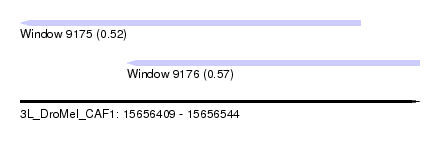
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 15,656,409 – 15,656,544 |
| Length | 135 |
| Max. P | 0.568961 |

| Location | 15,656,409 – 15,656,524 |
|---|---|
| Length | 115 |
| Sequences | 6 |
| Columns | 119 |
| Reading direction | reverse |
| Mean pairwise identity | 95.34 |
| Mean single sequence MFE | -27.08 |
| Consensus MFE | -24.20 |
| Energy contribution | -23.98 |
| Covariance contribution | -0.22 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.12 |
| Structure conservation index | 0.89 |
| SVM decision value | -0.02 |
| SVM RNA-class probability | 0.521825 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 15656409 115 - 23771897 UCUCGUUGUUUAUUUUUAUGAGCGUUGAGUUCGGUGGCCGCCUCACUCAACAUAUUUUGACUUCUUUCAUUGAUAAAGUUAUCAGUCAGCCUUCGGCAGUAUUUGUUUGUGC----UCG ...................(((((((((((..(((....)))..)))))))........(((......(((((((....)))))))..(((...))))))..........))----)). ( -26.50) >DroSec_CAF1 16160 119 - 1 UCUUGUUGUUUAUUUUUAUGAGCGUUGAGUUCGGUGGCCGCCUCACUCAACAUAUUUUGACUUCUUUCAUUGAUAAAGUUAUCAGUCAGCCUUCGGCAGUAUUUGUUUGUGCUCGCUCG ...................(((((((((((..(((....)))..))))))).................(((((((....)))))))..(((...)))(((((......))))).)))). ( -28.10) >DroSim_CAF1 16146 119 - 1 UCUUGUUGUUUAUUUUUAUGAGCGUUGAGUUCGGUGGCCGCCUCACUCAACAUAUUUUGACUUCUUUCAUUGAUAAAGUUAUCAGUCAGCCUUCGGCAGUAUUUGUUUGUGCUCGCUCG ...................(((((((((((..(((....)))..))))))).................(((((((....)))))))..(((...)))(((((......))))).)))). ( -28.10) >DroEre_CAF1 16324 115 - 1 UCUUGUUGUUUAUUUUUAUGAGCGUUGAGUUCGGUGGCCGCCUCACUCAACAUAUUUUGACUUCUUUCAUUGAUAAAGUUAUCAGUCAGCCUUCGGCAGCAUUUGUUUGUGC----UCG ....(((((((((....))))))(((((((..(((....)))..))))))).......))).......(((((((....)))))))..(((...)))(((((......))))----).. ( -27.80) >DroYak_CAF1 16195 115 - 1 UCUUGUUGUUUAUUUUUAUGAGCGUUGAGUUCGGUGGCCGCCUCACUCAACAUAUUUUGACUUCUUUCAUUGAUAAAGUUAUCAGUCAGCCUUCGGCAGUAUUUGUUUGUGC----UCG ...................(((((((((((..(((....)))..)))))))........(((......(((((((....)))))))..(((...))))))..........))----)). ( -26.50) >DroAna_CAF1 75736 115 - 1 UCUUGUUGUUUAUUUUUAUGAGCCCUGAGUUCGGUGGCCGCCGCGCUCAACAUAUUUUGACUUCUUUCAUUGAUAAAGUUAUCAGUCAGCCUUCGGCGGUGUUUGUUUGCGC----UUG ...((((((((((....))))))...((((.((((....)))).))))))))................(((((((....)))))))..(((...)))(((((......))))----).. ( -25.50) >consensus UCUUGUUGUUUAUUUUUAUGAGCGUUGAGUUCGGUGGCCGCCUCACUCAACAUAUUUUGACUUCUUUCAUUGAUAAAGUUAUCAGUCAGCCUUCGGCAGUAUUUGUUUGUGC____UCG ....(((((((((....))))))(((((((..(((....)))..))))))).......))).......(((((((....)))))))..(((...))).((((......))))....... (-24.20 = -23.98 + -0.22)


