
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 14,755,325 – 14,755,466 |
| Length | 141 |
| Max. P | 0.999504 |

| Location | 14,755,325 – 14,755,426 |
|---|---|
| Length | 101 |
| Sequences | 5 |
| Columns | 101 |
| Reading direction | forward |
| Mean pairwise identity | 92.87 |
| Mean single sequence MFE | -29.37 |
| Consensus MFE | -21.82 |
| Energy contribution | -22.22 |
| Covariance contribution | 0.40 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.32 |
| Structure conservation index | 0.74 |
| SVM decision value | 0.63 |
| SVM RNA-class probability | 0.804847 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 14755325 101 + 23771897 GCAAAGGUAACGCAAAAAGAGUGCAAAAGCCCGAAAUGUACGCACAAAUUGGCCAGCAAGUGUACAGCUGGCUGGCUGACUUGCCAUAAACGCGCUGGAAA ((.........))......(((((...((((.(...(((((((................))))))).).))))(((......)))......)))))..... ( -26.49) >DroSec_CAF1 7277 101 + 1 GCAAAGGUAACGCAAAAAGAGUGCAAAAGCCCGAAAUGUACGCACAAAUUGGCCAGCAAGUGUACAGCUGGCUGGCUGACUUGCCAUAAACGCGCUGUAAA ((.........))....(.(((((...((((.(...(((((((................))))))).).))))(((......)))......))))).)... ( -26.79) >DroSim_CAF1 7187 101 + 1 GCAAAGGUAACGCAAAAAGAGUGCAAAAGCCCGAAAUGUACGCACAAAUUGGCCAGCAAGUGUACAGCUGGCUGGCUGACUUGCCAUAAACGCGCUGUUAA ((.........))...((.(((((...((((.(...(((((((................))))))).).))))(((......)))......))))).)).. ( -27.69) >DroEre_CAF1 4633 101 + 1 GCAAAGGUAACGCAGAAAGAGUGCAAAAGUCCGAAAUGGACGCAAAAAUUGGCCAGCAAAUGUACAGCUGGCUGGCUGACUUGCGAUAAAAGCGCUGGAGA ((.........))......(((((....((((.....))))((((...(..((((((....(.....)..))))))..).)))).......)))))..... ( -30.30) >DroYak_CAF1 11178 101 + 1 GCAAAGGUAACGCAACAAGAGUGCAAAAACCCGAAAGGGACGCAAUAAUUGGCCAGCAAAUGUACAGCUGGCUGGCUGACUUGCGAUAAAUGCGCUGGAUA ((.........))......((((((....(((....))).(((((...(..((((((....(.....)..))))))..).))))).....))))))..... ( -35.60) >consensus GCAAAGGUAACGCAAAAAGAGUGCAAAAGCCCGAAAUGUACGCACAAAUUGGCCAGCAAGUGUACAGCUGGCUGGCUGACUUGCCAUAAACGCGCUGGAAA ((.........))......(((((..........................(((((((.........)))))))(((......)))......)))))..... (-21.82 = -22.22 + 0.40)


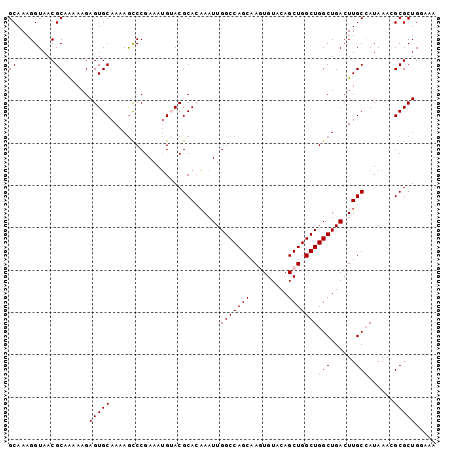
| Location | 14,755,325 – 14,755,426 |
|---|---|
| Length | 101 |
| Sequences | 5 |
| Columns | 101 |
| Reading direction | reverse |
| Mean pairwise identity | 92.87 |
| Mean single sequence MFE | -27.36 |
| Consensus MFE | -24.84 |
| Energy contribution | -24.96 |
| Covariance contribution | 0.12 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.68 |
| Structure conservation index | 0.91 |
| SVM decision value | 1.92 |
| SVM RNA-class probability | 0.982682 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 14755325 101 - 23771897 UUUCCAGCGCGUUUAUGGCAAGUCAGCCAGCCAGCUGUACACUUGCUGGCCAAUUUGUGCGUACAUUUCGGGCUUUUGCACUCUUUUUGCGUUACCUUUGC .....((((((......(((((..((((.((((((.........)))))).........((.......)))))))))))........))))))........ ( -27.54) >DroSec_CAF1 7277 101 - 1 UUUACAGCGCGUUUAUGGCAAGUCAGCCAGCCAGCUGUACACUUGCUGGCCAAUUUGUGCGUACAUUUCGGGCUUUUGCACUCUUUUUGCGUUACCUUUGC .....((((((......(((((..((((.((((((.........)))))).........((.......)))))))))))........))))))........ ( -27.54) >DroSim_CAF1 7187 101 - 1 UUAACAGCGCGUUUAUGGCAAGUCAGCCAGCCAGCUGUACACUUGCUGGCCAAUUUGUGCGUACAUUUCGGGCUUUUGCACUCUUUUUGCGUUACCUUUGC .(((((((.((..((((((((((..((((((.((.......)).))))))..)))))).)))).....)).)))...(((.......)))))))....... ( -27.60) >DroEre_CAF1 4633 101 - 1 UCUCCAGCGCUUUUAUCGCAAGUCAGCCAGCCAGCUGUACAUUUGCUGGCCAAUUUUUGCGUCCAUUUCGGACUUUUGCACUCUUUCUGCGUUACCUUUGC .....(((((.......(((((.......((((((.........)))))).....)))))((((.....))))...............)))))........ ( -26.50) >DroYak_CAF1 11178 101 - 1 UAUCCAGCGCAUUUAUCGCAAGUCAGCCAGCCAGCUGUACAUUUGCUGGCCAAUUAUUGCGUCCCUUUCGGGUUUUUGCACUCUUGUUGCGUUACCUUUGC .....((((((......(((((.......((((((.........)))))).......((((.(((....)))....))))..)))))))))))........ ( -27.60) >consensus UUUCCAGCGCGUUUAUGGCAAGUCAGCCAGCCAGCUGUACACUUGCUGGCCAAUUUGUGCGUACAUUUCGGGCUUUUGCACUCUUUUUGCGUUACCUUUGC .....((((((......(((((..((((.((((((.........)))))).........((.......)))))))))))........))))))........ (-24.84 = -24.96 + 0.12)



