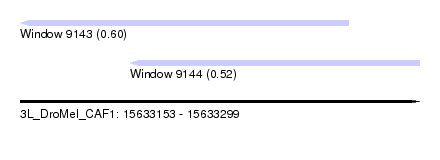
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 15,633,153 – 15,633,299 |
| Length | 146 |
| Max. P | 0.595477 |

| Location | 15,633,153 – 15,633,273 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 95.38 |
| Mean single sequence MFE | -37.00 |
| Consensus MFE | -30.30 |
| Energy contribution | -31.18 |
| Covariance contribution | 0.88 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.91 |
| Structure conservation index | 0.82 |
| SVM decision value | 0.12 |
| SVM RNA-class probability | 0.595477 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 15633153 120 - 23771897 GAUUCCCGGACUCCUUGCUGUUUGUUUUUAAUAUAAAUGCAAAUUUCGCCAACGGCAUUCGCCCGGAGCCGUGCUCCAUAGUACAUAUGUGUGUAUGCGAGCCUGUUCCGGAGAUCUGUC (((..(((((....((((.((((((.......))))))))))...........(((..((((..(((((...)))))...((((((....)))))))))))))...)))))..))).... ( -38.20) >DroSec_CAF1 55424 118 - 1 GAUUCCCGGACUCCUUGCUGUUUGUUUUUAAUAUAAAUGCAAAUUUCGACAACGGCAUUCGCCAGGCGCCAUGCUCCCUAGUACAUAUGUGUG--UGCAAGCCUGUGCCGGAGAUCUGUC ......((((((((((((.((((((.......))))))))))...........((((.....(((((((...))......((((((....)))--)))..))))))))))))).)))).. ( -34.50) >DroSim_CAF1 56381 118 - 1 GAUUCACGGACUCCUUGCUGUUUGUUUUUAAUAUAAAUGCAAAUUUCGCCAACGGCAUUCGCCAGGCGCCAUGCUCCAUAGUACAUAUGUGUG--UGCGAGCCUGUUCCGGAGAUCUGUC .....(((((((((((((.((((((.......))))))))))....((((...(((....))).)))).((.((((((((.(((....)))))--)).)))).))....)))).))))). ( -35.90) >DroYak_CAF1 60144 118 - 1 GAUUCCCGGACUCCUUGCUGUUUGUUUUUAAUAUAAAUGCAAAUUUCGCCAACGGCAUUCGCCAGGCGCCAUGCUCCACAGUACAUAUGUGUG--UGCGAGCCUGUUCCGGAGAUCUGUC (((..(((((....((((.((((((.......))))))))))....((((...(((....))).)))).((.((((((((.(((....)))))--)).)))).)).)))))..))).... ( -39.40) >consensus GAUUCCCGGACUCCUUGCUGUUUGUUUUUAAUAUAAAUGCAAAUUUCGCCAACGGCAUUCGCCAGGCGCCAUGCUCCAUAGUACAUAUGUGUG__UGCGAGCCUGUUCCGGAGAUCUGUC ......((((((((((((.((((((.......))))))))))...........(((....))).(((((...((((((...((((....))))..)).))))..))))))))).)))).. (-30.30 = -31.18 + 0.88)

