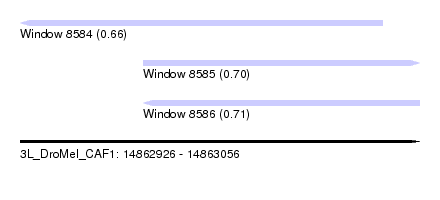
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 14,862,926 – 14,863,056 |
| Length | 130 |
| Max. P | 0.705526 |

| Location | 14,862,926 – 14,863,044 |
|---|---|
| Length | 118 |
| Sequences | 4 |
| Columns | 118 |
| Reading direction | reverse |
| Mean pairwise identity | 86.04 |
| Mean single sequence MFE | -25.30 |
| Consensus MFE | -21.08 |
| Energy contribution | -21.70 |
| Covariance contribution | 0.62 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.55 |
| Structure conservation index | 0.83 |
| SVM decision value | 0.27 |
| SVM RNA-class probability | 0.659744 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 14862926 118 - 23771897 AUGGCUGGAAGGCAAACCGACCACAAGCAGCCCAACAGCUUGGCCAACUUAUGUGAACGCACAUGCACUGCAAAACAAGAAGCGUUCGACAGAUUAAUCAAAUAAAAUAAAUAAAUAG .(((((((..((....))..)))(((((.........))))))))).....(((((((((.....................)))))).)))........................... ( -24.60) >DroSec_CAF1 44997 116 - 1 AUGGCUGGAAGGCAAACCGACCACAAGCAGCCCAACAGCUUGGCCAACUUAUGUGAGCGCACAUGCACUGCA--ACAAAAAGCGUUCGACAGAUUAAGCAAAUAAAAUAAAUAAAUGG .(((((((..((....))..)))(((((.........))))))))).(((((((((((((...(((...)))--.......)))))).)))...)))).................... ( -25.40) >DroSim_CAF1 40177 116 - 1 AUGGCUGGAAGGCAAACCGACCACAAGCAGCCCAACAGCUUGGCCAACUUAUGUGAACGCACAUGCACUGCA--GCAAAAAGCGUUCGACAGAUUAAGCAAAUAAAAUAAAUAAAUAG .(((((((..((....))..)))(((((.........))))))))).(((((((((((((...(((......--)))....)))))).)))...)))).................... ( -29.00) >DroEre_CAF1 41305 98 - 1 AUGGCUGGAAGGCAAACCGACCACAUGCAGCCCAACAGCUUGGCCAACUUAUGUGAACGCACAUGCACUGAA--ACAAAAGGCGCUAGCCAGAUUU-----------------UA-AG .(((((((...((........((((((.((.((((....))))....))))))))...((....))......--.......)).))))))).....-----------------..-.. ( -22.20) >consensus AUGGCUGGAAGGCAAACCGACCACAAGCAGCCCAACAGCUUGGCCAACUUAUGUGAACGCACAUGCACUGCA__ACAAAAAGCGUUCGACAGAUUAAGCAAAUAAAAUAAAUAAAUAG .(((((((..((....))..)))(((((.........))))))))).....(((((((((.....................))))))).))........................... (-21.08 = -21.70 + 0.62)

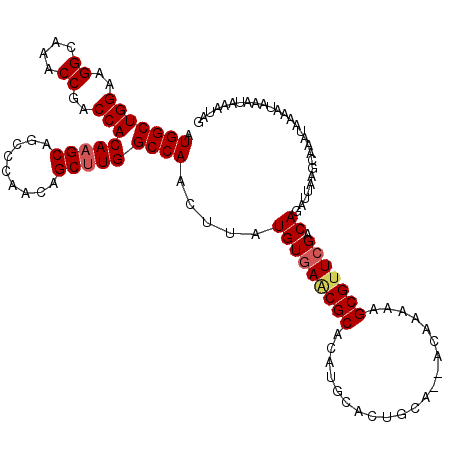
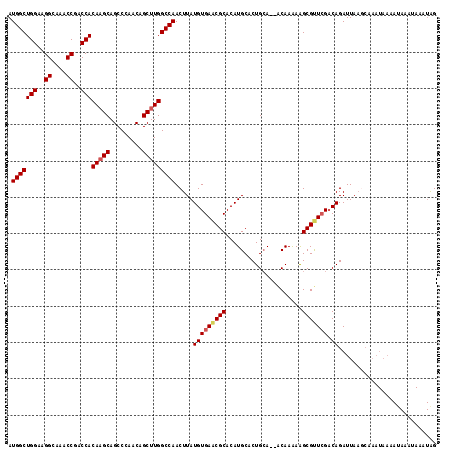
