
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 14,816,798 – 14,816,898 |
| Length | 100 |
| Max. P | 0.857183 |

| Location | 14,816,798 – 14,816,898 |
|---|---|
| Length | 100 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 81.51 |
| Mean single sequence MFE | -27.33 |
| Consensus MFE | -18.23 |
| Energy contribution | -18.62 |
| Covariance contribution | 0.39 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.40 |
| Structure conservation index | 0.67 |
| SVM decision value | 0.81 |
| SVM RNA-class probability | 0.857183 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 14816798 100 + 23771897 UUUCU------------CUGCGUAUUUUAUGCUGAUUUUGCCUCAGCUCUUGGGGCCAUCUAUUGUGUGUUUGGAUUUGCACAUUUCCAG--------AUUUUCUGGGGGCAAACACACA ....(------------(.(((((...))))).))....(((((((...))))))).......((((((((((..........(..((((--------.....))))..))))))))))) ( -29.70) >DroSec_CAF1 51696 100 + 1 UUUCU------------CUGCGUAUUUUAUGCUGAUUUUGCCUCAGCUCUUGGGGCCAUCUAUUGUGUGUUUGGAUUUGCACAUUUGCAG--------AUUUUCUGGGCCCAAACACACA ....(------------(.(((((...))))).))....(((((((...))))))).......((((((((((((((((((....)))))--------)).........))))))))))) ( -30.80) >DroSim_CAF1 44810 100 + 1 UUUCU------------CUGCGUAUUUUAUGCUGAUUUUGCCUCAGCUCUUGGGGCCAUCUAUUGUGUGUUUGGAUUUGCACAUUUGCAG--------AUUUUCUGGGGGCAAACACACA ....(------------(.(((((...))))).))....(((((((...))))))).......((((((((((((((((((....)))))--------))).((....)))))))))))) ( -28.90) >DroEre_CAF1 52222 106 + 1 CUUCA------------CUGCGUAUUUUAUGCUGAUUUUGCCUCAACUCUUGGGGCCGUCUAUUGUGUGUUUGGACUUGCACAUUUCCAGAUU--CGGAUUUUCUGUGGGCAAACACACA .....------------..(((((...))))).(((...((((((.....)))))).)))...((((((((((..((..((....(((.....--.))).....))..)))))))))))) ( -31.20) >DroYak_CAF1 54894 108 + 1 CUUCU------------UUGCGUAUUUUAUGCUGAUUUCGCCUCAACUCUUGGGGCCAUCUAUUGUGUGUUUGGAUUUGCACAUUUCCAGAUUUCCAGAUUUUCUGUGGGCAAACACACA ..((.------------..(((((...))))).))....((((((.....)))))).......((((((((((((((((........))))))(((((.....))).)).)))))))))) ( -28.00) >DroAna_CAF1 49262 112 + 1 UUCCUCUUCCUUAUCCUUUGCACAUUUUAUGCCGAUUUUGCUUCAACUCGUGGUGCCAUCUAUUGU--GUUUGGAUUUUCAAAUUUUCGC-CA--CCAACUAACUG---GCAAACAUACA ............((((...(((((.(..(((.((...(..(........)..))).)))..).)))--))..))))............((-((--.........))---))......... ( -15.40) >consensus UUUCU____________CUGCGUAUUUUAUGCUGAUUUUGCCUCAACUCUUGGGGCCAUCUAUUGUGUGUUUGGAUUUGCACAUUUCCAG________AUUUUCUGGGGGCAAACACACA ...................(((((...))))).(((...((((((.....)))))).)))...((((((((((.....................................)))))))))) (-18.23 = -18.62 + 0.39)