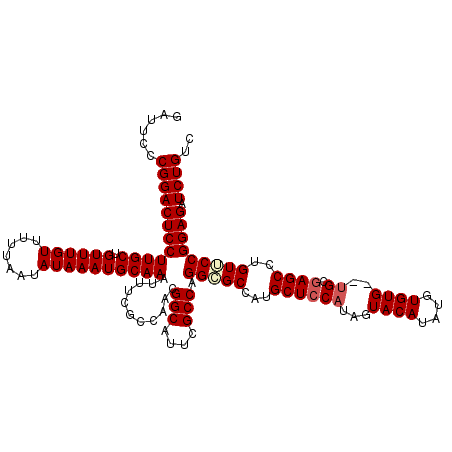
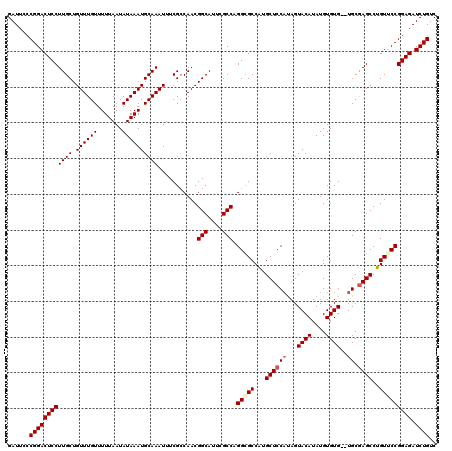
| Location | 15,633,193 – 15,633,299 |
|---|---|
| Length | 106 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 90.71 |
| Mean single sequence MFE | -31.80 |
| Consensus MFE | -25.01 |
| Energy contribution | -25.08 |
| Covariance contribution | 0.06 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.45 |
| Structure conservation index | 0.79 |
| SVM decision value | -0.04 |
| SVM RNA-class probability | 0.515420 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 15633193 106 - 23771897 GCUUCUGGUUUCUGGCUUCUG--------------GCCGAGAUUCCCGGACUCCUUGCUGUUUGUUUUUAAUAUAAAUGCAAAUUUCGCCAACGGCAUUCGCCCGGAGCCGUGCUCCAUA ((....((..((((((.....--------------))).)))..))(((.((((((((.((((((.......))))))))))...........(((....))).))))))).))...... ( -30.50) >DroSec_CAF1 55462 106 - 1 GCUUCUGGUUUCUGGCUUCUG--------------GCCGAGAUUCCCGGACUCCUUGCUGUUUGUUUUUAAUAUAAAUGCAAAUUUCGACAACGGCAUUCGCCAGGCGCCAUGCUCCCUA ((...((((.((((((.....--------------((((((.((....))))).((((.((((((.......))))))))))...........)))....)))))).)))).))...... ( -29.10) >DroSim_CAF1 56419 106 - 1 GCUUCUGGUUUCUGGCUUCUG--------------GCCGAGAUUCACGGACUCCUUGCUGUUUGUUUUUAAUAUAAAUGCAAAUUUCGCCAACGGCAUUCGCCAGGCGCCAUGCUCCAUA .....(((....((((.((((--------------((.((((((...((...)).(((.((((((.......)))))))))))))))(((...)))....)))))).))))....))).. ( -29.70) >DroYak_CAF1 60182 120 - 1 GCUUCUGGUUUCUGGCUUCUGGCUGCUGGAUGCUGGCCGAGAUUCCCGGACUCCUUGCUGUUUGUUUUUAAUAUAAAUGCAAAUUUCGCCAACGGCAUUCGCCAGGCGCCAUGCUCCACA .....(((....((((.((((((....((((((((..(((((((...((...)).(((.((((((.......))))))))))))))))....)))))))))))))).))))....))).. ( -37.90) >consensus GCUUCUGGUUUCUGGCUUCUG______________GCCGAGAUUCCCGGACUCCUUGCUGUUUGUUUUUAAUAUAAAUGCAAAUUUCGCCAACGGCAUUCGCCAGGCGCCAUGCUCCAUA ((...((((.((((((....................(((.......)))......((((((((((.((((...)))).))).........)))))))...)))))).)))).))...... (-25.01 = -25.08 + 0.06)


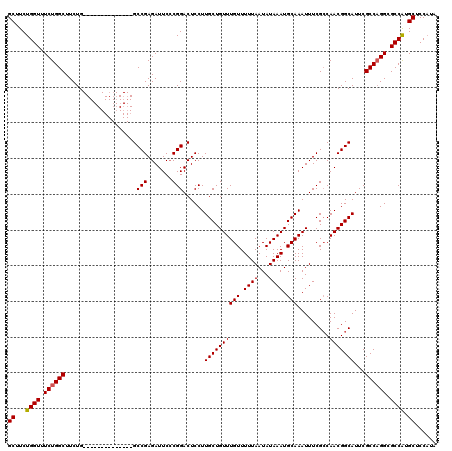
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:59:07 2006