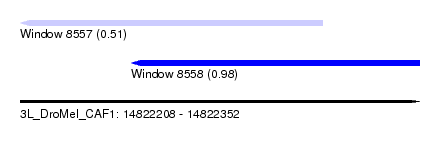
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 14,822,208 – 14,822,352 |
| Length | 144 |
| Max. P | 0.984764 |

| Location | 14,822,208 – 14,822,317 |
|---|---|
| Length | 109 |
| Sequences | 4 |
| Columns | 118 |
| Reading direction | reverse |
| Mean pairwise identity | 88.16 |
| Mean single sequence MFE | -18.20 |
| Consensus MFE | -15.00 |
| Energy contribution | -15.25 |
| Covariance contribution | 0.25 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.35 |
| Structure conservation index | 0.82 |
| SVM decision value | -0.06 |
| SVM RNA-class probability | 0.505401 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 14822208 109 - 23771897 ACCAAAA--------AAAAAGA-AAAAAAAUCAUAAAAAAGAUACCAAGUAAAACAAAUGCUUGGGGCACUUUGGAGCUCGUAUACCCCGUUGUACUAUUAAAUUUACUCAUGGGAAU .(((((.--------.......-......(((........))).(((((((.......))))))).....)))))...........(((((.(((..........)))..)))))... ( -18.10) >DroSec_CAF1 56984 118 - 1 ACCUAAAGAACCUAAAAAAAGAGAAAAAAAUCAUAAAAAAGAUACCAAGUAAAACAAAUGCUUGGGGCACUUUAGAGCUCGUAUACCCCGUUGUACUAUUAAAUUUACCCAUGGGAAU ..((((((..((.................(((........))).(((((((.......)))))))))..))))))...........(((((.(((..........)))..)))))... ( -20.40) >DroSim_CAF1 50174 118 - 1 ACCUAAAGAACCUAAAAAAAGAGAAAAAAAUCAUAAAAAAGAUACCAAGUAAAACAACUGCUUGGGGCACUUUAGAGCUCGUAUACCCCGUUGUACUAUUAAAUUUACCCAUGGGAAU ..((((((..((.................(((........))).(((((((.......)))))))))..))))))...........(((((.(((..........)))..)))))... ( -20.60) >DroAna_CAF1 54433 104 - 1 A-------------AAAAAGGAUGAAAAAAACAUAA-AAAGAUACGAAGUAAAACAAAUGCUUGGGGCACUUUAGAGCUCGUAUACCCCGUUGUACUAUUAAAUUUACCCAUGGGACU .-------------......................-....((((.(((((.......))))).((((........))))))))..(((((.(((..........)))..)))))... ( -13.70) >consensus ACCUAAA_______AAAAAAGAGAAAAAAAUCAUAAAAAAGAUACCAAGUAAAACAAAUGCUUGGGGCACUUUAGAGCUCGUAUACCCCGUUGUACUAUUAAAUUUACCCAUGGGAAU .........................................((((((((((.......))))))((((........))))))))..(((((.(((..........)))..)))))... (-15.00 = -15.25 + 0.25)

