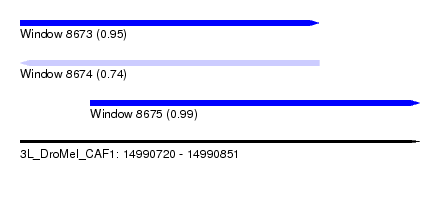

| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 14,990,720 – 14,990,851 |
| Length | 131 |
| Max. P | 0.994889 |

| Location | 14,990,720 – 14,990,818 |
|---|---|
| Length | 98 |
| Sequences | 5 |
| Columns | 99 |
| Reading direction | forward |
| Mean pairwise identity | 95.33 |
| Mean single sequence MFE | -24.12 |
| Consensus MFE | -21.42 |
| Energy contribution | -22.10 |
| Covariance contribution | 0.68 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.62 |
| Structure conservation index | 0.89 |
| SVM decision value | 1.43 |
| SVM RNA-class probability | 0.952904 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
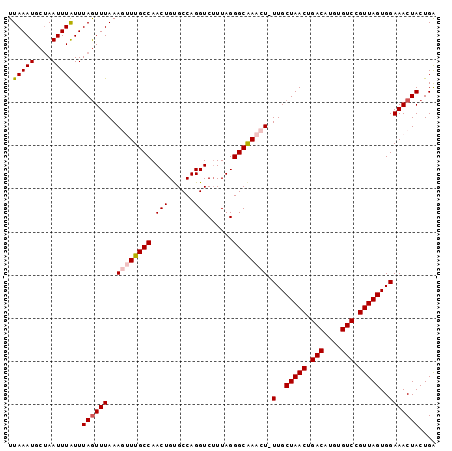
>3L_DroMel_CAF1 14990720 98 + 23771897 UUAAAUGCUAAUUUGUUUAGUUUAAAGUUUGCCAACUGUGCCAGGUCUUUAGGGCAAACU-UUGCUAACUGACAUGUGUCCGUUAGUGGAAACUACUGA .................((((((((((((((((.(((......)))......))))))))-))((((((.(((....))).))))))..)))))).... ( -25.90) >DroSec_CAF1 9103 98 + 1 UUAAAUGCUAAUUUAUUUAAUUUAAAGUUUGCCAACUGUGCCAGGUCAUUAGGGCAAACU-UUGCUAACUGACAUGUGUCCGUUAGUGGAAACUACUGA (((((((......)))))))...((((((((((....(((......)))...))))))))-))((((((.(((....))).))))))(....)...... ( -25.90) >DroSim_CAF1 8015 98 + 1 UUAAAUGCUAAUUUAUUUAGUUUAAAGUUUGCCAACUGUGCCAGGUCUUUAGGGCAAACU-UUGCUAACUGACAUGUGUCCGUUAGUGGAAACUACUGA .(((((....)))))..((((((((((((((((.(((......)))......))))))))-))((((((.(((....))).))))))..)))))).... ( -26.20) >DroEre_CAF1 8071 98 + 1 UUAAAUGCUAAUUUAUUUAGUUUAAAACUUGCCAACUGCGCCAGGUCUUUAGGGCGAACU-UUGCUAACUGACAUGUGUCCGUUAGUGGAAACUACUGG ......(((((.....)))))..........(((....((((..(....)..))))...(-(..(((((.(((....))).)))))..))......))) ( -21.90) >DroYak_CAF1 9629 99 + 1 UUAAAUGCAAAUUUAUUUAGUUUAAAACUUGCCAACUGCGCCAGGUCUUUAGGGCAAACUUUUGCUAACUGACAUGUGUCCGUUAGUGGAAACUACUGA .(((((....)))))..((((((.....(((((.(((......)))......)))))....(..(((((.(((....))).)))))..))))))).... ( -20.70) >consensus UUAAAUGCUAAUUUAUUUAGUUUAAAGUUUGCCAACUGUGCCAGGUCUUUAGGGCAAACU_UUGCUAACUGACAUGUGUCCGUUAGUGGAAACUACUGA .(((((....)))))..((((((..((((((((.(((......)))......)))))))).(..(((((.(((....))).)))))..))))))).... (-21.42 = -22.10 + 0.68)


