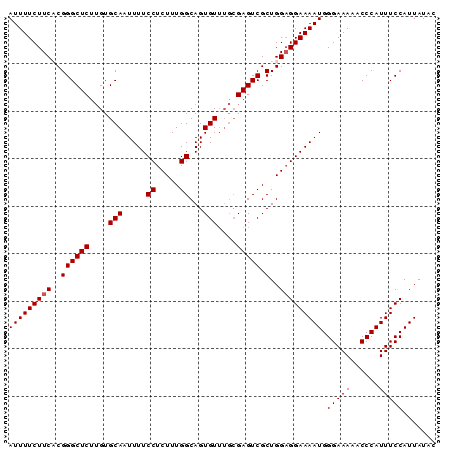
>3L_DroMel_CAF1 15760465 90 + 23771897 AUUUUCUUCUCGGGCUCUUGUGCAAUUUUCCUCUUUGGCAGUGUUUGCGAGUCGCUGGAGGAAAAUGGGAAAAACCCAUUUCCAUUAUAC ......(((((.(((.((((((((.....((.....))...)))..)))))..))).)))))(((((((.....)))))))......... ( -24.40) >DroSec_CAF1 12063 90 + 1 AUUUUCUUCACGGGCUCUUGUGCAAUUUUCCUCUUUGGCAGUGUUUGCGAGUCGCUGGAGGAAAAUGGGAAAAACCCAUUUCCAUUAUAC .(((((((((..(((((....((((..((((.....)).))...)))))))))..)))))))))(((((.....)))))........... ( -23.20) >DroSim_CAF1 12349 90 + 1 AUUUUCUUCACGGGCUCUUGUGCAAUUUUCCUCUUUGGCAGUGUUUGCGAGUCGCUGGAGGAAAAUGGGAAAAACCCAUUUCCAUUAUAC .(((((((((..(((((....((((..((((.....)).))...)))))))))..)))))))))(((((.....)))))........... ( -23.20) >DroEre_CAF1 12328 86 + 1 AUUUUCUUCUCGGGCUCCAGUGCAAUUUUCCUCUUUGGCAGUGU----GAGUCGCUGGUGGAAAAUGGGAAAAACCCAUUUCCAUUAUAC ............(((((((.(((..............))).)).----)))))..(((((((((.((((.....)))))))))))))... ( -26.44) >consensus AUUUUCUUCACGGGCUCUUGUGCAAUUUUCCUCUUUGGCAGUGUUUGCGAGUCGCUGGAGGAAAAUGGGAAAAACCCAUUUCCAUUAUAC (((((((((..((((((....(((.....((.....))...)))....))))).)..))))))))).(((((......)))))....... (-21.25 = -21.50 + 0.25)

| Location | 15,760,518 – 15,760,634 |
|---|---|
| Length | 116 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 94.90 |
| Mean single sequence MFE | -25.17 |
| Consensus MFE | -21.68 |
| Energy contribution | -22.16 |
| Covariance contribution | 0.48 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.38 |
| Structure conservation index | 0.86 |
| SVM decision value | 2.80 |
| SVM RNA-class probability | 0.997122 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
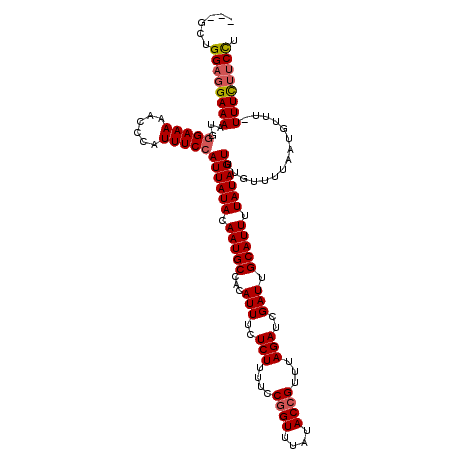
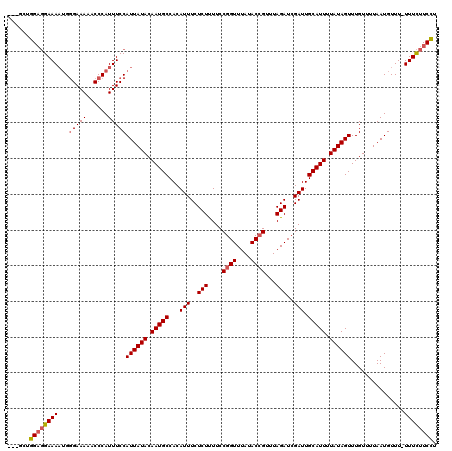
