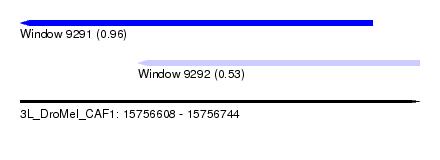
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 15,756,608 – 15,756,744 |
| Length | 136 |
| Max. P | 0.959499 |

| Location | 15,756,608 – 15,756,728 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 87.00 |
| Mean single sequence MFE | -39.26 |
| Consensus MFE | -26.98 |
| Energy contribution | -28.98 |
| Covariance contribution | 2.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -3.17 |
| Structure conservation index | 0.69 |
| SVM decision value | 1.50 |
| SVM RNA-class probability | 0.959499 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
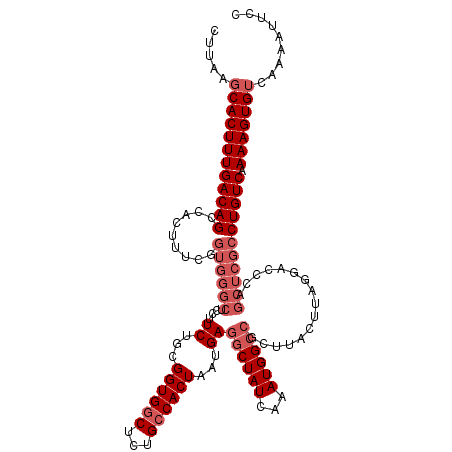
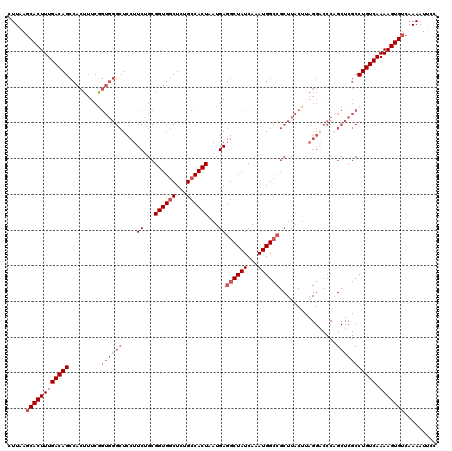
>3L_DroMel_CAF1 15756608 120 - 23771897 CUUAAGCACUUUGACAGCCACUUUCGGUGGGCUCCUUCUGCGGUGGCUCUGCCACUAAUGAGGCUAUCAAAUGGCCGCUUACUUAGGACCCAGCUCGCCUGUCAAAAGUGUCAAAAUUCC .....((((((((((((((......))((((.((((...((((((((...)))))).....((((((...))))))))......))))))))......))))).)))))))......... ( -41.50) >DroSec_CAF1 8294 120 - 1 CUUAAGCACUUUGACAGCCACUUUCGGUGGGCUCCUUCUGCGGUGGCUCUGCCACUAAUGAGGCUAUCAAAUGGCCGCUUACUUAGGUCCCAGCUCGCCUGUCAAAAGUGUCAAAAUUCC .....(((((((((((.........(((((((((((...((((((((...)))))).....((((((...))))))))......)))....)))))))))))).)))))))......... ( -40.00) >DroSim_CAF1 8538 120 - 1 CUUAAGCACUUUGACAGCCACUUUCGGUGGGCUCCUUCUGCGGUGGCUCUGCCACUAAUGAGGCUAUCAAAUGGCCGCUUACUUAGGACCCAGCUCGCCUGUCAAAAGUGUCAAAAUUCC .....((((((((((((((......))((((.((((...((((((((...)))))).....((((((...))))))))......))))))))......))))).)))))))......... ( -41.50) >DroEre_CAF1 8360 120 - 1 CUUGAGCACUUUGACAGCCACUUUCAGUGGGCUCCUUCCGCGGUGGCUCUGCCACUAAUGAGGCUAUCAAAUGGCCGCUUACUUAGGACCCCGCUCGCCUGUCAAAAGUGUCAAAAUUUC .((((.(((((((((((.(......((((((.((((...((((((((...)))))).....((((((...))))))))......)))).)))))).).))))).))))))))))...... ( -44.40) >DroAna_CAF1 3013 96 - 1 CUUAACCACUUUGACAGACACUCUCACCCAGAUCCUUCUGCGGUGGCUCAG-CACUAAUGAGGCUAUCAAAUGG-----------------------CCUGUCAAAAGUGUCAAAAUUCC ......(((((((((((......(((((((((....)))).)))))((((.-......))))(((((...))))-----------------------)))))).)))))).......... ( -28.90) >consensus CUUAAGCACUUUGACAGCCACUUUCGGUGGGCUCCUUCUGCGGUGGCUCUGCCACUAAUGAGGCUAUCAAAUGGCCGCUUACUUAGGACCCAGCUCGCCUGUCAAAAGUGUCAAAAUUCC .....((((((((((((.........((((((....((...((((((...))))))...))((((((...))))))................))))))))))).)))))))......... (-26.98 = -28.98 + 2.00)

