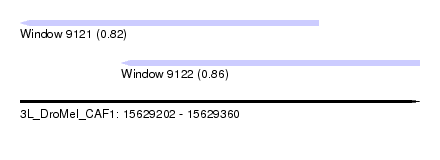
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 15,629,202 – 15,629,360 |
| Length | 158 |
| Max. P | 0.859277 |

| Location | 15,629,202 – 15,629,320 |
|---|---|
| Length | 118 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 94.67 |
| Mean single sequence MFE | -43.65 |
| Consensus MFE | -35.90 |
| Energy contribution | -35.40 |
| Covariance contribution | -0.50 |
| Combinations/Pair | 1.12 |
| Mean z-score | -2.28 |
| Structure conservation index | 0.82 |
| SVM decision value | 0.69 |
| SVM RNA-class probability | 0.823571 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 15629202 118 - 23771897 AAAACGUGUGCCCAUCUAUUGGGG-GGGUUUUUUGUGGGGUGUUUCGCAGUUAUGGAAGGGGGAG-CGCUCCAUAAAAGGGGGUGGUUCGCUGCACUGGGUGGGUUUCGGUGGUUUGGCC ....((...(((((((((.((.((-.((((((((((((((((((((.(..((...))..).))))-)))))))))))))).......)).)).)).)))))))))..))........... ( -42.91) >DroSec_CAF1 51431 117 - 1 AAAACGUGUGCCCAUCUAUUGGG--GGGUUUUUUGUGGGGUGUUUCGCAGUUAUGGAAUGGGGAG-CGCUCCAUAAAAGGGGGUGGUUCACUGCACUGGGUGGGUUUCGGUGGUUUGGCC ....((...(((((((((.((.(--(..((((((((((((((((((.((.........)).))))-)))))))))))))).((....)).)).)).)))))))))..))........... ( -43.20) >DroSim_CAF1 52438 119 - 1 AAAACGUGUGCCCAUCUAUUGGGGGGGGUUUUUUGUGGGGUGCUUCGCAGUUGUGGAAUGGGGAG-CGCUCCAUAAAAGGGGGUGGUUCACUGCACUGGGUGGGUUUUGGUGGUUUGGCC ....((...(((((((((.((.((.(..((((((((((((((((((.((.........)).))))-)))))))))))))......)..).)).)).)))))))))..))........... ( -44.30) >DroYak_CAF1 55998 118 - 1 AAAACGUGUGCCCAUCUAUUGGG--GGGUUUUUUGUGGGGUGCUUUGCAGUUAUGGAACGGGGGGCCGCUCCAUAAAAGGGGGUGGUUCCCUGCACUGGGUGGGUUUGGGUGGUUUGGCC ...((.((.(((((((((.((..--.......(((..((....))..)))........((((((((((((((.......)))))))))))))))).))))))))).)).))......... ( -44.20) >consensus AAAACGUGUGCCCAUCUAUUGGG__GGGUUUUUUGUGGGGUGCUUCGCAGUUAUGGAAUGGGGAG_CGCUCCAUAAAAGGGGGUGGUUCACUGCACUGGGUGGGUUUCGGUGGUUUGGCC ....((...(((((((((..........((((((((((((((((((.(...........).)))).))))))))))))))..(..(....)..)..)))))))))..))........... (-35.90 = -35.40 + -0.50)