| Location | 14,862,966 – 14,863,056 |
|---|---|
| Length | 90 |
| Sequences | 6 |
| Columns | 90 |
| Reading direction | forward |
| Mean pairwise identity | 77.39 |
| Mean single sequence MFE | -23.37 |
| Consensus MFE | -16.20 |
| Energy contribution | -16.20 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.21 |
| Structure conservation index | 0.69 |
| SVM decision value | 0.34 |
| SVM RNA-class probability | 0.696700 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 14862966 90 + 23771897 UUGUUUUGCAGUGCAUGUGCGUUCACAUAAGUUGGCCAAGCUGUUGGGCUGCUUGUGGUCGGUUUGCCUUCCAGCCAUAUUCCCAUCCAU .......(((((.((...((.....((.....)).....))...)).))))).((((((.((........)).))))))........... ( -22.50) >DroPse_CAF1 93988 69 + 1 ---------------------UACAUAUAAGUUGGCCAAGCUGCUGGCCUGCUUGUGGUCGGCUUGCCUGCCAGCCAUAUACCCAUCCAU ---------------------.....((((((.(((((......))))).))))))(((.(((......))).))).............. ( -24.10) >DroSec_CAF1 45037 88 + 1 UUGU--UGCAGUGCAUGUGCGCUCACAUAAGUUGGCCAAGCUGUUGGGCUGCUUGUGGUCGGUUUGCCUUCCAGCCAUAUACCCAUCCAU ....--.(((((.((...(((((.((....)).)))...))...)).))))).((((((.((........)).))))))........... ( -23.90) >DroSim_CAF1 40217 88 + 1 UUGC--UGCAGUGCAUGUGCGUUCACAUAAGUUGGCCAAGCUGUUGGGCUGCUUGUGGUCGGUUUGCCUUCCAGCCAUAUACCCAUCCAU ..((--((.((.(((....((....(((((((.(.((((....)))).).)))))))..))...)))))..))))............... ( -23.20) >DroAna_CAF1 32964 76 + 1 --AU--UGUAGUCUAAGUGC----------UUUGGCCAAGCUGUUGGCCUGCUUGUGGUCGGGUAGCCUUCCAGCCAUAUACCCAUCCAU --..--............((----------...((((((....)))))).))..((((..(((((((......))....)))))..)))) ( -22.40) >DroPer_CAF1 91429 69 + 1 ---------------------UACAUAUAAGUUGGCCAAGCUGCUGGCCUGCUUGUGGUCGGCUUGCCUGCCAGCCAUAUACCCAUCCAU ---------------------.....((((((.(((((......))))).))))))(((.(((......))).))).............. ( -24.10) >consensus __GU__UGCAGUGCAUGUGC_UUCACAUAAGUUGGCCAAGCUGUUGGCCUGCUUGUGGUCGGUUUGCCUUCCAGCCAUAUACCCAUCCAU ................................(((((((((.........)))))(((..((....))..)))))))............. (-16.20 = -16.20 + -0.00)

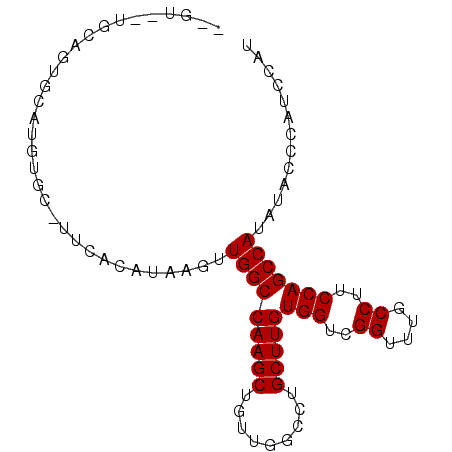
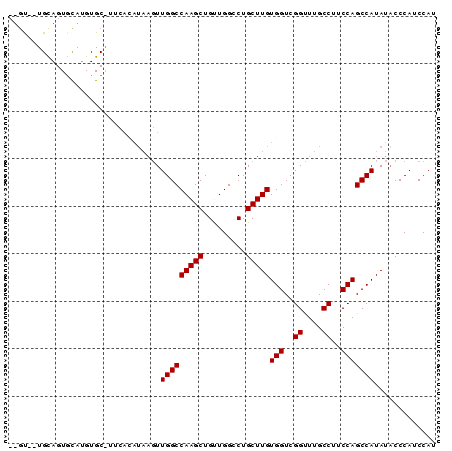
| Location | 14,862,966 – 14,863,056 |