| Location | 14,990,720 – 14,990,818 |
|---|---|
| Length | 98 |
| Sequences | 5 |
| Columns | 99 |
| Reading direction | reverse |
| Mean pairwise identity | 95.33 |
| Mean single sequence MFE | -20.28 |
| Consensus MFE | -17.12 |
| Energy contribution | -16.68 |
| Covariance contribution | -0.44 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.48 |
| Structure conservation index | 0.84 |
| SVM decision value | 0.45 |
| SVM RNA-class probability | 0.739941 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
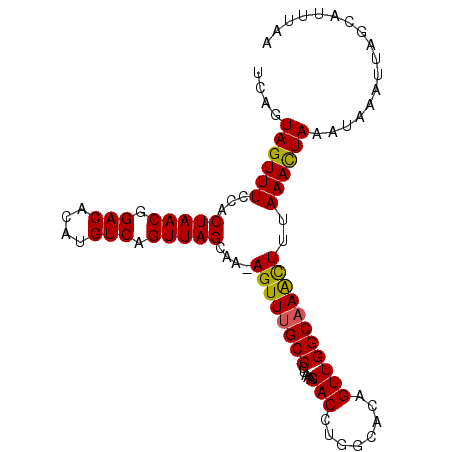
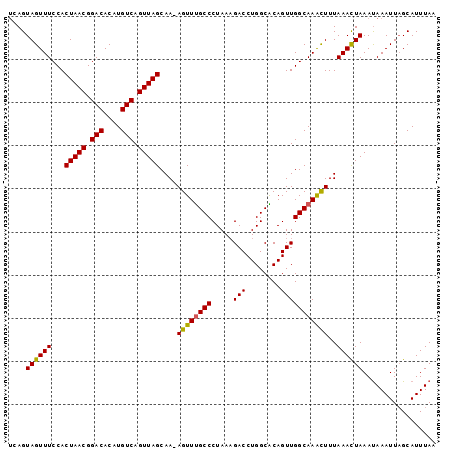
>3L_DroMel_CAF1 14990720 98 - 23771897 UCAGUAGUUUCCACUAACGGACACAUGUCAGUUAGCAA-AGUUUGCCCUAAAGACCUGGCACAGUUGGCAAACUUUAAACUAAACAAAUUAGCAUUUAA ....((((((...(((((.(((....))).))))).((-((((((((.....(((........)))))))))))))))))))................. ( -22.50) >DroSec_CAF1 9103 98 - 1 UCAGUAGUUUCCACUAACGGACACAUGUCAGUUAGCAA-AGUUUGCCCUAAUGACCUGGCACAGUUGGCAAACUUUAAAUUAAAUAAAUUAGCAUUUAA ....((((((...(((((.(((....))).))))).((-((((((((....((........))...))))))))))))))))................. ( -21.10) >DroSim_CAF1 8015 98 - 1 UCAGUAGUUUCCACUAACGGACACAUGUCAGUUAGCAA-AGUUUGCCCUAAAGACCUGGCACAGUUGGCAAACUUUAAACUAAAUAAAUUAGCAUUUAA ....((((((...(((((.(((....))).))))).((-((((((((.....(((........)))))))))))))))))))................. ( -22.50) >DroEre_CAF1 8071 98 - 1 CCAGUAGUUUCCACUAACGGACACAUGUCAGUUAGCAA-AGUUCGCCCUAAAGACCUGGCGCAGUUGGCAAGUUUUAAACUAAAUAAAUUAGCAUUUAA ....((((((...(((((.(((....))).)))))(((-....((((..........))))...))).........))))))................. ( -16.50) >DroYak_CAF1 9629 99 - 1 UCAGUAGUUUCCACUAACGGACACAUGUCAGUUAGCAAAAGUUUGCCCUAAAGACCUGGCGCAGUUGGCAAGUUUUAAACUAAAUAAAUUUGCAUUUAA .............(((((.(((....))).))))).......((((((.........)).))))...((((((((..........))))))))...... ( -18.80) >consensus UCAGUAGUUUCCACUAACGGACACAUGUCAGUUAGCAA_AGUUUGCCCUAAAGACCUGGCACAGUUGGCAAACUUUAAACUAAAUAAAUUAGCAUUUAA ....((((((...(((((.(((....))).)))))....((((((((.....(((........)))))))))))..))))))................. (-17.12 = -16.68 + -0.44)