>3L_DroMel_CAF1 15760518 116 + 23771897 ---GCUGGAGGAAAAUGGGAAAAACCCAUUUCCAUUAUACAAUGCCACAUUUCUCUUUUCCGGUUUAUACCGUUUAGAUCGAUUGCAUUUUAUAGUUUGUUUUAAUGUUU-UUUCUUCCU ---...(((((((((((((.....)))).....((((((.(((((...(((..(((....((((....))))...)))..))).))))).)))))).............)-)))))))). ( -27.80) >DroSec_CAF1 12116 117 + 1 ---GCUGGAGGAAAAUGGGAAAAACCCAUUUCCAUUAUACAAUGCCACAUUUCUCUUUUCCGGUUUAUACCGUUUAGAUCGAUUGCAUUUUAUAGUUUGUUUUAAUGUUUUUUUCUUCCU ---...(((((((((((((.....)))).....((((((.(((((...(((..(((....((((....))))...)))..))).))))).))))))..............))))))))). ( -27.80) >DroSim_CAF1 12402 116 + 1 ---GCUGGAGGAAAAUGGGAAAAACCCAUUUCCAUUAUACAAUGCCACAUUUCUCUUUUCCGGUUUAUACCGUUUAGAUCGAUUGCAUUUUAUAGUUUGUUUUAAUGUUU-UUUCUUCCU ---...(((((((((((((.....)))).....((((((.(((((...(((..(((....((((....))))...)))..))).))))).)))))).............)-)))))))). ( -27.80) >DroEre_CAF1 12377 116 + 1 ---GCUGGUGGAAAAUGGGAAAAACCCAUUUCCAUUAUACAAUGCCACAUUUCUCUUUUCCGGUUUAUACCGUUUAGAUCGAUUGCAUUUUAUAGUUUGUUUUAAUGUUU-UUUCUUCCU ---...((.(((((.((((.....)))))))))((((((.(((((...(((..(((....((((....))))...)))..))).))))).))))))..............-......)). ( -24.10) >DroAna_CAF1 7081 118 + 1 CCUGCAGGACGAAA-AGAGAAAAACCCAUUUCCAUUAUACAAUGCCACAUUUCUCUCUUCCGGUUUAUACAGUUUAGAUCGAUUGCAUUUUAUAGUUUGUUUUAAUGUUU-UUUUUUCUG ((....))......-(((((((((..((((...((((((.(((((...............(((((((.......)))))))...))))).)))))).......))))..)-)))))))). ( -18.37) >consensus ___GCUGGAGGAAAAUGGGAAAAACCCAUUUCCAUUAUACAAUGCCACAUUUCUCUUUUCCGGUUUAUACCGUUUAGAUCGAUUGCAUUUUAUAGUUUGUUUUAAUGUUU_UUUCUUCCU ......((((((((...(((((......)))))((((((.(((((...(((..(((....((((....))))...)))..))).))))).))))))...............)))))))). (-21.68 = -22.16 + 0.48)

| Location | 15,760,518 – 15,760,634 |
|---|---|
| Length | 116 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 94.90 |
| Mean single sequence MFE | -25.49 |
| Consensus MFE | -22.26 |
| Energy contribution | -23.10 |
| Covariance contribution | 0.84 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.73 |
| Structure conservation index | 0.87 |
| SVM decision value | 3.58 |
