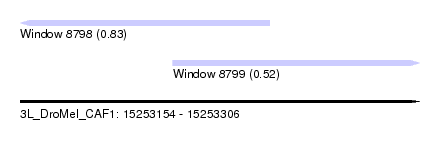
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 15,253,154 – 15,253,306 |
| Length | 152 |
| Max. P | 0.831021 |

| Location | 15,253,154 – 15,253,249 |
|---|---|
| Length | 95 |
| Sequences | 5 |
| Columns | 112 |
| Reading direction | reverse |
| Mean pairwise identity | 86.35 |
| Mean single sequence MFE | -25.68 |
| Consensus MFE | -18.92 |
| Energy contribution | -18.76 |
| Covariance contribution | -0.16 |
| Combinations/Pair | 1.23 |
| Mean z-score | -2.05 |
| Structure conservation index | 0.74 |
| SVM decision value | 0.71 |
| SVM RNA-class probability | 0.831021 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
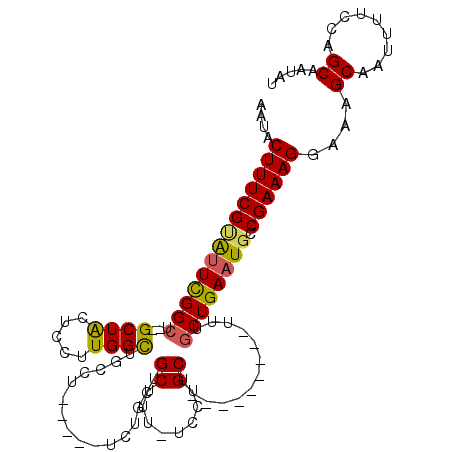
>3L_DroMel_CAF1 15253154 95 - 23771897 AAUACUUUCGUAUUCGGCU--GCUACUCCUUGGCUGCCU-----UCUGCUGCUUU-UCCUGCU---------UUUGCUGAAUGCCGAAAGGAAAGCAAUUUUCCAGCAAUAU (((((....))))).(((.--((((.....)))).))).-----..(((((....-...((((---------(((((.....))(....))))))))......))))).... ( -24.42) >DroSec_CAF1 142345 95 - 1 AAUACUUUCGUAUUCGGCU--GCUACUCCUUGGCUGCCU-----UCUGCUGCUUU-UCCUGCU---------CUUGCUGAAUGCCGAAAGGAAAGCAAUUUUCCAGCAAUAU ....((((((((((((((.--((........(((.((..-----...)).)))..-....)).---------...)))))))).))))))....((.........))..... ( -23.94) >DroSim_CAF1 150556 95 - 1 AAUACUUUCGUAUUCGGCU--GCUACUCCUUGGCUGCCU-----UCUGCUGCUUU-UCCUGCU---------UUUGCUGAAUGCCGAAAGGAAAGCAAUUUUCCAGCAAUAU (((((....))))).(((.--((((.....)))).))).-----..(((((....-...((((---------(((((.....))(....))))))))......))))).... ( -24.42) >DroEre_CAF1 142670 104 - 1 AAUACUUUCGCAUUCGGCU--GCUACUCCUUGGCUGCCU-----UCUGCUGCUCU-UCCUGCUCUUCCUGCUUUUGCUGAAUGCCGAAAGGAAAGCAAUUUUCCAGCAAUAU ...............(((.--((((.....)))).))).-----..(((((....-...((((.(((((..(((.((.....)).))))))))))))......))))).... ( -25.52) >DroAna_CAF1 134528 103 - 1 AAUACUUUCGGGCUUGGCUGUGCUGCUCCUUGGUUUUCUCAGCUUUGGCAGCUUUCUGGUGCC---------UUUCCUGAAUGCCGAAAGGAAAGCAAUUUUCCAGCAAUAU ...........(((.((...((((..((((((((....((((....((((.(.....).))))---------....))))..)))..))))).)))).....)))))..... ( -30.10) >consensus AAUACUUUCGUAUUCGGCU__GCUACUCCUUGGCUGCCU_____UCUGCUGCUUU_UCCUGCU_________UUUGCUGAAUGCCGAAAGGAAAGCAAUUUUCCAGCAAUAU ....((((((((((((((...((((.....))))................((........)).............)))))))).))))))....((.........))..... (-18.92 = -18.76 + -0.16)


| Location | 15,253,212 – 15,253,306 |
|---|---|
| Length | 94 |
| Sequences | 4 |
| Columns | 109 |
| Reading direction | forward |
| Mean pairwise identity | 85.85 |
| Mean single sequence MFE | -25.16 |
| Consensus MFE | -17.39 |
| Energy contribution | -17.20 |
| Covariance contribution | -0.19 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.46 |
| Structure conservation index | 0.69 |
| SVM decision value | -0.02 |
| SVM RNA-class probability | 0.523642 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 15253212 94 + 23771897 AGGCAGCCAAGGAGUAGCAGCCGAAUACGAAAGUAUUGUUAACCAACAAUAACAACAACACAAAGGGGGAUGGG------------UGCGGCAGGGGGUG---GCUAUA .(((.(((.....((.(((.((.(((((....)))))(((..((....................))..))))).------------))).))....))).---)))... ( -24.45) >DroSec_CAF1 142403 97 + 1 AGGCAGCCAAGGAGUAGCAGCCGAAUACGAAAGUAUUGUUAACCAACAAUAACAACAACACAAAGGGGGAUGUC------------UGCGGCAGGGGGUGGCGGCUAUA .(((.((((..(.((((((.((.(((((....)))))((((........))))..............)).)).)------------)))..)......)))).)))... ( -24.30) >DroSim_CAF1 150614 97 + 1 AGGCAGCCAAGGAGUAGCAGCCGAAUACGAAAGUAUUGUUAACCAACAAUAACAACAACACAAAGGGGGAUGGC------------UGCGGCAGGGGGUGGCGGCUAUA .(((.((((....((.((((((.(((((....)))))(((..((....................))..))))))------------))).))......)))).)))... ( -31.35) >DroEre_CAF1 142737 100 + 1 AGGCAGCCAAGGAGUAGCAGCCGAAUGCGAAAGUAUUGUUAACCAACAAUAACAACGACACAAAGGGGGAUGAGGUGGGAAAAUCGGGCGGCGGGGGGUG--------- ..((.(((...((....(.(((.(((((....)))))(((..((....................))..)))..)))..)....)).))).))........--------- ( -20.55) >consensus AGGCAGCCAAGGAGUAGCAGCCGAAUACGAAAGUAUUGUUAACCAACAAUAACAACAACACAAAGGGGGAUGGC____________UGCGGCAGGGGGUG___GCUAUA ...((.((..(......).(((((((((....)))))(((..((....................))..))).................))))...)).))......... (-17.39 = -17.20 + -0.19)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:53:28 2006