
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 15,072,784 – 15,072,874 |
| Length | 90 |
| Max. P | 0.993778 |

| Location | 15,072,784 – 15,072,874 |
|---|---|
| Length | 90 |
| Sequences | 5 |
| Columns | 97 |
| Reading direction | forward |
| Mean pairwise identity | 82.72 |
| Mean single sequence MFE | -27.32 |
| Consensus MFE | -21.34 |
| Energy contribution | -22.04 |
| Covariance contribution | 0.70 |
| Combinations/Pair | 1.22 |
| Mean z-score | -2.59 |
| Structure conservation index | 0.78 |
| SVM decision value | 2.03 |
| SVM RNA-class probability | 0.986244 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 15072784 90 + 23771897 ------UAGCUGCCGCCAGAAAUUGCUUUUGCUCGCUGGAAUAUCCAUACAUACAUAUGUAUAUGUAUGUCUGUGAAUGUGUGUGCU-UUGCGGCCU ------..(((((..((((.....((....))...))))....((((.(((((((((....))))))))).)).))...........-..))))).. ( -25.60) >DroSec_CAF1 37386 90 + 1 ------UAGCUGCCGCCAGAAAUUGCUUUUGCUCGCUGGAAUAUACAUGCAUACAUAUGUAUGUAUAUGCCUGUGAAUGUGUGUGCU-UUGCGGCCU ------..(((((.((((.(....((....))((((.((.((((((((((((....)))))))))))).)).))))...).)).)).-..))))).. ( -31.20) >DroSim_CAF1 38679 90 + 1 ------UAGCUGCCGCCAGAAAUUGCUUUUGCUCGCUGGAAUAUACAUACAUACAUAUGUAUGUAUAUGUCUGUGAAUGUGUGUGCU-UUGCGGCCU ------..(((((.((((.(....((....))((((.(..((((((((((((....))))))))))))..).))))...).)).)).-..))))).. ( -28.90) >DroEre_CAF1 38531 89 + 1 ------UAGCUGCCGCCAGAAAUUGCUUUUGCUCGCUGGAAUAUACAUACAUACAUAAGUAUGUAUGUUU--CUGUAUGUGUAUGCUUUUGCGGCCU ------.....(((((.((((...((...(((..((.(((((((((((((........))))))))))))--).))..)))...))))))))))).. ( -29.20) >DroAna_CAF1 36415 79 + 1 UUGCCGUAGCUGCCGCCAGAAAUUGCUUUUGCUCGCUGGAAUAUAUAUA---------GUAUGUGU------GUGUGUGUGUGUGCU---ACAGCCU ..((.(((((((((.((((.....((....))...)))).(((((((((---------.....)))------))))))).))).)))---)).)).. ( -21.70) >consensus ______UAGCUGCCGCCAGAAAUUGCUUUUGCUCGCUGGAAUAUACAUACAUACAUAUGUAUGUAUAUGUCUGUGAAUGUGUGUGCU_UUGCGGCCU ........(((((..((((.....((....))...)))).((((((((((((....))))))))))))......................))))).. (-21.34 = -22.04 + 0.70)

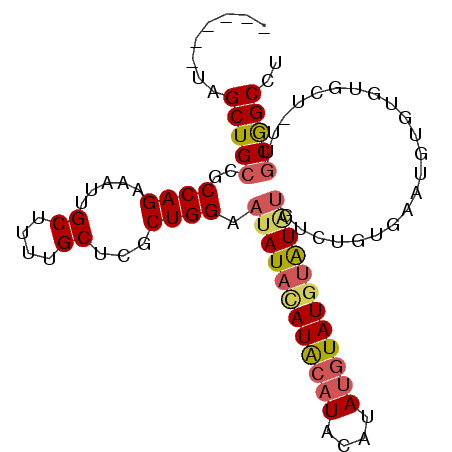

| Location | 15,072,784 – 15,072,874 |
|---|---|
| Length | 90 |
| Sequences | 5 |
| Columns | 97 |
| Reading direction | reverse |
| Mean pairwise identity | 82.72 |
| Mean single sequence MFE | -24.50 |
| Consensus MFE | -16.32 |
| Energy contribution | -17.98 |
| Covariance contribution | 1.66 |
| Combinations/Pair | 1.12 |
| Mean z-score | -3.28 |
| Structure conservation index | 0.67 |
| SVM decision value | 2.42 |
| SVM RNA-class probability | 0.993778 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 15072784 90 - 23771897 AGGCCGCAA-AGCACACACAUUCACAGACAUACAUAUACAUAUGUAUGUAUGGAUAUUCCAGCGAGCAAAAGCAAUUUCUGGCGGCAGCUA------ ..(((((..-.((..............(((((((((....))))))))).(((.....)))))..((....))........))))).....------ ( -26.30) >DroSec_CAF1 37386 90 - 1 AGGCCGCAA-AGCACACACAUUCACAGGCAUAUACAUACAUAUGUAUGCAUGUAUAUUCCAGCGAGCAAAAGCAAUUUCUGGCGGCAGCUA------ ..(((((..-................((.((((((((.(((....))).)))))))).)).((........))........))))).....------ ( -24.50) >DroSim_CAF1 38679 90 - 1 AGGCCGCAA-AGCACACACAUUCACAGACAUAUACAUACAUAUGUAUGUAUGUAUAUUCCAGCGAGCAAAAGCAAUUUCUGGCGGCAGCUA------ ..(((((..-.(......).....((((.((((((((((((....))))))))))))....((........))....))))))))).....------ ( -26.50) >DroEre_CAF1 38531 89 - 1 AGGCCGCAAAAGCAUACACAUACAG--AAACAUACAUACUUAUGUAUGUAUGUAUAUUCCAGCGAGCAAAAGCAAUUUCUGGCGGCAGCUA------ ..(((((....(......)...(((--((((((((((((....))))))))))........((........))...)))))))))).....------ ( -28.50) >DroAna_CAF1 36415 79 - 1 AGGCUGU---AGCACACACACACAC------ACACAUAC---------UAUAUAUAUUCCAGCGAGCAAAAGCAAUUUCUGGCGGCAGCUACGGCAA ..(((((---(((............------....(((.---------....)))...((.((.((..(((....))))).))))..)))))))).. ( -16.70) >consensus AGGCCGCAA_AGCACACACAUUCACAGACAUAUACAUACAUAUGUAUGUAUGUAUAUUCCAGCGAGCAAAAGCAAUUUCUGGCGGCAGCUA______ ..(((((......................((((((((((((....))))))))))))..(((.((((....))..)).))))))))........... (-16.32 = -17.98 + 1.66)


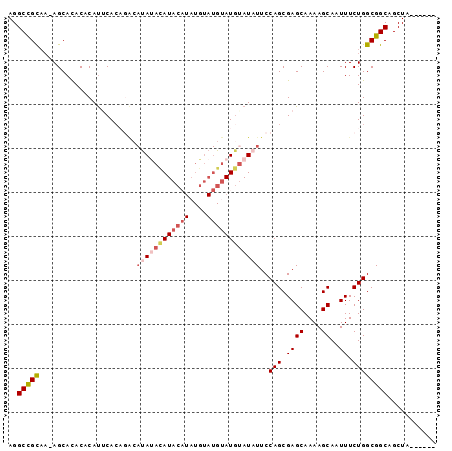
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:51:43 2006