
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 14,540,351 – 14,540,477 |
| Length | 126 |
| Max. P | 0.914881 |

| Location | 14,540,351 – 14,540,454 |
|---|---|
| Length | 103 |
| Sequences | 6 |
| Columns | 114 |
| Reading direction | reverse |
| Mean pairwise identity | 79.52 |
| Mean single sequence MFE | -19.28 |
| Consensus MFE | -13.35 |
| Energy contribution | -13.38 |
| Covariance contribution | 0.03 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.05 |
| Structure conservation index | 0.69 |
| SVM decision value | 1.10 |
| SVM RNA-class probability | 0.914881 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 14540351 103 - 23771897 GCAUGC-ACACAGAGACAAAGCCAUCUUGAGAAAUAAUUAAGAUGUAUGUAAAUGGAGAAAAUGCUGAAAUUGUUAAAAGCAUAAUAUUUCUUUGGA----GGC------GCUA ((..((-...((((((......((((((((.......))))))))................(((((............)))))......))))))..----.))------)).. ( -20.40) >DroVir_CAF1 18474 98 - 1 A----------------AAAACCAUCUUGAGAAAUAAUUAAGAUGUAUGUAAAUGGAGAAAAUGCUAAAAUUGUUAAAAGCAUAAUAUUUCUUUGGCUACAGCCCCGAGCACCC .----------------.....((((((((.......))))))))..((((..(..((((((((((............)))))....)))))..)..))))((.....)).... ( -18.30) >DroPse_CAF1 15541 103 - 1 GUACGCCACACAU-GGCAAAAGCAUCUUGAGAAAUAAUUAAGAUGUAUGUAAAUGGAGAAAAUGCUAAAAUUGUUAAAAGCAUAAUAUUUCUUUGGA----GGA------GCUG .(((((((....)-)))....(((((((((.......)))))))))..)))...((((((((((((............)))))....)))))))...----...------.... ( -21.90) >DroGri_CAF1 18057 98 - 1 A----------------AAAACCAUCUUGAGAAAUAAUUAAGAUGUAUGUAAAUGGAGAAAAUGCUAAAAUUGUUAAAAGCAUAAUAUUUCUUUGGCUACCACCCCGAGCAACA .----------------.....((((((((.......))))))))...(((..(..((((((((((............)))))....)))))..)..))).............. ( -15.70) >DroWil_CAF1 141733 88 - 1 A----------------AAAACCAUCUUGAGAAAUAAUUAAGAUGUAUGUAAAUGGAGAAAAUGCUAAAAUUGUUAAAAGCAUAAUAUUUCUUAAGA----GGA------GCCA .----------------....((.((((((((((((.......(.(((....))).)....(((((............)))))..))))))))))))----)).------.... ( -17.50) >DroPer_CAF1 19053 103 - 1 GUACGCCACACAU-GGCAAAAGCAUCUUGAGAAAUAAUUAAGAUGUAUGUAAAUGGAGAAAAUGCUAAAAUUGUUAAAAGCAUAAUAUUUCUUUGGA----GGA------GCUA .(((((((....)-)))....(((((((((.......)))))))))..)))...((((((((((((............)))))....)))))))...----...------.... ( -21.90) >consensus A________________AAAACCAUCUUGAGAAAUAAUUAAGAUGUAUGUAAAUGGAGAAAAUGCUAAAAUUGUUAAAAGCAUAAUAUUUCUUUGGA____GGA______GCCA ......................((((((((.......))))))))........(((((((((((((............)))))....))))))))................... (-13.35 = -13.38 + 0.03)



| Location | 14,540,375 – 14,540,477 |
|---|---|
| Length | 102 |
| Sequences | 6 |
| Columns | 114 |
| Reading direction | reverse |
| Mean pairwise identity | 83.45 |
| Mean single sequence MFE | -25.11 |
| Consensus MFE | -14.77 |
| Energy contribution | -15.67 |
| Covariance contribution | 0.89 |
| Combinations/Pair | 1.19 |
| Mean z-score | -2.74 |
| Structure conservation index | 0.59 |
| SVM decision value | 0.13 |
| SVM RNA-class probability | 0.598079 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
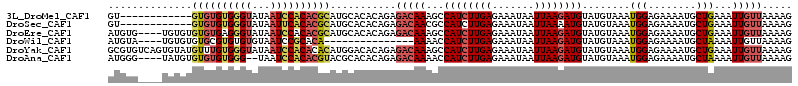
>3L_DroMel_CAF1 14540375 102 - 23771897 GU------------GUGUGUGGGUAUAAUCCACACGCAUGCACACAGAGACAAAGCCAUCUUGAGAAAUAAUUAAGAUGUAUGUAAAUGGAGAAAAUGCUGAAAUUGUUAAAAG ((------------((((((((((...))))))))))))(((..((...(((....((((((((.......))))))))..)))...)).......)))............... ( -26.00) >DroSec_CAF1 14586 102 - 1 GU------------GUGUGUGGGUAUAAUUCACACGCAUGCACACAGAGACAACGCCAUCUUGAGAAAUAAUUAAAAUGUAUGUAAAUGGAGAAAAUGCUGAAAUUGUUAAAAG ((------------((((((((((...)))))))))))).....(((........((((.(((....(((.........))).)))))))........)))............. ( -17.69) >DroEre_CAF1 14976 110 - 1 AUGUG----UGUGUGUGUGAGGGUAUAAUCCACACGCAUGCACACAGAGACAAAGCCAUCUUGAGAAAUAAUUAAGAUGUAUGUAAAUGGAGAAAAUGCUGAAAUUGUUAAAAG .((((----((((((((((.((((...)))).))))))))))))))..((((((((((((((((.......))))))))(((....)))........)))....)))))..... ( -34.30) >DroWil_CAF1 141757 95 - 1 AUGUA----UGUGUGUGCGUGUGUGUAAUCCGCACA---------------AAAACCAUCUUGAGAAAUAAUUAAGAUGUAUGUAAAUGGAGAAAAUGCUAAAAUUGUUAAAAG ..(((----(((.((((((..(.....)..))))))---------------...))((((((((.......))))))))................))))............... ( -18.30) >DroYak_CAF1 16000 114 - 1 GCGUGUCAGUGUAUGUUUGUGGGUAUAAUCCACACACAUGGACACAGAGACAAAGCCAUCUUGAGAAAUAAUUAAGAUGUAUGUAAAUGGAGAAAAUGCUGAAAUUGUUAAAAG ((((.((.((.(((((.(((((((...))))))).))))).)).((...(((....((((((((.......))))))))..)))...))..))..))))............... ( -28.90) >DroAna_CAF1 13179 108 - 1 AUGGG----UAUGUGUGUGUGGG--UAAUCCACACGUACGCACACAGAGACAAAACCAUCUUGAGAAAUAAUUAAGAUGUAUGUAAAUGGAGAAAAUGCUAAAAUUGUUAAAAG ...((----((((((((((((.(--(.......)).)))))))))....(((....((((((((.......))))))))..)))...........))))).............. ( -25.50) >consensus AUGUG____UGUGUGUGUGUGGGUAUAAUCCACACGCAUGCACACAGAGACAAAGCCAUCUUGAGAAAUAAUUAAGAUGUAUGUAAAUGGAGAAAAUGCUGAAAUUGUUAAAAG ..............((((((((((...))))))))))...........(((((...((((((((.......))))))))........(((........)))...)))))..... (-14.77 = -15.67 + 0.89)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:45:54 2006