
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 14,658,130 – 14,658,230 |
| Length | 100 |
| Max. P | 0.933213 |

| Location | 14,658,130 – 14,658,230 |
|---|---|
| Length | 100 |
| Sequences | 3 |
| Columns | 100 |
| Reading direction | forward |
| Mean pairwise identity | 95.33 |
| Mean single sequence MFE | -19.76 |
| Consensus MFE | -17.46 |
| Energy contribution | -18.24 |
| Covariance contribution | 0.78 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.07 |
| Structure conservation index | 0.88 |
| SVM decision value | 0.38 |
| SVM RNA-class probability | 0.712608 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
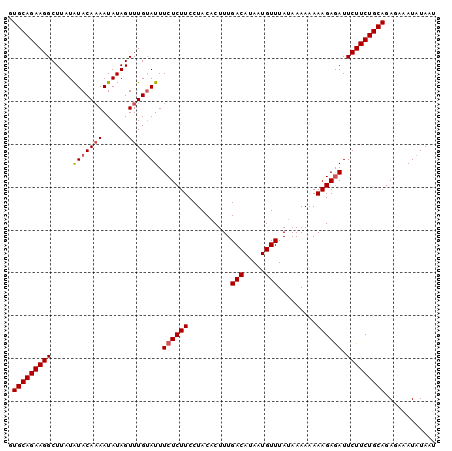
>3L_DroMel_CAF1 14658130 100 + 23771897 GUGCAGAAGGCUUAUAUACAUAAUAUAGUUUGAAUUUCUCUUCCUACCCUUUGACAUAAUGUUUAAAAAAAAAAGAAAUUCUUCUGCAGAGAAAUAUAAU .(((((((((((.((((.....)))))))))((((((((.............(((.....)))..........)))))))).))))))............ ( -16.60) >DroSec_CAF1 7910 99 + 1 GUGCAGAAGGCUUAUGUACAAAAUAUAGUUUGUAUUUCUCUUCCUACACUUUGACAUAAUGUUUAUAAAA-AAAGAGAUUCUUCUGCAGAGAAAUAUAAU .(((((((((.....(((((((......))))))).((((((..........(((.....))).......-.)))))).)))))))))............ ( -21.27) >DroSim_CAF1 7959 100 + 1 GUGCAGAAGGCUUAUAUACAAAAUAUAGUUUGUAUUUCUCUUCCUACACUUUGACAUAAUGUUUAUAAAAAAAAGAGAUUCUUCUGCAGAGAAAUAUAAU .(((((((((.....(((((((......))))))).((((((..........(((.....))).........)))))).)))))))))............ ( -21.41) >consensus GUGCAGAAGGCUUAUAUACAAAAUAUAGUUUGUAUUUCUCUUCCUACACUUUGACAUAAUGUUUAUAAAAAAAAGAGAUUCUUCUGCAGAGAAAUAUAAU .(((((((((.....(((((((......))))))).((((((..........(((.....))).........)))))).)))))))))............ (-17.46 = -18.24 + 0.78)


| Location | 14,658,130 – 14,658,230 |
|---|---|
| Length | 100 |
| Sequences | 3 |
| Columns | 100 |
| Reading direction | reverse |
| Mean pairwise identity | 95.33 |
| Mean single sequence MFE | -22.82 |
| Consensus MFE | -19.72 |
| Energy contribution | -20.17 |
| Covariance contribution | 0.45 |
| Combinations/Pair | 1.04 |
| Mean z-score | -3.17 |
| Structure conservation index | 0.86 |
| SVM decision value | 1.23 |
| SVM RNA-class probability | 0.933213 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
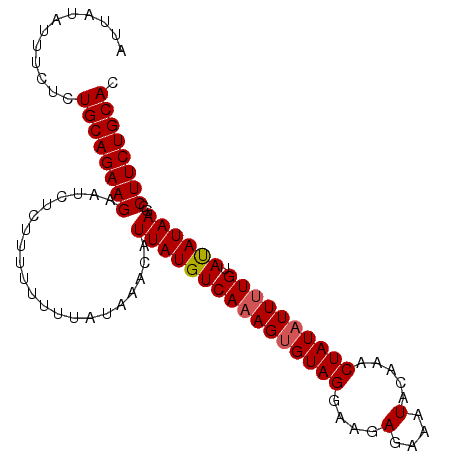
>3L_DroMel_CAF1 14658130 100 - 23771897 AUUAUAUUUCUCUGCAGAAGAAUUUCUUUUUUUUUUAAACAUUAUGUCAAAGGGUAGGAAGAGAAAUUCAAACUAUAUUAUGUAUAUAAGCCUUCUGCAC ............(((((((((((((((((((((......................))))))))))))))....(((((.....)))))....))))))). ( -20.55) >DroSec_CAF1 7910 99 - 1 AUUAUAUUUCUCUGCAGAAGAAUCUCUUU-UUUUAUAAACAUUAUGUCAAAGUGUAGGAAGAGAAAUACAAACUAUAUUUUGUACAUAAGCCUUCUGCAC ............((((((((......(((-......)))..((((((((((((((((....(....).....)))))))))).))))))..)))))))). ( -23.90) >DroSim_CAF1 7959 100 - 1 AUUAUAUUUCUCUGCAGAAGAAUCUCUUUUUUUUAUAAACAUUAUGUCAAAGUGUAGGAAGAGAAAUACAAACUAUAUUUUGUAUAUAAGCCUUCUGCAC ............((((((((..(((((((((.......(((((.......)))))))))))))).(((((((......)))))))......)))))))). ( -24.01) >consensus AUUAUAUUUCUCUGCAGAAGAAUCUCUUUUUUUUAUAAACAUUAUGUCAAAGUGUAGGAAGAGAAAUACAAACUAUAUUUUGUAUAUAAGCCUUCUGCAC ............((((((((.....................((((((((((((((((....(....).....)))))))))).))))))..)))))))). (-19.72 = -20.17 + 0.45)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:46:41 2006