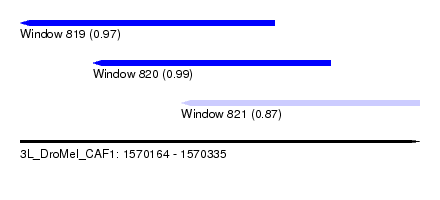

| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 1,570,164 – 1,570,335 |
| Length | 171 |
| Max. P | 0.988959 |

| Location | 1,570,164 – 1,570,273 |
|---|---|
| Length | 109 |
| Sequences | 5 |
| Columns | 110 |
| Reading direction | reverse |
| Mean pairwise identity | 92.14 |
| Mean single sequence MFE | -28.32 |
| Consensus MFE | -25.42 |
| Energy contribution | -25.22 |
| Covariance contribution | -0.20 |
| Combinations/Pair | 1.11 |
| Mean z-score | -1.64 |
| Structure conservation index | 0.90 |
| SVM decision value | 1.65 |
| SVM RNA-class probability | 0.969663 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
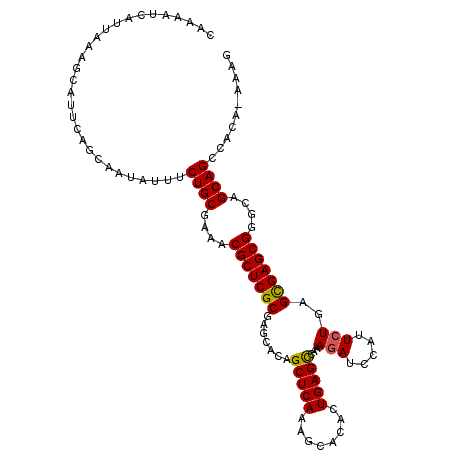
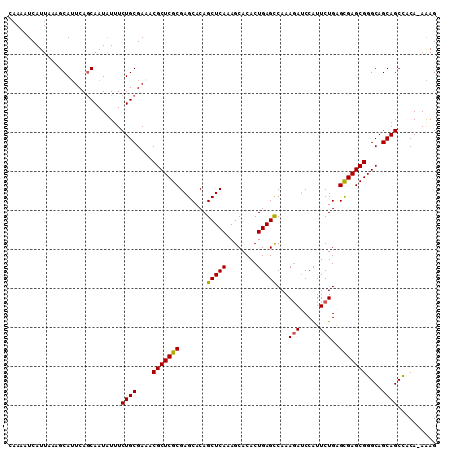
>3L_DroMel_CAF1 1570164 109 - 23771897 CAAAAUCAUUAAAGCAUUCAGCAAUCUUUCUGCGAAACGCUCGCGAGCACAGCUCAAAGCACACUGAGCCAAAGAUCCAUUCUGAGCGAGCGGGCAGCAGCCACA-AAAG .............((.....)).......((((....(((((((.......(((((........)))))...(((.....)))..)))))))....)))).....-.... ( -29.00) >DroSec_CAF1 14618 109 - 1 CAAAAUCAUUAAAGCAUUCAGCAAUAUAUCUGCGAAACGCUCGCGAGCACAGCUCAAAGCACACUGAGCCAAAAAGCCAUUCUGAGCGAGCGGGCAGCAGCCACA-AAAG .............((.....)).......((((....(((((((((((...)))).........((.((......))))......)))))))....)))).....-.... ( -27.60) >DroSim_CAF1 14871 109 - 1 CAAAAUCAUUCAAGCAUUCAGCAAUAUUUCUGCGAAACGCUCGCGAGCACAGCUCAAAGCACACUGAGCCAAAGAUCCAUUCUGAGCGAGCGGGCAGCAGCCACA-AAAG .............((.....)).......((((....(((((((.......(((((........)))))...(((.....)))..)))))))....)))).....-.... ( -29.00) >DroEre_CAF1 20827 109 - 1 UAAAAUCAUUUAGGCACUCAACAAUAUUUCUGCGAACCGCUCGCGAGCACAGCUCAAAGCACACUGAGCCAAAGAUCCGUUCUGAGUGAGCGGGCAGCAGCUUCA-AGAG .............................((((...((((((((.......(((((........)))))...(((.....)))..))))))))...)))).....-.... ( -29.50) >DroYak_CAF1 15168 110 - 1 CAAAAUCAUUAAAGCAUUCAGCAAUAUUUCUGCGAUACGCUCGCGAGCACAGCUCAAAGCACACUGAGUCAAAGAUCCAUUCUGGGUGAGCGGGCAGCAGCCCCAAAGAG .....((......((.....((.........((((.....))))((((...))))...)).((((.((.............)).)))).))((((....))))....)). ( -26.52) >consensus CAAAAUCAUUAAAGCAUUCAGCAAUAUUUCUGCGAAACGCUCGCGAGCACAGCUCAAAGCACACUGAGCCAAAGAUCCAUUCUGAGCGAGCGGGCAGCAGCCACA_AAAG .............................((((....(((((((.......(((((........)))))...(((.....)))..)))))))....)))).......... (-25.42 = -25.22 + -0.20)

