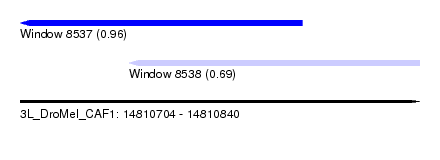
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 14,810,704 – 14,810,840 |
| Length | 136 |
| Max. P | 0.956936 |

| Location | 14,810,704 – 14,810,800 |
|---|---|
| Length | 96 |
| Sequences | 5 |
| Columns | 97 |
| Reading direction | reverse |
| Mean pairwise identity | 94.32 |
| Mean single sequence MFE | -25.97 |
| Consensus MFE | -21.46 |
| Energy contribution | -21.86 |
| Covariance contribution | 0.40 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.98 |
| Structure conservation index | 0.83 |
| SVM decision value | 1.47 |
| SVM RNA-class probability | 0.956936 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
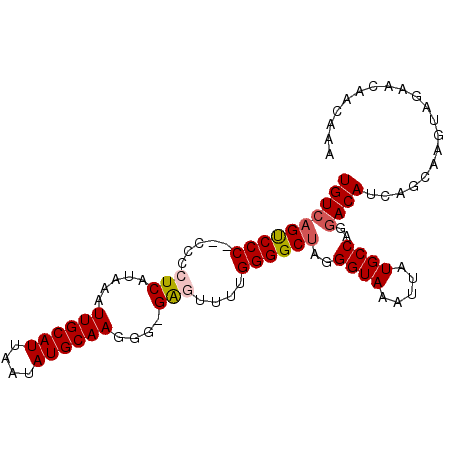
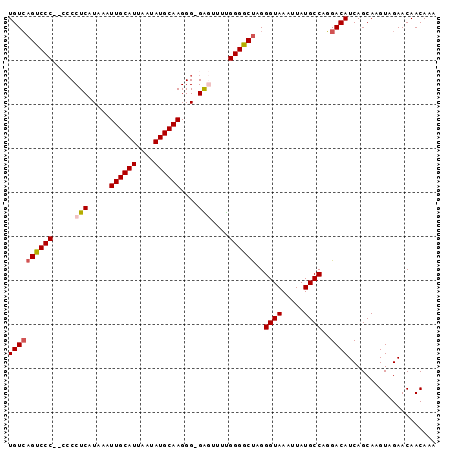
>3L_DroMel_CAF1 14810704 96 - 23771897 UGUAAGCCCCCCCCCCUCAUAAAUUGCAUUAAUAUGCAAGGG-GAGUUUUGGGGCUAGGGUAAAUUAUGCCAUGACAUCAGCAAGUAGAACAACAAA (((.((((((.((((((.......(((((....)))))))))-).)....))))))..((((.....)))).........))).............. ( -26.41) >DroSec_CAF1 45932 94 - 1 UGUCAGUCCC--CCCCUCAUAAAUUGCAUUAAUAUGCAAGGG-GAGUUUUGGGGCUAGGGUAAAUUAUGCCAGGACAUCAGCAAGUAGAACAACAAA ((((((((((--(((((.......(((((....)))))))))-)......))))))..((((.....))))..)))).................... ( -26.21) >DroSim_CAF1 38880 94 - 1 UGUCAGUCCC--CCCCUCAUAAAUUGCAUUAAUAUGCAAGGG-GAGUUUUGGGGCUAGGGUAAAUUAUGCCAGGACAUCAGCAAGUAGAACAACAAA ((((((((((--(((((.......(((((....)))))))))-)......))))))..((((.....))))..)))).................... ( -26.21) >DroEre_CAF1 45312 93 - 1 UGUCAGUCCC---CCCUCAUAAAUUGCAUUAAUAUGCAAGGG-GGUUUUUGGGGCCAGGGUAAAUUAUGCCAGGACAUCAGCAAGUAGAACAACAAA ((((.(((((---(((((.....((((((....)))))))))-)).....)))))...((((.....))))..)))).................... ( -25.20) >DroYak_CAF1 48840 93 - 1 UGUCAGUCCC----CCUCAUAAAUUGCAUUAAUAUGCAAGGGGGGUUUUCGGGGCUAGGGUAAAUUAUGCCAGGACAUCAGCAAGUAGAACAACAAA ((((((((((----((((.....((((((....))))))..)))).....))))))..((((.....))))..)))).................... ( -25.80) >consensus UGUCAGUCCC__CCCCUCAUAAAUUGCAUUAAUAUGCAAGGG_GAGUUUUGGGGCUAGGGUAAAUUAUGCCAGGACAUCAGCAAGUAGAACAACAAA ((((((((((.....(((.....((((((....))))))....)))....))))))..((((.....))))..)))).................... (-21.46 = -21.86 + 0.40)

