
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 15,446,237 – 15,446,342 |
| Length | 105 |
| Max. P | 0.988666 |

| Location | 15,446,237 – 15,446,342 |
|---|---|
| Length | 105 |
| Sequences | 6 |
| Columns | 118 |
| Reading direction | forward |
| Mean pairwise identity | 78.71 |
| Mean single sequence MFE | -38.90 |
| Consensus MFE | -22.77 |
| Energy contribution | -22.37 |
| Covariance contribution | -0.40 |
| Combinations/Pair | 1.18 |
| Mean z-score | -3.31 |
| Structure conservation index | 0.59 |
| SVM decision value | 2.13 |
| SVM RNA-class probability | 0.988666 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
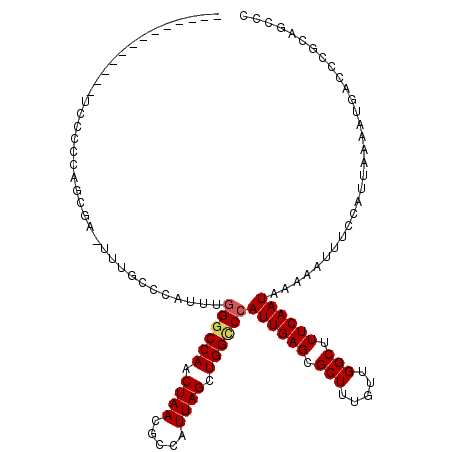
>3L_DroMel_CAF1 15446237 105 + 23771897 GCGGGGUGGGUCAAUUUAAUGGAAAUUUUUAUUGAAAGCCAACAAAGCGCUCAAUGGGCCAGCUAAUGGCGUUAGUUGGCCCAAAUGGGCCAAGUCGCUGGGGGA------------- (((((((...(((((..(((....)))...)))))..)))......(.(((((.(((((((((((((...)))))))))))))..))))))...)))).......------------- ( -40.60) >DroVir_CAF1 172011 94 + 1 AGAAU-GGGGCCAUUUUAAUGGAAAUUUUUAUUGAAAGCCAACAAAGCGCUCAAUGGCCCAGCUAAUGGCGUUAGUUGGCC-AAC-------------UGGAGGA---------ACCA ....(-((..((.((((((((((....))))))))))((((((..(((((.((.((((...)))).))))))).)))))).-...-------------....)).---------.))) ( -31.20) >DroPse_CAF1 174444 116 + 1 GGGCUGCGGGUCAUUUCAGUGGAAAUU-UUAUUGAAAGCCAGCAAAGCGCUCAAUGGACCAGCUAAUGGCGUUAGUUGGCCCAAAUGGUCAAA-AGGCAAGGGCAGACCGAGAGAACU (((((((.(((..((((((((((...)-)))))))))))).))..)))((((..(((.(((((((((...))))))))).)))....(((...-.)))..))))...))......... ( -37.50) >DroEre_CAF1 174397 105 + 1 GCGGGGCGGGUCAAUUUAAUGGAAAUUUUUAUUGAAAGCCAACAAAGCGCUCAAUGGGCCAGCUAAUGGCGUUAGUUGGCCCAAAUGGGCCAAGUCGCUGGGGGA------------- (((((((...(((((..(((....)))...)))))..)))......(.(((((.(((((((((((((...)))))))))))))..))))))...)))).......------------- ( -42.20) >DroAna_CAF1 315872 104 + 1 GGGCGGCGGGUCAUUUUAAUGGAAAUUUUUAUUGAAAGCCAACAAAGCGCUCAAUGGGCCAGCUAAUGGCGUUAGUUGGCCCAAAUGGGCAAAGUCGCUAGGGG-------------- .((((((.(((..((((((((((....)))))))))))))........(((((.(((((((((((((...)))))))))))))..)))))...)))))).....-------------- ( -44.40) >DroPer_CAF1 172564 116 + 1 GGGCUGCGGGUCAUUUCAGUGGAAAUU-UUAUUGAAAGCCAGCAAAGCGCUCAAUGGACCAGCUAAUGGCGUUAGUUGGCCCAAGUGGUCAAA-AGGCAAGGGCAGACCGAGAGAACU (((((((.(((..((((((((((...)-)))))))))))).))..)))((((..(((.(((((((((...))))))))).)))....(((...-.)))..))))...))......... ( -37.50) >consensus GGGCGGCGGGUCAUUUUAAUGGAAAUUUUUAUUGAAAGCCAACAAAGCGCUCAAUGGGCCAGCUAAUGGCGUUAGUUGGCCCAAAUGGGCAAA_UCGCUAGGGGA_____________ ....(((.......(((((((((....))))))))).((((((..(((((.((.(((.....))).))))))).))))))................)))................... (-22.77 = -22.37 + -0.40)



