| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
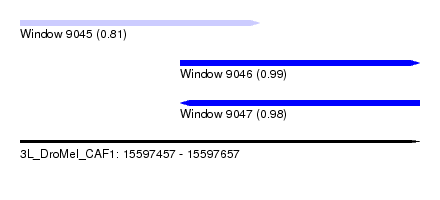
| Location | 15,597,457 – 15,597,657 |
| Length | 200 |
| Max. P | 0.994916 |

| Location | 15,597,457 – 15,597,577 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 96.94 |
| Mean single sequence MFE | -45.68 |
| Consensus MFE | -45.41 |
| Energy contribution | -45.41 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.73 |
| Structure conservation index | 0.99 |
| SVM decision value | 0.65 |
| SVM RNA-class probability | 0.810509 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 15597457 120 + 23771897 CGAAUCCAAGCCCAAGGCCAGCUGCUCCAGUUUGUUAAUGAGUAUGGAGUAGUUGCCGGGCAGAUCGAAGCCGCCAAGAUCGAGUUUUUCGCCACUACUGGUGGAUGCUGCGGCAUCCAA (((.((...((((..(((.((((((((((...............))))))))))))))))).)))))..(((((........(((..(((((((....))))))).))))))))...... ( -45.66) >DroSec_CAF1 20717 120 + 1 CGAAUCCAAGCCCAAGGCCAGCUGCUCCAGUUUGUUAAUGAGUAUGGAGUAGUUGCCGGGCAGAUCGAAGCCGCCAAGAUCGAGUUUUUCGCCACUACUGGUGGAUGCUGCGGCAUCCAA (((.((...((((..(((.((((((((((...............))))))))))))))))).)))))..(((((........(((..(((((((....))))))).))))))))...... ( -45.66) >DroEre_CAF1 20713 120 + 1 CGAGUCCAAGCCCAGGGCCAGCUGCUCCAGUUUGUUAAUGAGUAUGGAGUAGUUGCCGGGCAGAUCGAAGCCGCCCAGAUCGAGUUUUUCGCCACUAUUGGUGGAUGCUGCGGCGUCCAA (((.((...((((..(((.((((((((((...............))))))))))))))))).)))))..(((((........(((..(((((((....))))))).))))))))...... ( -45.66) >DroYak_CAF1 22111 120 + 1 CGAGUCCAAGCCCAAGGCCAGCUGCUCCAGUUUGUUAAUGAGUAUGGAGUAGUUGCCGGGCAGAUCGAAGCCGCCCAGAUCGAGUUUUUCGCCACUAUUGGUGGACGCUGCGGCGUCCAA .....((((((((..(((.((((((((((...............))))))))))))))))).((((...........))))................))))(((((((....))))))). ( -45.76) >consensus CGAAUCCAAGCCCAAGGCCAGCUGCUCCAGUUUGUUAAUGAGUAUGGAGUAGUUGCCGGGCAGAUCGAAGCCGCCAAGAUCGAGUUUUUCGCCACUACUGGUGGAUGCUGCGGCAUCCAA (((.((...((((..(((.((((((((((...............))))))))))))))))).)))))..(((((........(((..(((((((....))))))).))))))))...... (-45.41 = -45.41 + 0.00)



