
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 15,337,128 – 15,337,273 |
| Length | 145 |
| Max. P | 0.836089 |

| Location | 15,337,128 – 15,337,237 |
|---|---|
| Length | 109 |
| Sequences | 5 |
| Columns | 109 |
| Reading direction | forward |
| Mean pairwise identity | 92.00 |
| Mean single sequence MFE | -32.88 |
| Consensus MFE | -28.12 |
| Energy contribution | -28.12 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.48 |
| Structure conservation index | 0.86 |
| SVM decision value | 0.73 |
| SVM RNA-class probability | 0.836089 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 15337128 109 + 23771897 AGGGCAUGGCGAGGCUCAGUUUGAGAACUGUUCAGCUCAGCUUCUUCGUGGCUUAGAAGCCACUGCCACAGCAUGUUAACUAGAGCACUGAGCAAAAUUUAAUUAAUGU ...((.((((((((((.((((.(((.....))))))).))))))...((((((....)))))).))))(((..((((......))))))).))................ ( -32.40) >DroSec_CAF1 69185 109 + 1 AGGGCAUGGCGAGGCUCAGUUUGAGAACUGUUCAGCUCAGCUUCUUCGUGGCUUAGGAGCCACUGCCACAGCAUGUUAACUAGAGCACUGAGCAAAAUUUAAUUAAUGU ...((.((((((((((.((((.(((.....))))))).))))))...((((((....)))))).))))(((..((((......))))))).))................ ( -32.60) >DroSim_CAF1 70586 109 + 1 AGGGCAUGGCGAGGCUCAGUUUGAGAACUGUUCAGCUCAGCUUCUUCGUGGCUUAGGAGCCACUGCCACAGCAUGUUAACUAGAGCACUGAGCAAAAUUUAAUUAAUGU ...((.((((((((((.((((.(((.....))))))).))))))...((((((....)))))).))))(((..((((......))))))).))................ ( -32.60) >DroEre_CAF1 69485 107 + 1 GGGGCGUGGCGAGGCUCAGU-UGAGAACUGUUCAGCUCAGCUUCUUCGUGGCUUAGGAGCCUCUGCCACAGCAUGUUAACUAGAACACUGAACGAAAUUUA-AAUACGC ...(((((((((((((.(((-((((.....))))))).))))))...(((((..((....))..))))).)).((((......))))..............-..))))) ( -35.00) >DroYak_CAF1 71612 107 + 1 GGGGCAUGGCGAGGCUCAGU-UGAGAACUGUUCAGCUCAGCUUCUUCGUGGCUUAGGAGUCUGUGCCACAGCAUGUUAACUAGAACACUGAAAAAAAUUAA-UUAAAGU ..((((((.(((((((.(((-((((.....))))))).))))))...(((((.(((....))).))))).)))))))........................-....... ( -31.80) >consensus AGGGCAUGGCGAGGCUCAGUUUGAGAACUGUUCAGCUCAGCUUCUUCGUGGCUUAGGAGCCACUGCCACAGCAUGUUAACUAGAGCACUGAGCAAAAUUUAAUUAAUGU ...(((((((((((((.(((.((((.....))))))).))))))...((((((....)))))).))).(((..((((......)))))))...............)))) (-28.12 = -28.12 + -0.00)



| Location | 15,337,128 – 15,337,237 |
|---|---|
| Length | 109 |
| Sequences | 5 |
| Columns | 109 |
| Reading direction | reverse |
| Mean pairwise identity | 92.00 |
| Mean single sequence MFE | -29.62 |
| Consensus MFE | -24.44 |
| Energy contribution | -24.16 |
| Covariance contribution | -0.28 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.49 |
| Structure conservation index | 0.83 |
| SVM decision value | 0.46 |
| SVM RNA-class probability | 0.747019 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 15337128 109 - 23771897 ACAUUAAUUAAAUUUUGCUCAGUGCUCUAGUUAACAUGCUGUGGCAGUGGCUUCUAAGCCACGAAGAAGCUGAGCUGAACAGUUCUCAAACUGAGCCUCGCCAUGCCCU ............(((.((((((((((.((((......)))).))).((((((....))))))......))))))).)))(((((....)))))................ ( -31.60) >DroSec_CAF1 69185 109 - 1 ACAUUAAUUAAAUUUUGCUCAGUGCUCUAGUUAACAUGCUGUGGCAGUGGCUCCUAAGCCACGAAGAAGCUGAGCUGAACAGUUCUCAAACUGAGCCUCGCCAUGCCCU ............(((.((((((((((.((((......)))).))).((((((....))))))......))))))).)))(((((....)))))................ ( -31.50) >DroSim_CAF1 70586 109 - 1 ACAUUAAUUAAAUUUUGCUCAGUGCUCUAGUUAACAUGCUGUGGCAGUGGCUCCUAAGCCACGAAGAAGCUGAGCUGAACAGUUCUCAAACUGAGCCUCGCCAUGCCCU ............(((.((((((((((.((((......)))).))).((((((....))))))......))))))).)))(((((....)))))................ ( -31.50) >DroEre_CAF1 69485 107 - 1 GCGUAUU-UAAAUUUCGUUCAGUGUUCUAGUUAACAUGCUGUGGCAGAGGCUCCUAAGCCACGAAGAAGCUGAGCUGAACAGUUCUCA-ACUGAGCCUCGCCACGCCCC ((((...-.....((((((((((.(((((((......)))(((((..((....))..)))))..))))))))))).)))..((((...-...))))......))))... ( -31.70) >DroYak_CAF1 71612 107 - 1 ACUUUAA-UUAAUUUUUUUCAGUGUUCUAGUUAACAUGCUGUGGCACAGACUCCUAAGCCACGAAGAAGCUGAGCUGAACAGUUCUCA-ACUGAGCCUCGCCAUGCCCC .......-......((((((.(((((......)))))...(((((..((....))..)))))))))))((((.((.((...((((...-...)))).)))))).))... ( -21.80) >consensus ACAUUAAUUAAAUUUUGCUCAGUGCUCUAGUUAACAUGCUGUGGCAGUGGCUCCUAAGCCACGAAGAAGCUGAGCUGAACAGUUCUCAAACUGAGCCUCGCCAUGCCCU ................(((((((((((((((......)))(((((...(....)...)))))..)).))).(((((....)))))....)))))))............. (-24.44 = -24.16 + -0.28)