| Location | 15,446,237 – 15,446,342 |
|---|---|
| Length | 105 |
| Sequences | 6 |
| Columns | 118 |
| Reading direction | reverse |
| Mean pairwise identity | 78.71 |
| Mean single sequence MFE | -27.67 |
| Consensus MFE | -18.90 |
| Energy contribution | -19.07 |
| Covariance contribution | 0.17 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.89 |
| Structure conservation index | 0.68 |
| SVM decision value | 1.08 |
| SVM RNA-class probability | 0.911536 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 15446237 105 - 23771897 -------------UCCCCCAGCGACUUGGCCCAUUUGGGCCAACUAACGCCAUUAGCUGGCCCAUUGAGCGCUUUGUUGGCUUUCAAUAAAAAUUUCCAUUAAAUUGACCCACCCCGC -------------.......(((...(((..((...((((((.((((.....)))).))))))((((((.(((.....))).)))))).................))..)))...))) ( -27.60) >DroVir_CAF1 172011 94 - 1 UGGU---------UCCUCCA-------------GUU-GGCCAACUAACGCCAUUAGCUGGGCCAUUGAGCGCUUUGUUGGCUUUCAAUAAAAAUUUCCAUUAAAAUGGCCCC-AUUCU (((.---------.((((((-------------(((-(((........))))...))))))..((((((.(((.....))).))))))..................))..))-).... ( -24.10) >DroPse_CAF1 174444 116 - 1 AGUUCUCUCGGUCUGCCCUUGCCU-UUUGACCAUUUGGGCCAACUAACGCCAUUAGCUGGUCCAUUGAGCGCUUUGCUGGCUUUCAAUAA-AAUUUCCACUGAAAUGACCCGCAGCCC .((((....((((.((....))..-...))))...(((((((.((((.....)))).)))))))..))))(((..((.((((((((....-.........))))).).)).))))).. ( -29.52) >DroEre_CAF1 174397 105 - 1 -------------UCCCCCAGCGACUUGGCCCAUUUGGGCCAACUAACGCCAUUAGCUGGCCCAUUGAGCGCUUUGUUGGCUUUCAAUAAAAAUUUCCAUUAAAUUGACCCGCCCCGC -------------.......(((((..((((((..(((((((.((((.....)))).))))))).)).).)))..)).(((..(((((...............)))))...))).))) ( -28.36) >DroAna_CAF1 315872 104 - 1 --------------CCCCUAGCGACUUUGCCCAUUUGGGCCAACUAACGCCAUUAGCUGGCCCAUUGAGCGCUUUGUUGGCUUUCAAUAAAAAUUUCCAUUAAAAUGACCCGCCGCCC --------------......(((.....((.((..(((((((.((((.....)))).))))))).)).))(((((((((.....)))))))......(((....)))....))))).. ( -26.90) >DroPer_CAF1 172564 116 - 1 AGUUCUCUCGGUCUGCCCUUGCCU-UUUGACCACUUGGGCCAACUAACGCCAUUAGCUGGUCCAUUGAGCGCUUUGCUGGCUUUCAAUAA-AAUUUCCACUGAAAUGACCCGCAGCCC .((((....((((.((....))..-...))))...(((((((.((((.....)))).)))))))..))))(((..((.((((((((....-.........))))).).)).))))).. ( -29.52) >consensus _____________UCCCCCAGCGA_UUUGCCCAUUUGGGCCAACUAACGCCAUUAGCUGGCCCAUUGAGCGCUUUGUUGGCUUUCAAUAAAAAUUUCCAUUAAAAUGACCCGCAGCCC ....................................((((((.((((.....)))).))))))((((((.(((.....))).)))))).............................. (-18.90 = -19.07 + 0.17)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:54:58 2006