| Location | 15,597,537 – 15,597,657 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 94.03 |
| Mean single sequence MFE | -45.35 |
| Consensus MFE | -45.55 |
| Energy contribution | -44.67 |
| Covariance contribution | -0.87 |
| Combinations/Pair | 1.14 |
| Mean z-score | -2.48 |
| Structure conservation index | 1.00 |
| SVM decision value | 2.52 |
| SVM RNA-class probability | 0.994916 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 15597537 120 + 23771897 CGAGUUUUUCGCCACUACUGGUGGAUGCUGCGGCAUCCAAGGUCGUUGCAUUGCCAUUAAGGCCUUCACUUAAAGUUGUGGUUGUGGCAGUUGCAAUCGUUAGAUUGCCGGCAUCUUCGA ((((......((.((.(((..(((((((....))))))).))).)).))..((((......(((.((((........))))....)))....((((((....)))))).))))..)))). ( -42.30) >DroSec_CAF1 20797 120 + 1 CGAGUUUUUCGCCACUACUGGUGGAUGCUGCGGCAUCCAAGGUCGUUGCAUUGCCAUUGAGACCUUCGCUUAAAGUUGUGGUUGUGGCAGUUGCAAUCGUUAGAUUGCCGGCAUCUUCGA ((((.....(((((....)))))(((((..(((((((.((.(..(((((((((((((...((((...((.....))...))))))))))).))))))).)).)).)))))))))))))). ( -42.80) >DroEre_CAF1 20793 120 + 1 CGAGUUUUUCGCCACUAUUGGUGGAUGCUGCGGCGUCCAAGGUCGUCGCAUUGCCAUUAAGGCCUUCACUUAAAGUUGUGGCUGUGGCAGUUGCAAUCGUCAGAUUGCCGGCAUCUUCGA ((((.....(((((....)))))((((((((((((.(....).))))))((((((((...((((...((.....))...)))))))))))).((((((....)))))).)))))))))). ( -49.50) >DroYak_CAF1 22191 120 + 1 CGAGUUUUUCGCCACUAUUGGUGGACGCUGCGGCGUCCAAGGUCGUCGCAUUGCCAUUGAGGCCUUCACUUAGAGUUGUGGUUGUGGCAGUUGCAAUCGUUAGAUUGCUGGUGUCUUCGA ((((.....(((((....)))))((((((((((((.(....).))))))((((((((...((((...((.....))...)))))))))))).((((((....)))))).)))))))))). ( -46.80) >consensus CGAGUUUUUCGCCACUACUGGUGGAUGCUGCGGCAUCCAAGGUCGUCGCAUUGCCAUUAAGGCCUUCACUUAAAGUUGUGGUUGUGGCAGUUGCAAUCGUUAGAUUGCCGGCAUCUUCGA ((((.....(((((....)))))((((((((((((.(....).))))))((((((((...((((...((.....))...)))))))))))).((((((....)))))).)))))))))). (-45.55 = -44.67 + -0.87)



| Location | 15,597,537 – 15,597,657 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 94.03 |
| Mean single sequence MFE | -36.05 |
| Consensus MFE | -34.95 |
| Energy contribution | -34.58 |
| Covariance contribution | -0.38 |
| Combinations/Pair | 1.11 |
| Mean z-score | -2.14 |
| Structure conservation index | 0.97 |
| SVM decision value | 1.82 |
| SVM RNA-class probability | 0.978857 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 15597537 120 - 23771897 UCGAAGAUGCCGGCAAUCUAACGAUUGCAACUGCCACAACCACAACUUUAAGUGAAGGCCUUAAUGGCAAUGCAACGACCUUGGAUGCCGCAGCAUCCACCAGUAGUGGCGAAAAACUCG .(((........((((((....))))))...((((((.(((((........)))...(((.....))).............(((((((....)))))))...)).))))))......))) ( -36.40) >DroSec_CAF1 20797 120 - 1 UCGAAGAUGCCGGCAAUCUAACGAUUGCAACUGCCACAACCACAACUUUAAGCGAAGGUCUCAAUGGCAAUGCAACGACCUUGGAUGCCGCAGCAUCCACCAGUAGUGGCGAAAAACUCG .(((........((((((....))))))...((((((.((..............((((((......((...))...))))))((((((....))))))....)).))))))......))) ( -36.20) >DroEre_CAF1 20793 120 - 1 UCGAAGAUGCCGGCAAUCUGACGAUUGCAACUGCCACAGCCACAACUUUAAGUGAAGGCCUUAAUGGCAAUGCGACGACCUUGGACGCCGCAGCAUCCACCAAUAGUGGCGAAAAACUCG (((..(((((..((((((....))))))...(((((..(((...((.....))...))).....))))).((((.((.(....).)).)))))))))(((.....))).)))........ ( -33.80) >DroYak_CAF1 22191 120 - 1 UCGAAGACACCAGCAAUCUAACGAUUGCAACUGCCACAACCACAACUCUAAGUGAAGGCCUCAAUGGCAAUGCGACGACCUUGGACGCCGCAGCGUCCACCAAUAGUGGCGAAAAACUCG .(((........((((((....))))))...((((((...(((........)))...(((.....))).............(((((((....)))))))......))))))......))) ( -37.80) >consensus UCGAAGAUGCCGGCAAUCUAACGAUUGCAACUGCCACAACCACAACUUUAAGUGAAGGCCUCAAUGGCAAUGCAACGACCUUGGACGCCGCAGCAUCCACCAAUAGUGGCGAAAAACUCG .(((........((((((....))))))...((((((..............(((...(((.....)))..)))........(((((((....)))))))......))))))......))) (-34.95 = -34.58 + -0.38)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:57:31 2006