
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 15,289,333 – 15,289,431 |
| Length | 98 |
| Max. P | 0.995575 |

| Location | 15,289,333 – 15,289,431 |
|---|---|
| Length | 98 |
| Sequences | 6 |
| Columns | 98 |
| Reading direction | forward |
| Mean pairwise identity | 70.41 |
| Mean single sequence MFE | -23.24 |
| Consensus MFE | -16.08 |
| Energy contribution | -16.89 |
| Covariance contribution | 0.81 |
| Combinations/Pair | 1.40 |
| Mean z-score | -1.70 |
| Structure conservation index | 0.69 |
| SVM decision value | 1.79 |
| SVM RNA-class probability | 0.977223 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
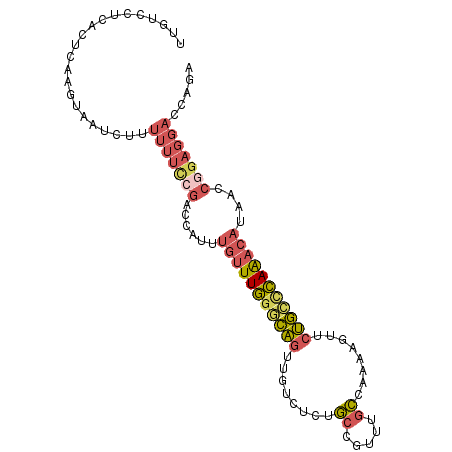
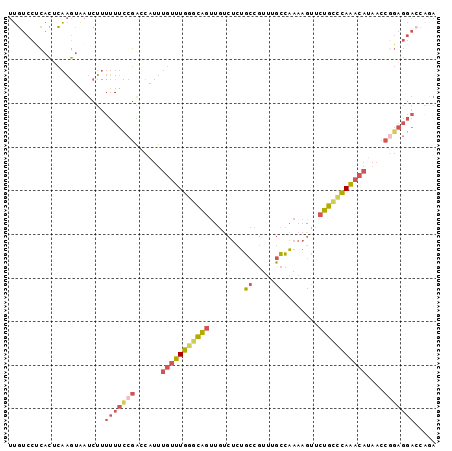
>3L_DroMel_CAF1 15289333 98 + 23771897 UUGUCCUCACUCAAGUAAUCUUUUUUCCGACCAUUUGUUUGGGCAGUUGUCUCUGCCGUUUGCCAAAAGUUCUGCCCAAACAUAACCGGAGGACCAGA ..((((((((....))...........((......(((((((((((........((.....))........)))))))))))....)))))))).... ( -28.29) >DroSim_CAF1 21157 98 + 1 UUGUCCUCACUCAAGUAAUCUUUUUUCCGACCAUUUGUUUGGGCAGUUGUCUCUGCCGUUUGCCAAAAGUUCUGCCCAAACAUAACCGGAGGACCAGA ..((((((((....))...........((......(((((((((((........((.....))........)))))))))))....)))))))).... ( -28.29) >DroEre_CAF1 21103 98 + 1 UUGUCCUCACACAAGUAAUCUUUUUUCCGACCAUUUGUUUGGGCAGUUGUCUCUGCCGUUUGCCAAAAGUUCUGCCCAAACACAACCGGAGGACCAGA ..((((((....(((....))).............(((((((((((........((.....))........)))))))))))......)))))).... ( -27.99) >DroWil_CAF1 18321 79 + 1 UGGCUUUUCUCUCUACUAUCGUUAUAUGGUUUUCUUAUUUCUUUUAUUUUCCUCUUGGCUCUAGGCCCAGAAAAACGAG------------------- ..................(((((....((.....................))(((.(((.....))).)))..))))).------------------- ( -9.80) >DroYak_CAF1 18337 98 + 1 UUGUCCCCACUCAAGUAAUCUUUUUUCCGACCAUUUGUUUGGACAGUUGUCUCUGCCGUUUGCCAAAAGUUCUGCCCAAACAUAACCGGAGGACCAGA ..((((..((....)).........((((......(((((((.(((........((.....))........))).)))))))....)))))))).... ( -20.59) >DroAna_CAF1 19835 98 + 1 UUGUCCCCACUCAAGUAAUCUUUUUUUCGGGCAUUUGUUUGGGCAGUUGUCUCUGCCGUUUGCCAAAAGUUCUGCUCAAACAUCACCAGAGGAGGCAA .((((.((((....))............((.....(((((((((((........((.....))........)))))))))))...))...)).)))). ( -24.49) >consensus UUGUCCUCACUCAAGUAAUCUUUUUUCCGACCAUUUGUUUGGGCAGUUGUCUCUGCCGUUUGCCAAAAGUUCUGCCCAAACAUAACCGGAGGACCAGA ......................(((((((......(((((((((((........((.....))........)))))))))))....)))))))..... (-16.08 = -16.89 + 0.81)

