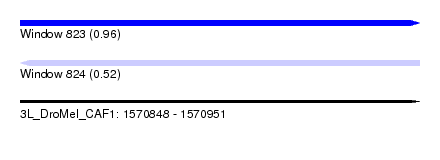

| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 1,570,848 – 1,570,951 |
| Length | 103 |
| Max. P | 0.962118 |

| Location | 1,570,848 – 1,570,951 |
|---|---|
| Length | 103 |
| Sequences | 4 |
| Columns | 114 |
| Reading direction | forward |
| Mean pairwise identity | 92.47 |
| Mean single sequence MFE | -32.38 |
| Consensus MFE | -29.65 |
| Energy contribution | -29.27 |
| Covariance contribution | -0.37 |
| Combinations/Pair | 1.07 |
| Mean z-score | -2.08 |
| Structure conservation index | 0.92 |
| SVM decision value | 1.54 |
| SVM RNA-class probability | 0.962118 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
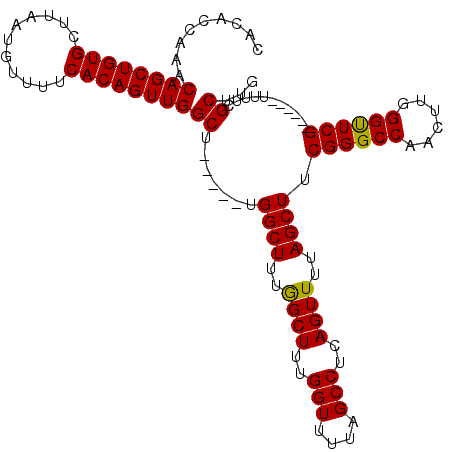
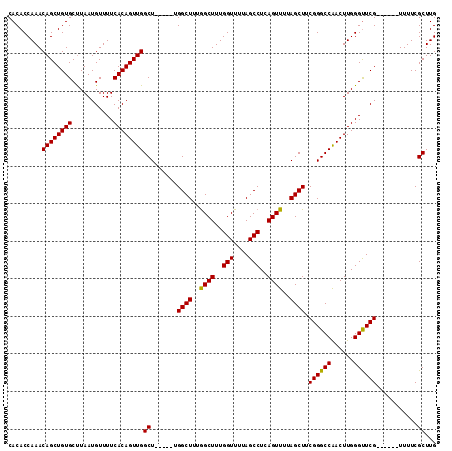
>3L_DroMel_CAF1 1570848 103 + 23771897 CACACCAAACAGCUGUGCUUAAUGUUUUCACAGUUGGCU-----UGGCUUUGGCUUUGGUUUUAGCCUCAGUUUUAGCUUCGGGCCAACUUGGGUUCG------UUUUCGCUUG ..............(((...((((..((((.((((((((-----(((((..((((..(((....)))..))))..)))..))))))))))))))..))------))..)))... ( -32.00) >DroSec_CAF1 15318 103 + 1 CACACCAAACAGCUGUGCCUAAUGUUUUCACAGUUGGCU-----UGGCUUUGGCUUUGGUUUUAGCCUCAGUUUUAGCUUCGGGCCGACUUGGGUUCG------UUUUCGCUUG ..............(((...((((..((((.((((((((-----(((((..((((..(((....)))..))))..)))..))))))))))))))..))------))..)))... ( -31.70) >DroEre_CAF1 21554 103 + 1 CACACCAAACAGCUGUGCUUAAUGUUUUCACAGUUGGCU-----UGGCUUUAGCUUUGGUUUUAGCCUCAGUUUUAGCUGCGGGCCGACUUGGGUUCG------UUUUCGCUUG ..............(((...((((..((((.((((((((-----(((((..((((..(((....)))..))))..)))..))))))))))))))..))------))..)))... ( -31.80) >DroYak_CAF1 15905 114 + 1 CACACCAAACAGCUGUGCUUAAUGUUUUCACAGUUGGCUUGGCUUGGCUUUGGCUUUGGUUUUAGCCUCAGUUUUAGCUUCGGGCCAACUUGGGCUCGUUUACGUUUUCGCUUG ((..((((.((((((((...........))))))))..))))..))(((..((((..(((....)))..))))..)))..((((((......))))))................ ( -34.00) >consensus CACACCAAACAGCUGUGCUUAAUGUUUUCACAGUUGGCU_____UGGCUUUGGCUUUGGUUUUAGCCUCAGUUUUAGCUUCGGGCCAACUUGGGUUCG______UUUUCGCUUG .........((((((((...........))))))))((.......((((..((((..(((....)))..))))..)))).((((((......))))))...........))... (-29.65 = -29.27 + -0.37)