| Location | 14,810,741 – 14,810,840 |
|---|---|
| Length | 99 |
| Sequences | 5 |
| Columns | 100 |
| Reading direction | reverse |
| Mean pairwise identity | 94.09 |
| Mean single sequence MFE | -32.17 |
| Consensus MFE | -25.37 |
| Energy contribution | -25.45 |
| Covariance contribution | 0.08 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.69 |
| Structure conservation index | 0.79 |
| SVM decision value | 0.32 |
| SVM RNA-class probability | 0.685853 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
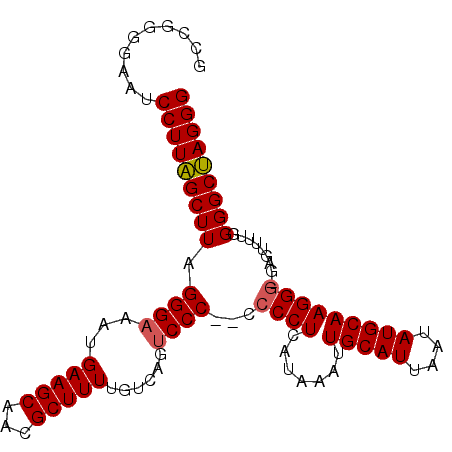
>3L_DroMel_CAF1 14810741 99 - 23771897 GCCGGGGAAUCCUUAGCUUAGGGAAAUGAAGCAACGCUUUUGUAAGCCCCCCCCCCUCAUAAAUUGCAUUAAUAUGCAAGGG-GAGUUUUGGGGCUAGGG (((((((..((((((...))))))...........((((....)))))))).(((((.......(((((....)))))))))-)........)))..... ( -31.31) >DroSec_CAF1 45969 97 - 1 GCCGGGGAAUCCUUAGCUUAGGGAAAUGAAGCAACGCUUUUGUCAGUCCC--CCCCUCAUAAAUUGCAUUAAUAUGCAAGGG-GAGUUUUGGGGCUAGGG ..........(((((((((.((((...(((((...)))))......))))--(((((.......(((((....)))))))))-).......))))))))) ( -31.71) >DroSim_CAF1 38917 97 - 1 GCCGGGGAAUCCUUAGCUUAGGGAAAUGAAGCAACGCUUUUGUCAGUCCC--CCCCUCAUAAAUUGCAUUAAUAUGCAAGGG-GAGUUUUGGGGCUAGGG ..........(((((((((.((((...(((((...)))))......))))--(((((.......(((((....)))))))))-).......))))))))) ( -31.71) >DroEre_CAF1 45349 96 - 1 GCCAGGGAAUCCUUGGCUUAGGGAAAUGAAGCAACGCUUUUGUCAGUCCC---CCCUCAUAAAUUGCAUUAAUAUGCAAGGG-GGUUUUUGGGGCCAGGG ..........(((((((((.((((...(((((...)))))......))))---(((((.....((((((....)))))))))-))......))))))))) ( -33.40) >DroYak_CAF1 48877 96 - 1 GCCGGGGAAUCCUUAGCUUAGGGAAAUGAAGCAACGCUUUUGUCAGUCCC----CCUCAUAAAUUGCAUUAAUAUGCAAGGGGGGUUUUCGGGGCUAGGG ..........(((((((((..(((((((((((...))))).......(((----(((.......(((((....))))))))))))))))).))))))))) ( -32.71) >consensus GCCGGGGAAUCCUUAGCUUAGGGAAAUGAAGCAACGCUUUUGUCAGUCCC__CCCCUCAUAAAUUGCAUUAAUAUGCAAGGG_GAGUUUUGGGGCUAGGG ..........(((((((((.((((...(((((...)))))......))))...((((.......(((((....))))))))).........))))))))) (-25.37 = -25.45 + 0.08)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:49:17 2006