| Location | 15,756,648 – 15,756,744 |
|---|---|
| Length | 96 |
| Sequences | 4 |
| Columns | 96 |
| Reading direction | reverse |
| Mean pairwise identity | 98.44 |
| Mean single sequence MFE | -32.53 |
| Consensus MFE | -32.69 |
| Energy contribution | -32.50 |
| Covariance contribution | -0.19 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.58 |
| Structure conservation index | 1.00 |
| SVM decision value | -0.00 |
| SVM RNA-class probability | 0.532491 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
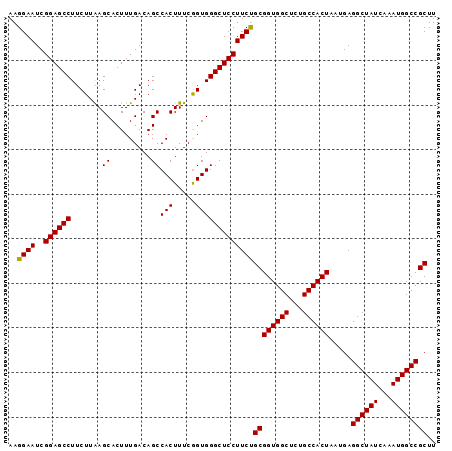
>3L_DroMel_CAF1 15756648 96 - 23771897 AAGGAAUCGGAGCCUUCUUAAGCACUUUGACAGCCACUUUCGGUGGGCUCCUUCUGCGGUGGCUCUGCCACUAAUGAGGCUAUCAAAUGGCCGCUU ..((((..((((((.......((.........))(((.....)))))))))))))((((((((...)))))).....((((((...)))))))).. ( -31.80) >DroSec_CAF1 8334 96 - 1 AAGGAAUCGGAGCCUUCUUAAGCACUUUGACAGCCACUUUCGGUGGGCUCCUUCUGCGGUGGCUCUGCCACUAAUGAGGCUAUCAAAUGGCCGCUU ..((((..((((((.......((.........))(((.....)))))))))))))((((((((...)))))).....((((((...)))))))).. ( -31.80) >DroSim_CAF1 8578 96 - 1 AAGGAAUCGGAGCCUUCUUAAGCACUUUGACAGCCACUUUCGGUGGGCUCCUUCUGCGGUGGCUCUGCCACUAAUGAGGCUAUCAAAUGGCCGCUU ..((((..((((((.......((.........))(((.....)))))))))))))((((((((...)))))).....((((((...)))))))).. ( -31.80) >DroEre_CAF1 8400 96 - 1 AAGGAAUCGGAGCCUUCUUGAGCACUUUGACAGCCACUUUCAGUGGGCUCCUUCCGCGGUGGCUCUGCCACUAAUGAGGCUAUCAAAUGGCCGCUU ..((((..(((((((.((((.((.........)))).....)).)))))))))))((((((((...)))))).....((((((...)))))))).. ( -34.70) >consensus AAGGAAUCGGAGCCUUCUUAAGCACUUUGACAGCCACUUUCGGUGGGCUCCUUCUGCGGUGGCUCUGCCACUAAUGAGGCUAUCAAAUGGCCGCUU ..((((..((((((.......((.........))(((.....)))))))))))))((((((((...)))))).....((((((...)))))))).. (-32.69 = -32.50 + -0.19)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:01:27 2006