
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 15,739,219 – 15,739,317 |
| Length | 98 |
| Max. P | 0.626349 |

| Location | 15,739,219 – 15,739,317 |
|---|---|
| Length | 98 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 70.11 |
| Mean single sequence MFE | -26.92 |
| Consensus MFE | -10.48 |
| Energy contribution | -9.48 |
| Covariance contribution | -1.00 |
| Combinations/Pair | 1.30 |
| Mean z-score | -1.80 |
| Structure conservation index | 0.39 |
| SVM decision value | 0.19 |
| SVM RNA-class probability | 0.626349 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
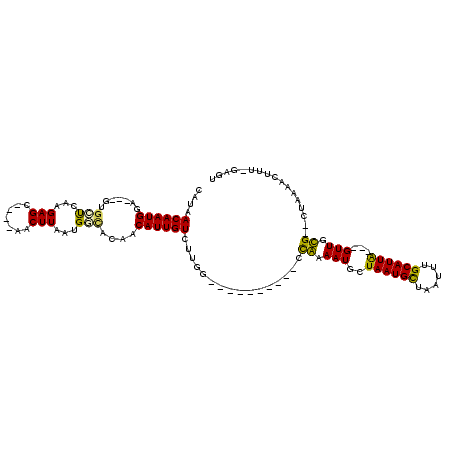
>3L_DroMel_CAF1 15739219 98 - 23771897 CAUAACAAUGGA---GUGCUCGAGAGC----AACUUAAUGGCACAACAUUGUCUUGU----------CCGAAAUGCUAAUGCUAAUUUGCAUUG---GUUGCG--CUAAAACUUUGGAGU ((..((((((..---(((((.(((...----..)))...)))))..))))))..))(----------(((((..((((((((......))))))---))....--.......)))))).. ( -27.82) >DroVir_CAF1 84545 112 - 1 AAUAACAAUGCAGA-GAGCUUCGGAGC----AUCUUAAUGUUAUACCAUUGUCUAGCAGCAGAACGCACAAAAUGCUAAUGCUCAUUUGCAUUACAAGUUGUGG-GCAGAAGCU--AUGU .........(((..-.(((((((((((----((....)))))...))..((((((.((((.....(((.....)))((((((......))))))...)))))))-)))))))))--.))) ( -30.70) >DroWil_CAF1 119894 96 - 1 CAUAACAAUGGAAAAGUUUUCAAGAGCUUGGUACUUAAUGAUAUACCAUUGUCUCGG----------UUAAAAUGCUAAUGCUAAUUUGCAUUA---GUUGUG-----------UUGCGU ....(((((((..((((((....)))))).((((.....).))).)))))))....(----------(......((((((((......))))))---))....-----------..)).. ( -21.10) >DroYak_CAF1 155115 98 - 1 CAUAACAAUGGA---GUGCUCAAGAGC----AACUUAAUGGCACAACAUUGUCUUGU----------CCGAAAUGCUAAUGCUAAUUUGCAUUG---GUUGCG--CUAAAACUUUGGAGU ((..((((((..---(((((.(((...----..)))...)))))..))))))..))(----------(((((..((((((((......))))))---))....--.......)))))).. ( -27.42) >DroMoj_CAF1 99220 115 - 1 CAUAACAAUGCAGA-GCGUUCCAGAGCUUC-AUCUUAAUGUUAUACCAUUGUCUAGGAGCAGAGCGCACAAAAUGCUAAUGUUCAUUUGCAUUACGAGUUGUGG-CCAAAAGUU--ACGU ((((((((((((((-((((((....(((((-....(((((......)))))....))))).)))))).......((....))...))))))))....)))))).-.........--.... ( -27.60) >DroAna_CAF1 50631 99 - 1 CAUAACAAUGGG---G-GCCUGAGAGC----AACUUAAUGGCACAACAUUGUCUUCG----------CCGAAAUGCUAAUGCUAAUUUGCAUUG---GUUGCGCACUCAAACUUUCGAGU ....((((((..---(-(((((((...----..))))..))).)..))))))....(----------(.(....((((((((......))))))---))..)))((((........)))) ( -26.90) >consensus CAUAACAAUGGA___GUGCUCAAGAGC____AACUUAAUGGCACAACAUUGUCUUGG__________CCAAAAUGCUAAUGCUAAUUUGCAUUA___GUUGCG__CUAAAACUUU_GAGU ....((((((.......(((...(((.......)))...)))....))))))................((.(((..((((((......))))))...))).))................. (-10.48 = -9.48 + -1.00)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:00:35 2006