| SVM RNA-class probability | 0.999407 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
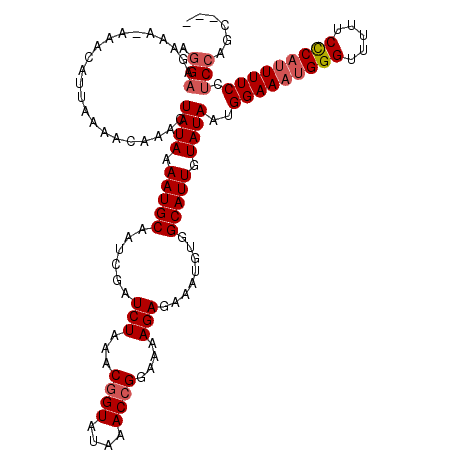
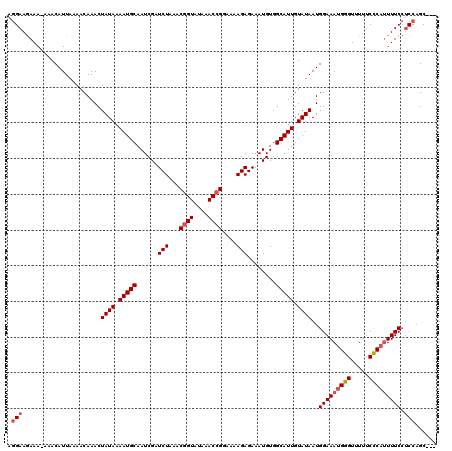
>3L_DroMel_CAF1 15760518 116 - 23771897 AGGAAGAAA-AAACAUUAAAACAAACUAUAAAAUGCAAUCGAUCUAAACGGUAUAAACCGGAAAAGAGAAAUGUGGCAUUGUAUAAUGGAAAUGGGUUUUUCCCAUUUUCCUCCAGC--- .(((.....-................((((.(((((......(((...((((....))))....)))........))))).))))..(((((((((.....))))))))).)))...--- ( -27.04) >DroSec_CAF1 12116 117 - 1 AGGAAGAAAAAAACAUUAAAACAAACUAUAAAAUGCAAUCGAUCUAAACGGUAUAAACCGGAAAAGAGAAAUGUGGCAUUGUAUAAUGGAAAUGGGUUUUUCCCAUUUUCCUCCAGC--- .(((......................((((.(((((......(((...((((....))))....)))........))))).))))..(((((((((.....))))))))).)))...--- ( -27.04) >DroSim_CAF1 12402 116 - 1 AGGAAGAAA-AAACAUUAAAACAAACUAUAAAAUGCAAUCGAUCUAAACGGUAUAAACCGGAAAAGAGAAAUGUGGCAUUGUAUAAUGGAAAUGGGUUUUUCCCAUUUUCCUCCAGC--- .(((.....-................((((.(((((......(((...((((....))))....)))........))))).))))..(((((((((.....))))))))).)))...--- ( -27.04) >DroEre_CAF1 12377 116 - 1 AGGAAGAAA-AAACAUUAAAACAAACUAUAAAAUGCAAUCGAUCUAAACGGUAUAAACCGGAAAAGAGAAAUGUGGCAUUGUAUAAUGGAAAUGGGUUUUUCCCAUUUUCCACCAGC--- .((......-................((((.(((((......(((...((((....))))....)))........))))).)))).((((((((((.....)))).))))))))...--- ( -26.54) >DroAna_CAF1 7081 118 - 1 CAGAAAAAA-AAACAUUAAAACAAACUAUAAAAUGCAAUCGAUCUAAACUGUAUAAACCGGAAGAGAGAAAUGUGGCAUUGUAUAAUGGAAAUGGGUUUUUCUCU-UUUCGUCCUGCAGG .........-......................................(((((...((..((((((((((((.((.((((....))))....)).))))))))))-))..))..))))). ( -19.80) >consensus AGGAAGAAA_AAACAUUAAAACAAACUAUAAAAUGCAAUCGAUCUAAACGGUAUAAACCGGAAAAGAGAAAUGUGGCAUUGUAUAAUGGAAAUGGGUUUUUCCCAUUUUCCUCCAGC___ .(((......................((((.(((((......(((...((((....))))....)))........))))).))))..(((((((((.....))))))))).)))...... (-22.26 = -23.10 + 0.84)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:01:38 2006