| Location | 1,570,848 – 1,570,951 |
|---|---|
| Length | 103 |
| Sequences | 4 |
| Columns | 114 |
| Reading direction | reverse |
| Mean pairwise identity | 92.47 |
| Mean single sequence MFE | -24.73 |
| Consensus MFE | -20.92 |
| Energy contribution | -21.05 |
| Covariance contribution | 0.13 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.42 |
| Structure conservation index | 0.85 |
| SVM decision value | -0.03 |
| SVM RNA-class probability | 0.516650 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
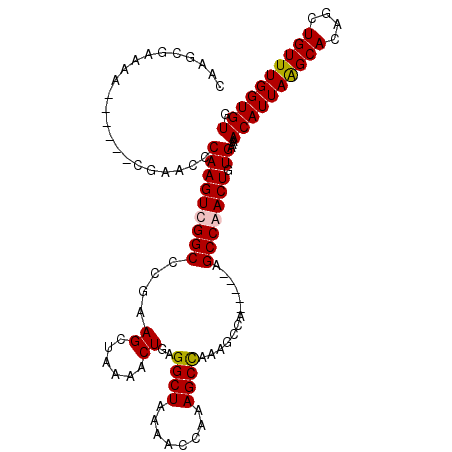
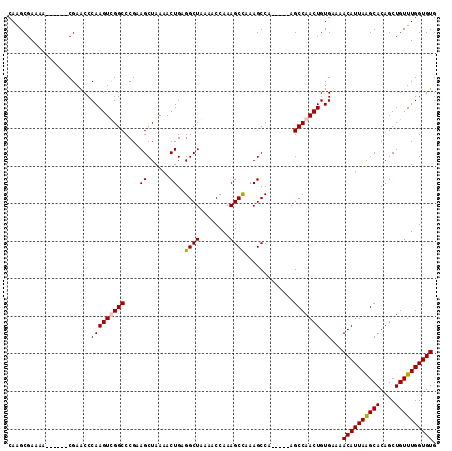
>3L_DroMel_CAF1 1570848 103 - 23771897 CAAGCGAAAA------CGAACCCAAGUUGGCCCGAAGCUAAAACUGAGGCUAAAACCAAAGCCAAAGCCA-----AGCCAACUGUGAAAACAUUAAGCACAGCUGUUUGGUGUG ..........------......(((((((((....((......))..((((........)))).......-----.))))))).))...((((((((((....)))))))))). ( -23.30) >DroSec_CAF1 15318 103 - 1 CAAGCGAAAA------CGAACCCAAGUCGGCCCGAAGCUAAAACUGAGGCUAAAACCAAAGCCAAAGCCA-----AGCCAACUGUGAAAACAUUAGGCACAGCUGUUUGGUGUG ..........------...(((((((.((((............((..((((........))))..))...-----.(((...(((....)))...)))...))))))))).)). ( -24.30) >DroEre_CAF1 21554 103 - 1 CAAGCGAAAA------CGAACCCAAGUCGGCCCGCAGCUAAAACUGAGGCUAAAACCAAAGCUAAAGCCA-----AGCCAACUGUGAAAACAUUAAGCACAGCUGUUUGGUGUG ..(((.....------(((.......)))(((..(((......))).)))..........)))...((((-----(((...(((((...........)))))..)))))))... ( -23.80) >DroYak_CAF1 15905 114 - 1 CAAGCGAAAACGUAAACGAGCCCAAGUUGGCCCGAAGCUAAAACUGAGGCUAAAACCAAAGCCAAAGCCAAGCCAAGCCAACUGUGAAAACAUUAAGCACAGCUGUUUGGUGUG ...(((....))).........(((((((((.....(((....((..((((........))))..))...)))...))))))).))...((((((((((....)))))))))). ( -27.50) >consensus CAAGCGAAAA______CGAACCCAAGUCGGCCCGAAGCUAAAACUGAGGCUAAAACCAAAGCCAAAGCCA_____AGCCAACUGUGAAAACAUUAAGCACAGCUGUUUGGUGUG ......................(((((((((....((......))..((((........)))).............))))))).))...((((((((((....)))))))))). (-20.92 = -21.05 + 0.13)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:44:49 2006