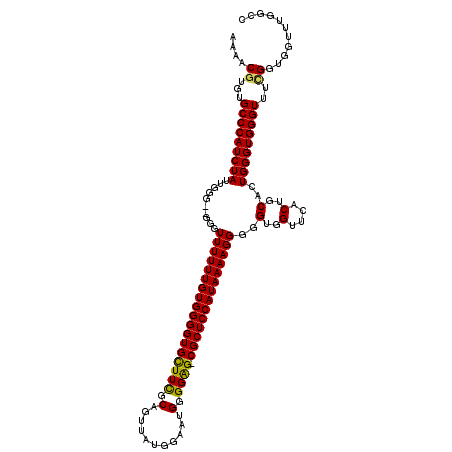
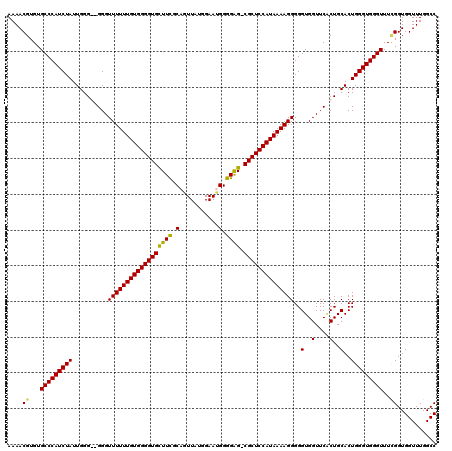
| Location | 15,629,242 – 15,629,360 |
|---|---|
| Length | 118 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 95.65 |
| Mean single sequence MFE | -40.30 |
| Consensus MFE | -35.83 |
| Energy contribution | -35.45 |
| Covariance contribution | -0.38 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.02 |
| Structure conservation index | 0.89 |
| SVM decision value | 0.82 |
| SVM RNA-class probability | 0.859277 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 15629242 118 - 23771897 UAUUACAUGCGGAUUAGCAUUUCCUCAUCGGGAUUGCAACAAAACGUGUGCCCAUCUAUUGGGG-GGGUUUUUUGUGGGGUGUUUCGCAGUUAUGGAAGGGGGAG-CGCUCCAUAAAAGG .....((((((..((.(((.((((.....)))).)))...))..))))))((((.....)))).-....(((((((((((((((((.(..((...))..).))))-))))))))))))). ( -40.30) >DroSec_CAF1 51471 117 - 1 UAUUACAUGCGGAUUAGCAUUUCCUCAUCGGUAUUGCAACAAAACGUGUGCCCAUCUAUUGGG--GGGUUUUUUGUGGGGUGUUUCGCAGUUAUGGAAUGGGGAG-CGCUCCAUAAAAGG .....((((((..((.(((...((.....))...)))...))..))))))((((.....))))--....(((((((((((((((((.((.........)).))))-))))))))))))). ( -38.90) >DroSim_CAF1 52478 119 - 1 UAUUACAUGCGGAUUAGCAUUUCCUCAUCGGGAUUGCAACAAAACGUGUGCCCAUCUAUUGGGGGGGGUUUUUUGUGGGGUGCUUCGCAGUUGUGGAAUGGGGAG-CGCUCCAUAAAAGG .....((((((..((.(((.((((.....)))).)))...))..))))))((((.....))))......(((((((((((((((((.((.........)).))))-))))))))))))). ( -44.20) >DroYak_CAF1 56038 118 - 1 UAUUACAUGCGGAUUAGCAUUUCCUCAUCGGGAUUGCAACAAAACGUGUGCCCAUCUAUUGGG--GGGUUUUUUGUGGGGUGCUUUGCAGUUAUGGAACGGGGGGCCGCUCCAUAAAAGG .....((((((..((.(((.((((.....)))).)))...))..))))))((((.....))))--....(((((((((((((((((.(.(((....)))).)))).))))))))))))). ( -37.80) >consensus UAUUACAUGCGGAUUAGCAUUUCCUCAUCGGGAUUGCAACAAAACGUGUGCCCAUCUAUUGGG__GGGUUUUUUGUGGGGUGCUUCGCAGUUAUGGAAUGGGGAG_CGCUCCAUAAAAGG .....((((((..((.(((.((((.....)))).)))...))..))))))((((.....))))......(((((((((((((((((.(...........).)))).))))))))))))). (-35.83 = -35.45 + -0.38)

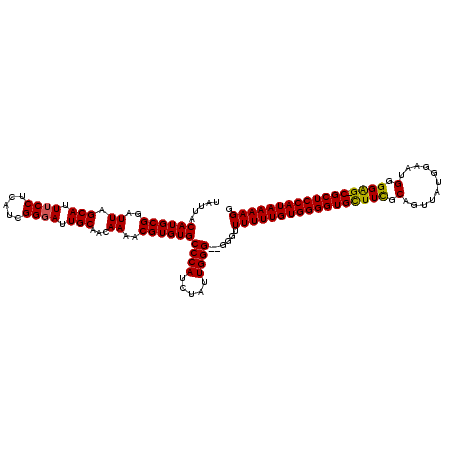
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:58:46 2006