| Location | 1,570,195 – 1,570,297 |
|---|---|
| Length | 102 |
| Sequences | 5 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 95.00 |
| Mean single sequence MFE | -22.03 |
| Consensus MFE | -21.37 |
| Energy contribution | -21.41 |
| Covariance contribution | 0.04 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.56 |
| Structure conservation index | 0.97 |
| SVM decision value | 2.14 |
| SVM RNA-class probability | 0.988959 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
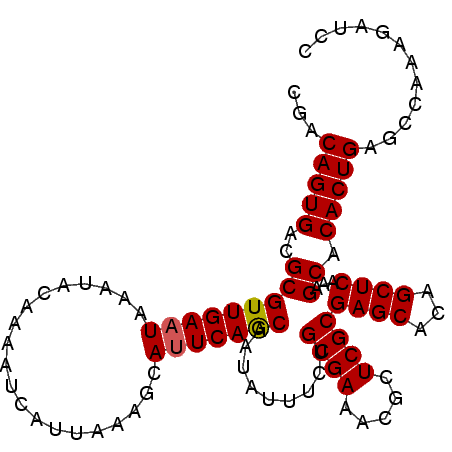
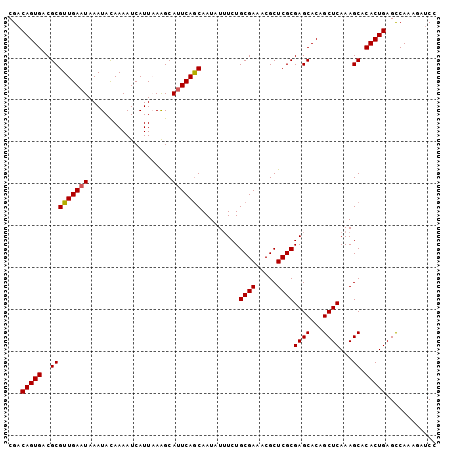
>3L_DroMel_CAF1 1570195 102 - 23771897 CGACAGUGACGCGUUGAAUAAAUACAAAAUCAUUAAAGCAUUCAGCAAUCUUUCUGCGAAACGCUCGCGAGCACAGCUCAAAGCACACUGAGCCAAAGAUCC ...(((((..(((((((((....................))))))).........((((.....))))((((...))))...)).)))))............ ( -21.85) >DroSec_CAF1 14649 102 - 1 CGACAGUGACGCGUUGAAUAAAUACAAAAUCAUUAAAGCAUUCAGCAAUAUAUCUGCGAAACGCUCGCGAGCACAGCUCAAAGCACACUGAGCCAAAAAGCC ...(((((..(((((((((....................))))))).........((((.....))))((((...))))...)).)))))............ ( -21.85) >DroSim_CAF1 14902 102 - 1 CGACAGUGACGCGUUGAAUAAAUACAAAAUCAUUCAAGCAUUCAGCAAUAUUUCUGCGAAACGCUCGCGAGCACAGCUCAAAGCACACUGAGCCAAAGAUCC ...(((((..((.((((((............)))))))).....((.........((((.....))))((((...))))...)).)))))............ ( -23.40) >DroEre_CAF1 20858 102 - 1 CGACAGUGACGCGUUGAAUAAAUAUAAAAUCAUUUAGGCACUCAACAAUAUUUCUGCGAACCGCUCGCGAGCACAGCUCAAAGCACACUGAGCCAAAGAUCC ...(((((..(((((((.(((((........))))).....))))).........((((.....))))((((...))))...)).)))))............ ( -21.40) >DroYak_CAF1 15200 102 - 1 CGACAGUGACGCGUUGAAUAAAUACAAAAUCAUUAAAGCAUUCAGCAAUAUUUCUGCGAUACGCUCGCGAGCACAGCUCAAAGCACACUGAGUCAAAGAUCC ...(((((..(((((((((....................))))))).........((((.....))))((((...))))...)).)))))............ ( -21.65) >consensus CGACAGUGACGCGUUGAAUAAAUACAAAAUCAUUAAAGCAUUCAGCAAUAUUUCUGCGAAACGCUCGCGAGCACAGCUCAAAGCACACUGAGCCAAAGAUCC ...(((((..(((((((((....................))))))).........((((.....))))((((...))))...)).)))))............ (-21.37 = -21.41 + 0.04)