| Location | 14,816,798 – 14,816,898 |
|---|---|
| Length | 100 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 81.51 |
| Mean single sequence MFE | -24.19 |
| Consensus MFE | -15.34 |
| Energy contribution | -16.01 |
| Covariance contribution | 0.67 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.97 |
| Structure conservation index | 0.63 |
| SVM decision value | 0.36 |
| SVM RNA-class probability | 0.705893 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 14816798 100 - 23771897 UGUGUGUUUGCCCCCAGAAAAU--------CUGGAAAUGUGCAAAUCCAAACACACAAUAGAUGGCCCCAAGAGCUGAGGCAAAAUCAGCAUAAAAUACGCAG------------AGAAA ((((((((((((.((((.....--------))))...((((............))))......))).......(((((.......)))))...))))))))).------------..... ( -26.20) >DroSec_CAF1 51696 100 - 1 UGUGUGUUUGGGCCCAGAAAAU--------CUGCAAAUGUGCAAAUCCAAACACACAAUAGAUGGCCCCAAGAGCUGAGGCAAAAUCAGCAUAAAAUACGCAG------------AGAAA (((((((((((((((((.....--------)))....((((............))))......))))).....(((((.......)))))...))))))))).------------..... ( -28.80) >DroSim_CAF1 44810 100 - 1 UGUGUGUUUGCCCCCAGAAAAU--------CUGCAAAUGUGCAAAUCCAAACACACAAUAGAUGGCCCCAAGAGCUGAGGCAAAAUCAGCAUAAAAUACGCAG------------AGAAA ((((((((((.....((.....--------))(((....))).....))))))))))...(((.(((.((.....)).)))...))).((.........))..------------..... ( -22.20) >DroEre_CAF1 52222 106 - 1 UGUGUGUUUGCCCACAGAAAAUCCG--AAUCUGGAAAUGUGCAAGUCCAAACACACAAUAGACGGCCCCAAGAGUUGAGGCAAAAUCAGCAUAAAAUACGCAG------------UGAAG ((((((((((((((((.....((((--....))))..))))...(((.............)))))).......(((((.......)))))...))))))))).------------..... ( -23.12) >DroYak_CAF1 54894 108 - 1 UGUGUGUUUGCCCACAGAAAAUCUGGAAAUCUGGAAAUGUGCAAAUCCAAACACACAAUAGAUGGCCCCAAGAGUUGAGGCGAAAUCAGCAUAAAAUACGCAA------------AGAAG ((((((((((..((((.....((..(....)..))..))))......))))))))))...(((.(((.((.....)).)))...))).((.........))..------------..... ( -25.00) >DroAna_CAF1 49262 112 - 1 UGUAUGUUUGC---CAGUUAGUUGG--UG-GCGAAAAUUUGAAAAUCCAAAC--ACAAUAGAUGGCACCACGAGUUGAAGCAAAAUCGGCAUAAAAUGUGCAAAGGAUAAGGAAGAGGAA ......(((((---((.(.....).--))-))))).........((((....--......(((.((..((.....))..))...))).((((.....))))...))))............ ( -19.80) >consensus UGUGUGUUUGCCCCCAGAAAAU________CUGGAAAUGUGCAAAUCCAAACACACAAUAGAUGGCCCCAAGAGCUGAGGCAAAAUCAGCAUAAAAUACGCAG____________AGAAA ((((((((((.....................................))))))))))...(((.(((.((.....)).)))...))).((.........))................... (-15.34 = -16.01 + 0.67)
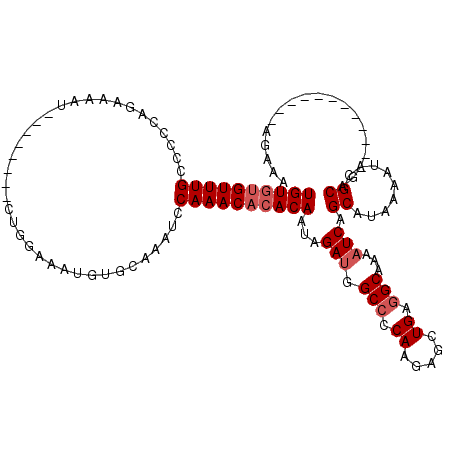



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:49:23 2006