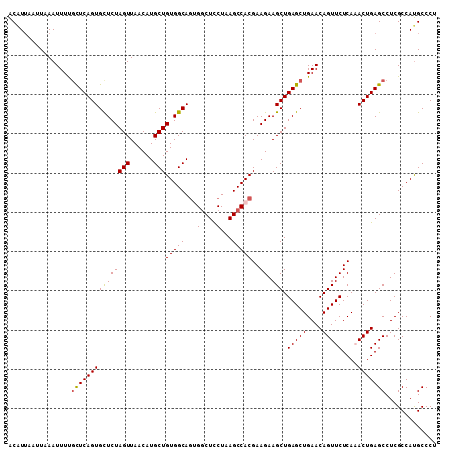
| Location | 15,337,158 – 15,337,273 |
|---|---|
| Length | 115 |
| Sequences | 5 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 83.95 |
| Mean single sequence MFE | -30.36 |
| Consensus MFE | -21.02 |
| Energy contribution | -21.54 |
| Covariance contribution | 0.52 |
| Combinations/Pair | 1.10 |
| Mean z-score | -2.11 |
| Structure conservation index | 0.69 |
| SVM decision value | 0.71 |
| SVM RNA-class probability | 0.829156 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 15337158 115 - 23771897 GGAUCUAGCAACCUUAGUGCAUCCUAUUAAAUA--CUAACAUUAAUUAAAUUUUGCUCAGUGCUCUAGUUAACAUGCUGUGGCAGUGGCUUCUAAGCCACGAAGAAGCUGAGCUGAA ((((...(((.......))))))).........--...................((((((((((.((((......)))).))).((((((....))))))......))))))).... ( -32.10) >DroSec_CAF1 69215 115 - 1 GGCUCUAGCAACCUUAGUGCAUCCUAUUAAAUG--CUAACAUUAAUUAAAUUUUGCUCAGUGCUCUAGUUAACAUGCUGUGGCAGUGGCUCCUAAGCCACGAAGAAGCUGAGCUGAA .....(((((...((((((.....)))))).))--)))................((((((((((.((((......)))).))).((((((....))))))......))))))).... ( -32.80) >DroSim_CAF1 70616 115 - 1 GGCUCUAGCAACCUUAGUGCAUCCUAUUAAAUG--CUAACAUUAAUUAAAUUUUGCUCAGUGCUCUAGUUAACAUGCUGUGGCAGUGGCUCCUAAGCCACGAAGAAGCUGAGCUGAA .....(((((...((((((.....)))))).))--)))................((((((((((.((((......)))).))).((((((....))))))......))))))).... ( -32.80) >DroEre_CAF1 69514 111 - 1 GGCUGU----UCCUUAGCGC-UCCUAUAAAAUAUAUUUGCGUAUU-UAAAUUUCGUUCAGUGUUCUAGUUAACAUGCUGUGGCAGAGGCUCCUAAGCCACGAAGAAGCUGAGCUGAA .((((.----....))))..-.......((((((......)))))-)....((((((((((.(((((((......)))(((((..((....))..)))))..))))))))))).))) ( -28.10) >DroYak_CAF1 71641 109 - 1 AGC-------ACAUCAGCACAUCUUAUAAAAUUGAAUAACUUUAA-UUAAUUUUUUUCAGUGUUCUAGUUAACAUGCUGUGGCACAGACUCCUAAGCCACGAAGAAGCUGAGCUGAA ...-------...(((((...........(((((((....)))))-))........(((((.(((((((......)))(((((..((....))..)))))..)))))))))))))). ( -26.00) >consensus GGCUCUAGCAACCUUAGUGCAUCCUAUUAAAUA__CUAACAUUAAUUAAAUUUUGCUCAGUGCUCUAGUUAACAUGCUGUGGCAGUGGCUCCUAAGCCACGAAGAAGCUGAGCUGAA ......................................................((((((((((.((((......)))).))).((((((....))))))......))))))).... (-21.02 = -21.54 + 0.52)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:54:10 2006