| Location | 14,755,357 – 14,755,466 |
|---|---|
| Length | 109 |
| Sequences | 5 |
| Columns | 109 |
| Reading direction | forward |
| Mean pairwise identity | 92.11 |
| Mean single sequence MFE | -39.00 |
| Consensus MFE | -32.78 |
| Energy contribution | -33.30 |
| Covariance contribution | 0.52 |
| Combinations/Pair | 1.11 |
| Mean z-score | -3.23 |
| Structure conservation index | 0.84 |
| SVM decision value | 3.66 |
| SVM RNA-class probability | 0.999504 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 14755357 109 + 23771897 GAAAUGUACGCACAAAUUGGCCAGCAAGUGUACAGCUGGCUGGCUGACUUGCCAUAAACGCGCUGGAAAGGGUUCAGCGCAGUAAAUUUUGCUUGCUGAGCAUGCAAAA ......(((...(((.(..((((((.(((.....))).))))))..).)))........((((((((.....)))))))).)))..((((((.(((...))).)))))) ( -39.60) >DroSec_CAF1 7309 109 + 1 GAAAUGUACGCACAAAUUGGCCAGCAAGUGUACAGCUGGCUGGCUGACUUGCCAUAAACGCGCUGUAAAGGGUUUAGCGCAGUAAAUGUCGCUUGCUGAGCAUGCAAAA .........((((.....)(((((((((((.(((....((((((......)))......((((((.........)))))))))...)))))))))))).)).))).... ( -37.90) >DroSim_CAF1 7219 109 + 1 GAAAUGUACGCACAAAUUGGCCAGCAAGUGUACAGCUGGCUGGCUGACUUGCCAUAAACGCGCUGUUAAGGGUUCAGCGCAGUAAAUGCCGCUUGCUGAGCAUGCAAAA .........((((.....)(((((((((((....((..((((((......)))......((((((.........)))))))))....))))))))))).)).))).... ( -36.30) >DroEre_CAF1 4665 109 + 1 GAAAUGGACGCAAAAAUUGGCCAGCAAAUGUACAGCUGGCUGGCUGACUUGCGAUAAAAGCGCUGGAGAAGGUUCAGCGCUGUAAAUGUCGCUUGCUGAGCAUGCAAAA .........(((......(((((((.........))))))).(((.((..((((((..(((((((((.....))))))))).....))))))..).).))).))).... ( -41.30) >DroYak_CAF1 11210 109 + 1 GAAAGGGACGCAAUAAUUGGCCAGCAAAUGUACAGCUGGCUGGCUGACUUGCGAUAAAUGCGCUGGAUAAGGUUCAGCGUAGUAAAUGUCGCUUGCUGAGCAUGCAAAA .........(((......(((((((.........))))))).(((.((..((((((..((((((((((...)))))))))).....))))))..).).))).))).... ( -39.90) >consensus GAAAUGUACGCACAAAUUGGCCAGCAAGUGUACAGCUGGCUGGCUGACUUGCCAUAAACGCGCUGGAAAGGGUUCAGCGCAGUAAAUGUCGCUUGCUGAGCAUGCAAAA .........(((.......(((((((((((..(((((....))))).............((((((((.....)))))))).........))))))))).)).))).... (-32.78 = -33.30 + 0.52)

