
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 14,891,922 – 14,892,013 |
| Length | 91 |
| Max. P | 0.526526 |

| Location | 14,891,922 – 14,892,013 |
|---|---|
| Length | 91 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 77.22 |
| Mean single sequence MFE | -33.35 |
| Consensus MFE | -14.28 |
| Energy contribution | -14.70 |
| Covariance contribution | 0.42 |
| Combinations/Pair | 1.22 |
| Mean z-score | -2.08 |
| Structure conservation index | 0.43 |
| SVM decision value | -0.01 |
| SVM RNA-class probability | 0.526526 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
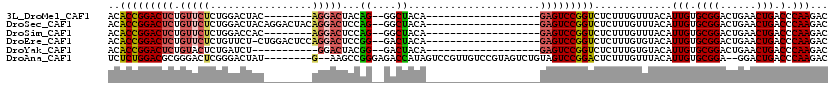
>3L_DroMel_CAF1 14891922 91 - 23771897 ACACCGGACUCUGUUCUCUGGACUAC--------AGGACUACAG--GGCUACA-------------------GAGUCCGGUCUCUUUGUUUACAUUGUGCGGACUGAACUGACCCAAGAC .....(((((((((.(((((......--------.......)))--))..)))-------------------))))))((((..((((((..(.....)..))).)))..))))...... ( -30.62) >DroSec_CAF1 73863 99 - 1 ACACCGGACUCUGUUCUCUGGACUACAGGACUACAGGACUCCAG--GGCUACA-------------------GAGUCCGGUCUCUUUGUUUACAUUGUGCGGACUGAACUGACCCAAGAC .....(((((((((.(((((((.....(..(....)..))))))--))..)))-------------------))))))((((..((((((..(.....)..))).)))..))))...... ( -34.90) >DroSim_CAF1 69860 91 - 1 ACACCGGACUCUGUUCUCUGGACCAC--------AGGACUCCAG--GGCUACA-------------------GAGUCCGGUCUCUUUGUUUACAUUGUGCGGACUGAACUGACCCAAGAC .....(((((((((.(((((((((..--------.))..)))))--))..)))-------------------))))))((((..((((((..(.....)..))).)))..))))...... ( -35.00) >DroEre_CAF1 70487 98 - 1 ACACCGGACUCUGUUCUCUGUUCU-CUGGACUCCAGGACUCCGG--GACUACA-------------------GAGUCCGGUCUCUUUGUGUACAUUGUGCGGACUGAACUGACCCAAGAC ..((((((((((((((((.(.((.-((((...))))))..).))--))..)))-------------------))))))))).((..(.(((((...))))).)..))............. ( -33.70) >DroYak_CAF1 66455 88 - 1 ACACCGGACUCUGUACUCUGAUCU-----------GGACUACGG--GACUACA-------------------GAGUCCGGUCUCUUUGUGUACAUUGUGCGGACUGAACUGACCCAAGAC ..(((((((((((((.((....((-----------(.....)))--)).))))-------------------))))))))).((..(.(((((...))))).)..))............. ( -29.00) >DroAna_CAF1 61822 108 - 1 UCUCUGGACGCGGGACUCGGGACUAU--------G--AAGCCGGGAGACCAUAGUCCGUUGUCCGUAGUCUGUAGUCCGGACUCUUUGUUUACAUUGUGCGGA--GGACUGACCCAAGAC (((.(((.....((((...(((((((--------(--....)((....)))))))))...))))...(((.((..((((.((....((....))..)).))))--..)).))))))))). ( -36.90) >consensus ACACCGGACUCUGUUCUCUGGACUAC________AGGACUCCAG__GACUACA___________________GAGUCCGGUCUCUUUGUUUACAUUGUGCGGACUGAACUGACCCAAGAC ..(((((((((.(((((.................)))))...((....))......................))))))))).............(((.((((......))).).)))... (-14.28 = -14.70 + 0.42)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:50:18 2006