| Location | 15,289,333 – 15,289,431 |
|---|---|
| Length | 98 |
| Sequences | 6 |
| Columns | 98 |
| Reading direction | reverse |
| Mean pairwise identity | 70.41 |
| Mean single sequence MFE | -30.78 |
| Consensus MFE | -20.74 |
| Energy contribution | -22.75 |
| Covariance contribution | 2.01 |
| Combinations/Pair | 1.35 |
| Mean z-score | -3.23 |
| Structure conservation index | 0.67 |
| SVM decision value | 2.59 |
| SVM RNA-class probability | 0.995575 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
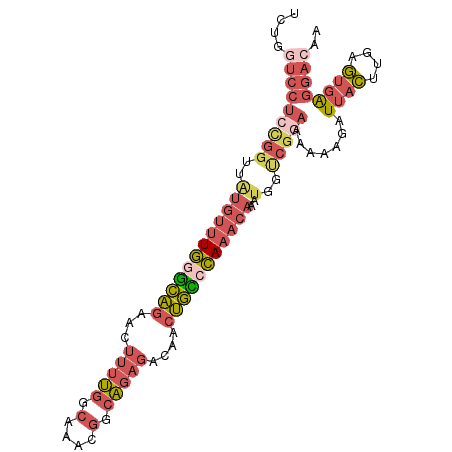
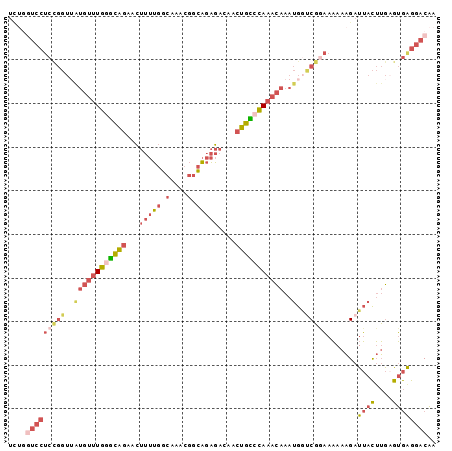
>3L_DroMel_CAF1 15289333 98 - 23771897 UCUGGUCCUCCGGUUAUGUUUGGGCAGAACUUUUGGCAAACGGCAGAGACAACUGCCCAAACAAAUGGUCGGAAAAAAGAUUACUUGAGUGAGGACAA ....((((((((..(((((((((((((...(((((.(....).)))))....)))))))))))..))..)))).......((((....)))))))).. ( -37.00) >DroSim_CAF1 21157 98 - 1 UCUGGUCCUCCGGUUAUGUUUGGGCAGAACUUUUGGCAAACGGCAGAGACAACUGCCCAAACAAAUGGUCGGAAAAAAGAUUACUUGAGUGAGGACAA ....((((((((..(((((((((((((...(((((.(....).)))))....)))))))))))..))..)))).......((((....)))))))).. ( -37.00) >DroEre_CAF1 21103 98 - 1 UCUGGUCCUCCGGUUGUGUUUGGGCAGAACUUUUGGCAAACGGCAGAGACAACUGCCCAAACAAAUGGUCGGAAAAAAGAUUACUUGUGUGAGGACAA ....((((((((..(.(((((((((((...(((((.(....).)))))....)))))))))))...)..)))).......((((....)))))))).. ( -37.10) >DroWil_CAF1 18321 79 - 1 -------------------CUCGUUUUUCUGGGCCUAGAGCCAAGAGGAAAAUAAAAGAAAUAAGAAAACCAUAUAACGAUAGUAGAGAGAAAAGCCA -------------------(((.(((((((.(((.....))).)))))))..........................((....)).))).......... ( -11.50) >DroYak_CAF1 18337 98 - 1 UCUGGUCCUCCGGUUAUGUUUGGGCAGAACUUUUGGCAAACGGCAGAGACAACUGUCCAAACAAAUGGUCGGAAAAAAGAUUACUUGAGUGGGGACAA ....((((.(((.((((((((((((((...(((((.(....).)))))....)))))))))))..(((((........)))))..))).))))))).. ( -33.90) >DroAna_CAF1 19835 98 - 1 UUGCCUCCUCUGGUGAUGUUUGAGCAGAACUUUUGGCAAACGGCAGAGACAACUGCCCAAACAAAUGCCCGAAAAAAAGAUUACUUGAGUGGGGACAA .(((((((((.(((((((((((..((((...)))).)))))(((((......)))))......................)))))).))).)))).)). ( -28.20) >consensus UCUGGUCCUCCGGUUAUGUUUGGGCAGAACUUUUGGCAAACGGCAGAGACAACUGCCCAAACAAAUGGUCGGAAAAAAGAUUACUUGAGUGAGGACAA ....(((((((((..((((((((((((...(((((.(....).)))))....)))))))))))..)..))))).......((((....)))))))).. (-20.74 = -22.75 + 2.01)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:53:47 2006