
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 15,496,674 – 15,496,834 |
| Length | 160 |
| Max. P | 0.961205 |

| Location | 15,496,674 – 15,496,794 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 96.11 |
| Mean single sequence MFE | -32.03 |
| Consensus MFE | -29.41 |
| Energy contribution | -29.30 |
| Covariance contribution | -0.11 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.15 |
| Structure conservation index | 0.92 |
| SVM decision value | 0.74 |
| SVM RNA-class probability | 0.838571 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 15496674 120 - 23771897 CAGCUGUGCCUGUGGCUUUUGUGUGUUCCUCGGAAAAAUACUAGGAAAAUGCAUAAAAACAUAUGCCAUGGCAUCUAAAAAUAUGUGCUCGUGUUUAGCUAUUUCGAUAAACGAUGGGCA .....(((((.(((((((((((((.(((((.(........).)))))...))))))))......))))))))))...........((((((((((((.(......).))))).))))))) ( -34.10) >DroSec_CAF1 37841 120 - 1 CAGCUGUGCCUGUGGCUUUUGUGUUUUCCUCGGAAAAAUACUAGGAAAAUGCAUAAAAACAUAUGCCAUGGCAUCUAAAAAUAUGUGCUCGUAUUCAGCUAUUUCGAUAAACGAUGGCCA .(((((((((.(((((...(((((((((((.(........).)))))))))))...........)))))))))......((((((....)))))))))))...(((.....)))...... ( -32.44) >DroSim_CAF1 38039 118 - 1 CAGCUGUGCCUGUGGCUUUUGUGUUUUCCUCGGAA-AAUACUAGGAAAAUGCAUAAAAACAUAUGCCAUGGCAUCUAAAAAUAUGUGCUCGUAUUCAACUAUUUCGAUAAACGAUGGC-A ..((((((((.(((((...(((((((((((.(...-....).)))))))))))...........)))))))))).....((((((....))))))........(((.....))).)))-. ( -29.54) >consensus CAGCUGUGCCUGUGGCUUUUGUGUUUUCCUCGGAAAAAUACUAGGAAAAUGCAUAAAAACAUAUGCCAUGGCAUCUAAAAAUAUGUGCUCGUAUUCAGCUAUUUCGAUAAACGAUGGCCA .(((((((((.(((((((((((((.(((((.(........).)))))...))))))))......)))))))))......((((((....)))))))))))...(((.....)))...... (-29.41 = -29.30 + -0.11)

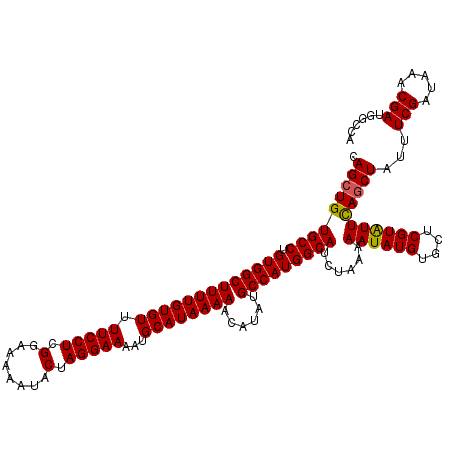

| Location | 15,496,714 – 15,496,834 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 98.89 |
| Mean single sequence MFE | -31.93 |
| Consensus MFE | -31.47 |
| Energy contribution | -31.80 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.41 |
| Structure conservation index | 0.99 |
| SVM decision value | 1.52 |
| SVM RNA-class probability | 0.961205 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 15496714 120 - 23771897 GUUUAAUGAUAUGCUAUUAUUAGCGGCACGUAUGAAAAAGCAGCUGUGCCUGUGGCUUUUGUGUGUUCCUCGGAAAAAUACUAGGAAAAUGCAUAAAAACAUAUGCCAUGGCAUCUAAAA ............((((....))))((((((..((......))..))))))((((..((((((((.(((((.(........).)))))...)))))))).))))(((....)))....... ( -31.30) >DroSec_CAF1 37881 120 - 1 GUUUAAUGAUAUGCUAUUAUUAGCGGCACGUAUGAAAAAGCAGCUGUGCCUGUGGCUUUUGUGUUUUCCUCGGAAAAAUACUAGGAAAAUGCAUAAAAACAUAUGCCAUGGCAUCUAAAA ............((((....))))((((((..((......))..))))))((((..((((((((((((((.(........).))))))))))).)))..))))(((....)))....... ( -32.40) >DroSim_CAF1 38078 119 - 1 GUUUAAUGAUAUGCUAUUAUUAGCGGCACGUAUGAAAAAGCAGCUGUGCCUGUGGCUUUUGUGUUUUCCUCGGAA-AAUACUAGGAAAAUGCAUAAAAACAUAUGCCAUGGCAUCUAAAA ............((((....))))((((((..((......))..))))))((((..((((((((((((((.(...-....).))))))))))).)))..))))(((....)))....... ( -32.10) >consensus GUUUAAUGAUAUGCUAUUAUUAGCGGCACGUAUGAAAAAGCAGCUGUGCCUGUGGCUUUUGUGUUUUCCUCGGAAAAAUACUAGGAAAAUGCAUAAAAACAUAUGCCAUGGCAUCUAAAA ............((((....))))((((((..((......))..))))))((((..((((((((((((((.(........).))))))))))).)))..))))(((....)))....... (-31.47 = -31.80 + 0.33)

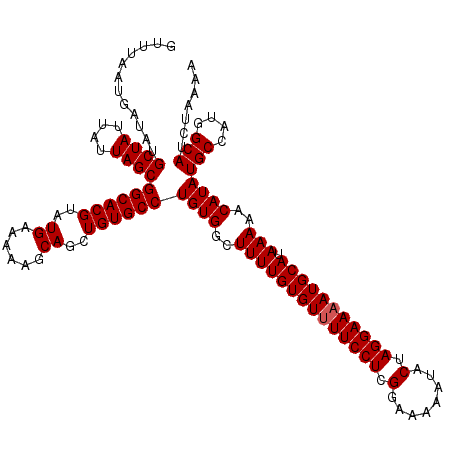

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:55:31 2006