| Location | 1,570,233 – 1,570,335 |
|---|---|
| Length | 102 |
| Sequences | 5 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 93.53 |
| Mean single sequence MFE | -23.16 |
| Consensus MFE | -19.22 |
| Energy contribution | -19.66 |
| Covariance contribution | 0.44 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.66 |
| Structure conservation index | 0.83 |
| SVM decision value | 0.88 |
| SVM RNA-class probability | 0.873409 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 1570233 102 - 23771897 CCUAAUUUACGAUCUAUUCGGUUUUAUGACAACGCGCGCGACAGUGACGCGUUGAAUAAAUACAAAAUCAUUAAAGCAUUCAGCAAUCUUUCUGCGAAACGC .........(((.....)))(((((((((((((((((((....))).))))))).............))).)))))).(((.(((.......)))))).... ( -22.21) >DroSec_CAF1 14687 102 - 1 CCUAAUUUACGAUCUAUUUGGUUUUAUCGCAACGCGCGCGACAGUGACGCGUUGAAUAAAUACAAAAUCAUUAAAGCAUUCAGCAAUAUAUCUGCGAAACGC ..........(((...((((..(((((..((((((((((....))).))))))).)))))..)))))))......((.((..(((.......)))..)).)) ( -22.60) >DroSim_CAF1 14940 102 - 1 CCUAAUUUACGAUCUAUUUGGUUUUAUGGCAACGCGCGCGACAGUGACGCGUUGAAUAAAUACAAAAUCAUUCAAGCAUUCAGCAAUAUUUCUGCGAAACGC ..........(((...((((..(((((..((((((((((....))).))))))).)))))..)))))))......((.((..(((.......)))..)).)) ( -23.80) >DroEre_CAF1 20896 102 - 1 CCUAAUUUACGAUCUUUUUGAUUUUACGUCAACGCGCGCGACAGUGACGCGUUGAAUAAAUAUAAAAUCAUUUAGGCACUCAACAAUAUUUCUGCGAACCGC ..........(.(((...((((((((..(((((((((((....))).)))))))).......))))))))...))))................(((...))) ( -22.70) >DroYak_CAF1 15238 102 - 1 CCUAAUUUACGAUCUAUUUGGUUUUAUGGCAACGCGCGCGACAGUGACGCGUUGAAUAAAUACAAAAUCAUUAAAGCAUUCAGCAAUAUUUCUGCGAUACGC .........(((((..((((..(((((..((((((((((....))).))))))).)))))..))))................(((.......)))))).)). ( -24.50) >consensus CCUAAUUUACGAUCUAUUUGGUUUUAUGGCAACGCGCGCGACAGUGACGCGUUGAAUAAAUACAAAAUCAUUAAAGCAUUCAGCAAUAUUUCUGCGAAACGC ..........(((...((((..(((((..((((((((((....))).))))))).)))))..)))))))......((.....(((.......))).....)) (-19.22 = -19.66 + 0.44)
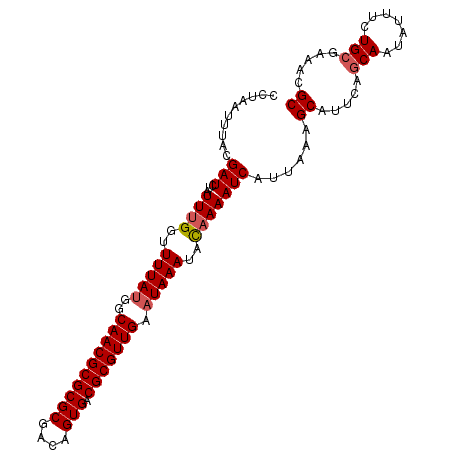


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:44:46 2006