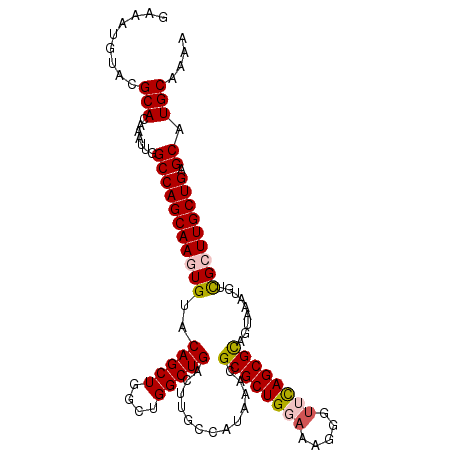

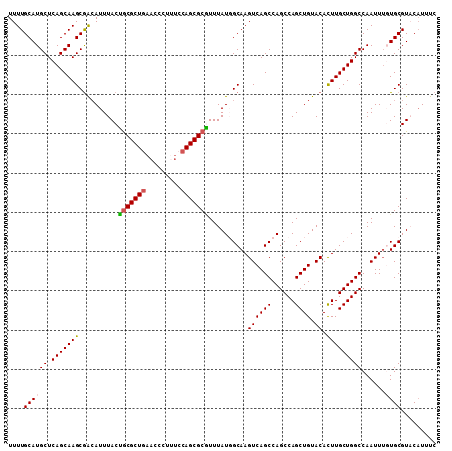
| Location | 14,755,357 – 14,755,466 |
|---|---|
| Length | 109 |
| Sequences | 5 |
| Columns | 109 |
| Reading direction | reverse |
| Mean pairwise identity | 92.11 |
| Mean single sequence MFE | -32.89 |
| Consensus MFE | -28.40 |
| Energy contribution | -28.52 |
| Covariance contribution | 0.12 |
| Combinations/Pair | 1.12 |
| Mean z-score | -2.56 |
| Structure conservation index | 0.86 |
| SVM decision value | 3.32 |
| SVM RNA-class probability | 0.999005 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 14755357 109 - 23771897 UUUUGCAUGCUCAGCAAGCAAAAUUUACUGCGCUGAACCCUUUCCAGCGCGUUUAUGGCAAGUCAGCCAGCCAGCUGUACACUUGCUGGCCAAUUUGUGCGUACAUUUC ((((((.(((...))).))))))......((((((.........))))))...((((((((((..((((((.((.......)).))))))..)))))).))))...... ( -35.70) >DroSec_CAF1 7309 109 - 1 UUUUGCAUGCUCAGCAAGCGACAUUUACUGCGCUAAACCCUUUACAGCGCGUUUAUGGCAAGUCAGCCAGCCAGCUGUACACUUGCUGGCCAAUUUGUGCGUACAUUUC ..((((.(((...))).))))(((..((.(((((...........)))))))..)))((((((..((((((.((.......)).))))))..))))))........... ( -31.80) >DroSim_CAF1 7219 109 - 1 UUUUGCAUGCUCAGCAAGCGGCAUUUACUGCGCUGAACCCUUAACAGCGCGUUUAUGGCAAGUCAGCCAGCCAGCUGUACACUUGCUGGCCAAUUUGUGCGUACAUUUC ....((((((.(((((((.(.(((..((.((((((.........))))))))..))).)..((((((......)))).)).)))))))))......))))......... ( -36.60) >DroEre_CAF1 4665 109 - 1 UUUUGCAUGCUCAGCAAGCGACAUUUACAGCGCUGAACCUUCUCCAGCGCUUUUAUCGCAAGUCAGCCAGCCAGCUGUACAUUUGCUGGCCAAUUUUUGCGUCCAUUUC ....((.(((...))).))(((......(((((((.........)))))))......(((((.......((((((.........)))))).....))))))))...... ( -32.60) >DroYak_CAF1 11210 109 - 1 UUUUGCAUGCUCAGCAAGCGACAUUUACUACGCUGAACCUUAUCCAGCGCAUUUAUCGCAAGUCAGCCAGCCAGCUGUACAUUUGCUGGCCAAUUAUUGCGUCCCUUUC ....((.(((...))).))(((........(((((.........)))))........((((........((((((.........))))))......)))))))...... ( -27.74) >consensus UUUUGCAUGCUCAGCAAGCGACAUUUACUGCGCUGAACCCUUUCCAGCGCGUUUAUGGCAAGUCAGCCAGCCAGCUGUACACUUGCUGGCCAAUUUGUGCGUACAUUUC ....((((((.(((((((..........(((((((.........)))))))..........((((((......)))).)).)))))))))......))))......... (-28.40 = -28.52 + 0.12)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:47:53 2006