| Location | 14,990,743 – 14,990,851 |
|---|---|
| Length | 108 |
| Sequences | 5 |
| Columns | 109 |
| Reading direction | forward |
| Mean pairwise identity | 94.19 |
| Mean single sequence MFE | -35.56 |
| Consensus MFE | -29.64 |
| Energy contribution | -29.44 |
| Covariance contribution | -0.20 |
| Combinations/Pair | 1.07 |
| Mean z-score | -2.49 |
| Structure conservation index | 0.83 |
| SVM decision value | 2.52 |
| SVM RNA-class probability | 0.994889 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
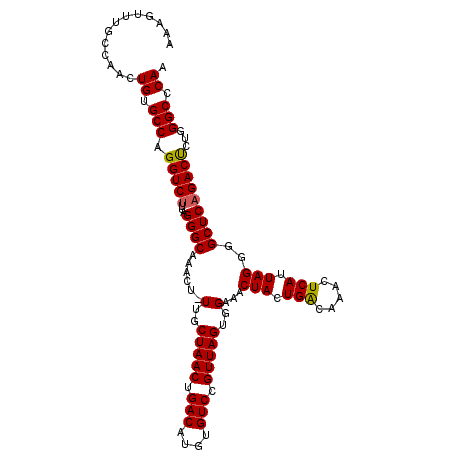
>3L_DroMel_CAF1 14990743 108 + 23771897 AAAGUUUGCCAACUGUGCCAGGUCUUUAGGGCAAACU-UUGCUAACUGACAUGUGUCCGUUAGUGGAAACUACUGACAAACUCAUUAGGGGCUCAGACUCUGGGCCCAA ((((((((((.(((......)))......))))))))-))((((((.(((....))).)))))).....(((.(((.....))).)))((((((((...)))))))).. ( -38.50) >DroSec_CAF1 9126 108 + 1 AAAGUUUGCCAACUGUGCCAGGUCAUUAGGGCAAACU-UUGCUAACUGACAUGUGUCCGUUAGUGGAAACUACUGACAAACUCAUUAGGGGCUCAGACUCCGGGCCCAA ((((((((((....(((......)))...))))))))-))((((((.(((....))).)))))).....(((.(((.....))).)))((((((.......)))))).. ( -37.20) >DroSim_CAF1 8038 108 + 1 AAAGUUUGCCAACUGUGCCAGGUCUUUAGGGCAAACU-UUGCUAACUGACAUGUGUCCGUUAGUGGAAACUACUGACAAACUCAUUAGGGGCUCAGACUCCGGGCCCAA ((((((((((.(((......)))......))))))))-))((((((.(((....))).)))))).....(((.(((.....))).)))((((((.......)))))).. ( -35.60) >DroEre_CAF1 8094 107 + 1 AAAACUUGCCAACUGCGCCAGGUCUUUAGGGCGAACU-UUGCUAACUGACAUGUGUCCGUUAGUGGAAACUACUGGCAAACUCAUUAGGUGCUCAGACCUUGGGC-CAA .............((.(((((((((...(((((...(-(..(((((.(((....))).)))))..))..(((.(((.....))).))).)))))))))))..)))-)). ( -32.30) >DroYak_CAF1 9652 109 + 1 AAAACUUGCCAACUGCGCCAGGUCUUUAGGGCAAACUUUUGCUAACUGACAUGUGUCCGUUAGUGGAAACUACUGACAAACUCAUUAGGCGCUCAGACCUUGGGCCCAA .............((.(((((((((...((((....(((..(((((.(((....))).)))))..))).(((.(((.....))).)))..))))))))))..))).)). ( -34.20) >consensus AAAGUUUGCCAACUGUGCCAGGUCUUUAGGGCAAACU_UUGCUAACUGACAUGUGUCCGUUAGUGGAAACUACUGACAAACUCAUUAGGGGCUCAGACUCUGGGCCCAA .............((.(((.(((((...((((......(..(((((.(((....))).)))))..)...(((.(((.....))).)))..)))))))))...))).)). (-29.64 = -29.44 + -0.20)

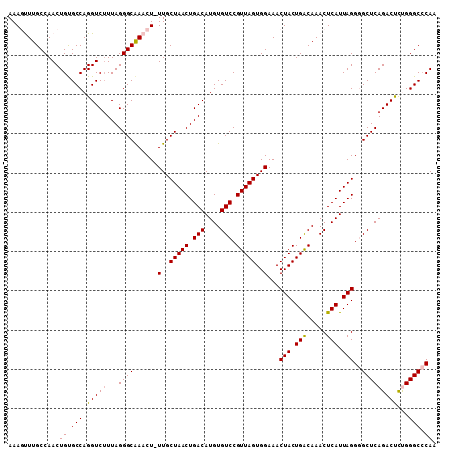
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:51:30 2006