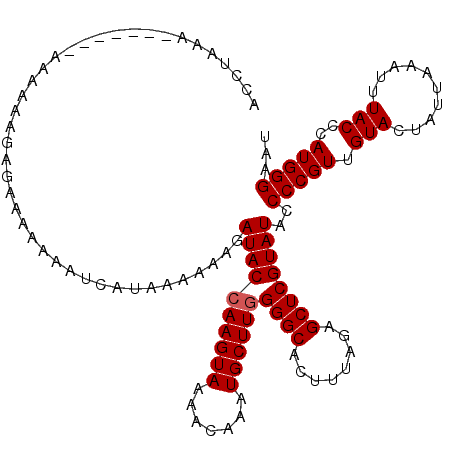
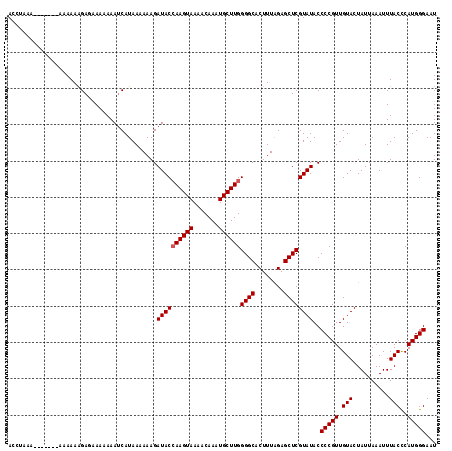
| Location | 14,822,248 – 14,822,352 |
|---|---|
| Length | 104 |
| Sequences | 4 |
| Columns | 118 |
| Reading direction | reverse |
| Mean pairwise identity | 83.53 |
| Mean single sequence MFE | -17.46 |
| Consensus MFE | -14.19 |
| Energy contribution | -13.88 |
| Covariance contribution | -0.31 |
| Combinations/Pair | 1.14 |
| Mean z-score | -2.53 |
| Structure conservation index | 0.81 |
| SVM decision value | 1.98 |
| SVM RNA-class probability | 0.984764 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 14822248 104 - 23771897 AACUUCAUCUAAGCCAAGUC-----AACUUUUUCCCAAAAACCAAAA--------AAAAAGA-AAAAAAAUCAUAAAAAAGAUACCAAGUAAAACAAAUGCUUGGGGCACUUUGGAGC ..(((((.....(((.....-----..((((((..............--------)))))).-.....................(((((((.......))))))))))....))))). ( -17.54) >DroSec_CAF1 57024 113 - 1 AACUUCAUCUAGGCCAAGUC-----AACUUUUUCCCAAAAACCUAAAGAACCUAAAAAAAGAGAAAAAAAUCAUAAAAAAGAUACCAAGUAAAACAAAUGCUUGGGGCACUUUAGAGC .......((((.(((..(((-----..((((((......................)))))).((......))........))).(((((((.......))))))))))....)))).. ( -16.35) >DroSim_CAF1 50214 113 - 1 AACUUCAUCUAAGCCAAGUC-----AACUUUUUCCCAAAAACCUAAAGAACCUAAAAAAAGAGAAAAAAAUCAUAAAAAAGAUACCAAGUAAAACAACUGCUUGGGGCACUUUAGAGC .......((((((((..(((-----..((((((......................)))))).((......))........))).(((((((.......))))))))))...))))).. ( -16.65) >DroAna_CAF1 54473 104 - 1 AACUUCAUCUAAGCCCAGGCUAGCUAACUUUUUUUGAGAAA-------------AAAAAGGAUGAAAAAAACAUAA-AAAGAUACGAAGUAAAACAAAUGCUUGGGGCACUUUAGAGC .......(((((((((((((.......((((((((....))-------------)))))).(((.......)))..-......................)))))))....)))))).. ( -19.30) >consensus AACUUCAUCUAAGCCAAGUC_____AACUUUUUCCCAAAAACCUAAA_______AAAAAAGAGAAAAAAAUCAUAAAAAAGAUACCAAGUAAAACAAAUGCUUGGGGCACUUUAGAGC .......((((((((............((((((......................)))))).......................(((((((.......))))))))))...))))).. (-14.19 = -13.88 + -0.31)

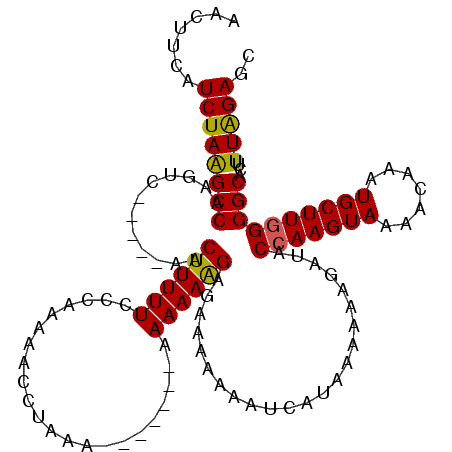
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:49:37 2006