|---|---|
| Length | 90 |
| Sequences | 6 |
| Columns | 90 |
| Reading direction | reverse |
| Mean pairwise identity | 77.39 |
| Mean single sequence MFE | -21.82 |
| Consensus MFE | -16.33 |
| Energy contribution | -16.33 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.06 |
| Structure conservation index | 0.75 |
| SVM decision value | 0.36 |
| SVM RNA-class probability | 0.705526 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
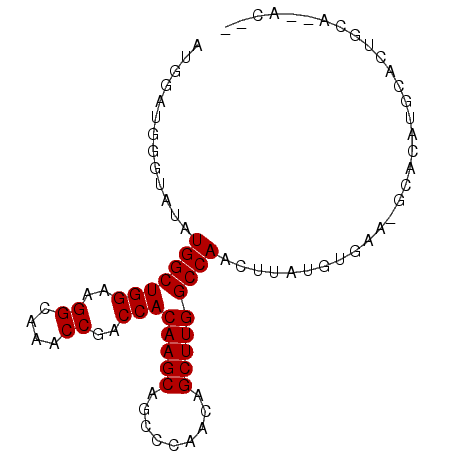
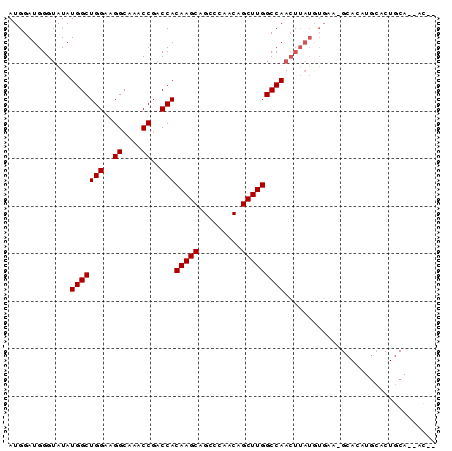
>3L_DroMel_CAF1 14862966 90 - 23771897 AUGGAUGGGAAUAUGGCUGGAAGGCAAACCGACCACAAGCAGCCCAACAGCUUGGCCAACUUAUGUGAACGCACAUGCACUGCAAAACAA .............(((((((..((....))..)))(((((.........)))))))))...((((((....))))))............. ( -20.30) >DroPse_CAF1 93988 69 - 1 AUGGAUGGGUAUAUGGCUGGCAGGCAAGCCGACCACAAGCAGGCCAGCAGCUUGGCCAACUUAUAUGUA--------------------- ....((((((...(((.((((......)))).)))......((((((....)))))).)))))).....--------------------- ( -21.40) >DroSec_CAF1 45037 88 - 1 AUGGAUGGGUAUAUGGCUGGAAGGCAAACCGACCACAAGCAGCCCAACAGCUUGGCCAACUUAUGUGAGCGCACAUGCACUGCA--ACAA ....((((((...(((((((..((....))..)))(((((.........)))))))))))))))((.((.((....)).)))).--.... ( -22.10) >DroSim_CAF1 40217 88 - 1 AUGGAUGGGUAUAUGGCUGGAAGGCAAACCGACCACAAGCAGCCCAACAGCUUGGCCAACUUAUGUGAACGCACAUGCACUGCA--GCAA ....((((((...(((((((..((....))..)))(((((.........)))))))))))))))(((....))).(((......--))). ( -22.50) >DroAna_CAF1 32964 76 - 1 AUGGAUGGGUAUAUGGCUGGAAGGCUACCCGACCACAAGCAGGCCAACAGCUUGGCCAAA----------GCACUUAGACUACA--AU-- .(((.(((((....((((....))))))))).)))...((.((((((....))))))...----------))............--..-- ( -23.20) >DroPer_CAF1 91429 69 - 1 AUGGAUGGGUAUAUGGCUGGCAGGCAAGCCGACCACAAGCAGGCCAGCAGCUUGGCCAACUUAUAUGUA--------------------- ....((((((...(((.((((......)))).)))......((((((....)))))).)))))).....--------------------- ( -21.40) >consensus AUGGAUGGGUAUAUGGCUGGAAGGCAAACCGACCACAAGCAGCCCAACAGCUUGGCCAACUUAUGUGAA_GCACAUGCACUGCA__AC__ .............(((((((..((....))..)))(((((.........)))))))))................................ (-16.33 = -16.33 + -0.00)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:50:02 2006