| Location | 15,656,445 – 15,656,544 |
|---|---|
| Length | 99 |
| Sequences | 6 |
| Columns | 119 |
| Reading direction | reverse |
| Mean pairwise identity | 89.40 |
| Mean single sequence MFE | -20.30 |
| Consensus MFE | -18.11 |
| Energy contribution | -18.08 |
| Covariance contribution | -0.03 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.30 |
| Structure conservation index | 0.89 |
| SVM decision value | 0.07 |
| SVM RNA-class probability | 0.568961 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 15656445 99 - 23771897 UUCCGUGGCUCC--------------------UUAUUCAUUCUCGUUGUUUAUUUUUAUGAGCGUUGAGUUCGGUGGCCGCCUCACUCAACAUAUUUUGACUUCUUUCAUUGAUAAAGU ....((((((((--------------------..(((((..(((((...........)))))...)))))..)).))))))...........((((.(((......)))..)))).... ( -20.20) >DroSec_CAF1 16200 99 - 1 UUCCGUGACUCC--------------------UUAUUCAUUCUUGUUGUUUAUUUUUAUGAGCGUUGAGUUCGGUGGCCGCCUCACUCAACAUAUUUUGACUUCUUUCAUUGAUAAAGU .((.((((....--------------------...............((((((....))))))(((((((..(((....)))..)))))))...............)))).))...... ( -19.70) >DroSim_CAF1 16186 99 - 1 UUCCGUGGCUCC--------------------UUAUUCAUUCUUGUUGUUUAUUUUUAUGAGCGUUGAGUUCGGUGGCCGCCUCACUCAACAUAUUUUGACUUCUUUCAUUGAUAAAGU .((.((((....--------------------...............((((((....))))))(((((((..(((....)))..)))))))...............)))).))...... ( -18.80) >DroEre_CAF1 16360 99 - 1 UUCCGUGGCUCC--------------------UUAUUCAUUCUUGUUGUUUAUUUUUAUGAGCGUUGAGUUCGGUGGCCGCCUCACUCAACAUAUUUUGACUUCUUUCAUUGAUAAAGU .((.((((....--------------------...............((((((....))))))(((((((..(((....)))..)))))))...............)))).))...... ( -18.80) >DroYak_CAF1 16231 99 - 1 UUCCGUGGCUCC--------------------UUAUUCAUUCUUGUUGUUUAUUUUUAUGAGCGUUGAGUUCGGUGGCCGCCUCACUCAACAUAUUUUGACUUCUUUCAUUGAUAAAGU .((.((((....--------------------...............((((((....))))))(((((((..(((....)))..)))))))...............)))).))...... ( -18.80) >DroAna_CAF1 75772 119 - 1 CUCAGUGAUUUAUUGAUGGCAGUGCAUUAUUUCUCUUCAUUCUUGUUGUUUAUUUUUAUGAGCCCUGAGUUCGGUGGCCGCCGCGCUCAACAUAUUUUGACUUCUUUCAUUGAUAAAGU .(((((((.....(((((......))))).......(((....((((((((((....))))))...((((.((((....)))).)))))))).....)))......)))))))...... ( -25.50) >consensus UUCCGUGGCUCC____________________UUAUUCAUUCUUGUUGUUUAUUUUUAUGAGCGUUGAGUUCGGUGGCCGCCUCACUCAACAUAUUUUGACUUCUUUCAUUGAUAAAGU .((.((((.......................................((((((....))))))(((((((..(((....)))..)))))))...............)))).))...... (-18.11 = -18.08 + -0.03)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:59:38 2006